Transcription factor NKX2-1 drives serine and glycine synthesis addiction in cancer
Published in Cancer

BACKGROUND
As cancer researchers, we have always been fascinated by the remarkable ability of cancer cells to adapt and thrive in the face of adversity. One of the most striking examples of this is the way in which cancer cells reprogram their metabolism to fuel their growth and ensure their survival (1). During a process known as tumorigenesis, cancer cells shift their energy metabolism to meet their ever-increasing demands for nutrients, including the production of certain amino acids. One such example is the synthesis of serine and glycine, two important building blocks for the proliferation and survival of cancer cells (2).
In contrast to non-transformed cells, which take up serine and glycine from their environment, some cancer cells prefer to synthesize their own de novo serine and glycine. The serine/glycine synthesis pathway branches from glycolysis via the glycolytic intermediate 3-phospho-glycerate, which can be converted into serine via three enzymatic reactions catalyzed by PHGDH, PSAT1, and PSPH, followed by the reversible interconversion of serine into glycine by SHMT1/2 (Figure 1)(2).
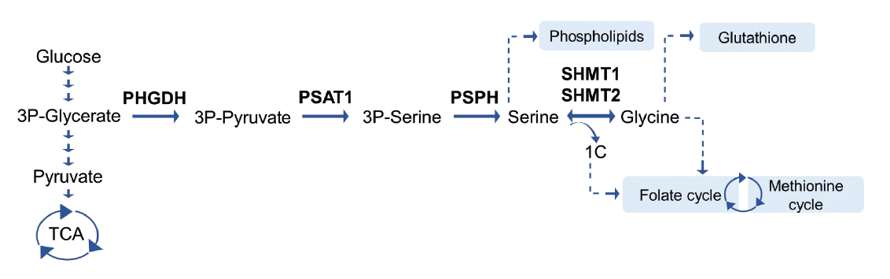
Figure 1. Schematic overview of the serine/glycine synthesis pathway and downstream metabolites.
But why are certain cancer cells so crazy about serine and glycine? As it turns out, these two amino acids are crucial building blocks for a variety of metabolic pathways that are essential for cancer cell proliferation and survival. Serine and glycine are necessary for the production of redox homeostasis molecules, ATP, nucleotides, and phospholipids, and are also involved in DNA and histone methylation (2). And because cancer cells often have to survive in nutrient-scarce environments, being able to produce these building blocks can provide them with a significant advantage.
In recent years, a growing number of oncogenic genomic alterations that promote serine synthesis have been identified. For instance, our group recently showed that a mutation in the ribosomal RPL10 gene, known as RPL10 R98S, enhances serine/glycine synthesis in T-cell leukaemia by increasing both PSPH transcriptional and translational levels (3). Interestingly, we noted that expression levels of serine/glycine synthesis enzymes are elevated by at least 4-fold in 80% of all T-cell leukaemia’s, whereas the RPL10 R98S mutation is only present in 8% of the samples, suggesting that other factors may be driving serine/glycine synthesis addiction in this type of cancer.
To find out what else might be at play, we employed a mixture of public dataset analyses, in vitro and in vivo experimental models and discovered a potential driver: transcription factor NKX2-1 overexpression. In this “behind the paper”, we will guide you through our main findings and the implications for cancer treatment.
KEY FINDINGS
To start, we analysed publically available sequencing data of cancer patient samples and found that the expression of serine and glycine synthesis enzymes was significantly increased in T-cell leukaemia patient samples that present NKX2-1 overexpression (Figure 2). NKX2-1 is a transcription factor that is normally not expressed in developing T-cells, but 5% of T-cell leukaemia samples show re-expression due to chromosomal rearrangements or genomic amplifications acquired during tumorigenesis (4). Interestingly, NKX2-1 alterations are also observed in 12% of lung adenocarcinoma (5) and 16% of neuroendocrine prostate cancer patients (6), and NKX2-1 overexpression is associated with increased expression of serine and glycine synthesis enzymes in these cancer types as well.
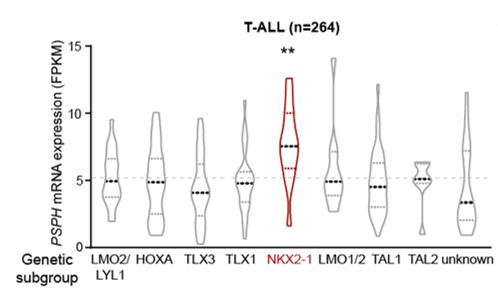
Figure 2. PSPH mRNA expression levels according to genetic subgroup in 264 T-ALL patients (P = 0.0014; Dunnett’s multiple comparisons test).
Next, we demonstrated in T-cell leukemia and lung cancer cell models that NKX2-1 binds and transcriptionally upregulates serine/glycine synthesis enzyme genes including phosphoserine phosphatase (PSPH) and serine hydroxymethyltransferase (SHMT). Fascinatingly, this binding and regulation even further increases when cancer cell have no access to serine and glycine.
Then, we explored whether the NKX2-1-driven increased expression of serine/glycine synthesis enzymes causes metabolic rewiring. To this end, we carried out mass spectrometry based glucose isotope tracing experiments, enabling us to track where the 6 labeled carbons from glucose are being used for. This analysis confirmed that NKX2-1 expressing cells synthesize more de novo serine and glycine (M+2/M+3) starting from glucose compared to control cells (Figure 3). In addition, we found that NKX2-1-driven serine/glycine synthesis mainly generates downstream nucleotides and redox molecule glutathione and subsequently, is associated with an altered cellular lipidome and methylome.

Figure 3. Schematic overview of 13C6-glucose isotope tracing and metabolic tracer analysis, measuring labelled serine/glycine in empty vector control versus NKX2–1 overexpressing Ba/F3 cells (n=3).
Given the metabolic advantages conferred by NKX2-1 overexpression, we investigated whether this trait could provide the cells with a proliferation benefit. When cultured under conditions of sufficient exogenous serine and glycine, NKX2-1 overexpression did not confer a significant proliferation advantage compared to control cells. However, since many cell lines are strictly dependent on exogenous serine for proliferation, we hypothesized that NKX2-1 overexpression could overcome this limitation. Indeed, when cultured in absence of serine/glycine, control cells exhibited impaired proliferation, while NKX2-1 overexpressing cells continued to proliferate, showing that increased serine/glycine synthesis enables cancer cells to become independent of exogenous serine for proliferation.
In a next step, we wanted to investigate the effects of NKX2-1 expression on tumour metabolism in an in vivo setting. In the past, NKX2–1 has often been referred to as a double-edged sword, given its described role as both a tumour suppressor as well as an oncogene (7). We also encountered this enigma in our in vivo experiments, making this one of the most challenging parts of the paper. We observed that at early stages of tumour progression, NKX2-1 functions as a tumor suppressor, but later becomes an aggressive oncogene, contributing to cancer progression when nutrients such as serine and glycine become scarce. Consistent with our earlier findings, NKX2-1 tumor-bearing mice showed systemically elevated reductive glutathione serum levels.
Now, armed with this knowledge, we wanted to verify whether NKX2-1 overexpressing cancer cells could be sensitive for metabolically interfering drugs since targeted treatments are crucial for improving treatment outcomes by reducing toxicity and preventing therapy resistance. Interestingly, our lab recently identified the widely clinically used anti-depressant sertraline as a potent SHMT inhibitor that selectively inhibits proliferation of serine synthesis addicted cancer cells (8). Indeed, we could show that sertraline, alone or even more effectively in combination with a serine/glycine-free diet, significantly decreased disease progression in NKX2–1 positive T-cell leukemia in vivo (Figure 4). Additionally, we showed that etoposide, a topoisomerase II inhibitor that restrains DNA replication, effectively inhibits proliferation of NKX2–1 overexpressing cells. Conformingly, while NKX2-1 positive LUAD is associated with early disease progression, etoposide treated patients with high NKX2-1 expression present improved outcomes.
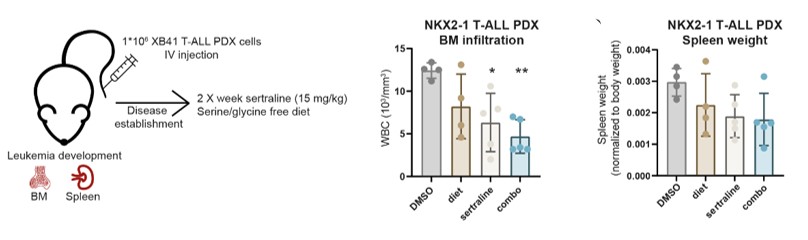
Figure 4. Schematic overview of the experimental set-up to test the effects of sertraline treatment and/or a serine/glycine-free diet on T-cell leukaemia disease progression. White blood cell counts (WBC) measured at disease end stage in the bone marrow (BM) of femurs and spleen weight normalised to body weight of the leukemic mice (n = 4 for DMSO and diet group; n = 5 for sertraline and combo group).
TAKE HOME MESSAGE
Reprogramming of cancer cell metabolism is a fascinating and complex process, and the synthesis of serine and glycine by cancer cells is a striking example of this. Serine and glycine are critical building blocks for various metabolic pathways essential for cancer cell survival and proliferation. In our recent research, we discovered that NKX2-1 overexpression drives increased expression of serine and glycine synthesis enzymes across cancers. This increased synthesis provides these cancer cells with a significant advantage in nutrient-scarce environments, enabling them to overcome the dependence on exogenous serine for proliferation and survival. Our findings identify NKX2–1 tumours as an extra subset of cancers that could benefit from the therapeutic targeting of serine/glycine synthesis hyperactivation with drugs such as sertraline. We hope our work may contribute to the development of personalized and effective treatments for cancer patients, and we look forward to further investigations into the role of NKX2-1 in tumorigenesis.

Figure 5. Transcription factor NKX2–1 fuels cancer cell proliferation and survival by hyperactivating serine/glycine synthesis, highlighting this pathway as a novel therapeutic target in NKX2–1-positive cancers.
REFERENCES
- Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23:27–47.
- Geeraerts SL, Heylen E, De Keersmaecker K, Kampen KR. The ins and outs of serine and glycine metabolism in cancer. Nat Metab. 2021;3:131–41. https://www.nature.com/articles/s42255-020-00329-9
- Kampen KR, Fancello L, Girardi T, Rinaldi G, Planque M, Sulima SO, et al. Translatome analysis reveals altered serine and glycine metabolism in T-cell acute lymphoblastic leukemia cells. Nat Commun. 2019;10:2542. https://www.nature.com/articles/s41467-019-10508-2
- Homminga I, Pieters R, Langerak AW, de Rooi JJ, Stubbs A, Verstegen M, et al. Integrated transcript and genome analyses reveal NKX2-1 and MEF2C as potential oncogenes in T cell acute lymphoblastic leukemia. Cancer Cell. 2011;19:484–97.
- Weir BA, Woo MS, Getz G, Perner S, Ding L, Beroukhim R, et al. Characterizing the cancer genome in lung adenocarcinoma. Nature. 2007;450:893–8.
- Baca SC, Takeda DY, Seo J-H, Hwang J, Ku SY, Arafeh R, et al. Reprogramming of the FOXA1 cistrome in treatment-emergent neuroendocrine prostate cancer. Nat Commun. 2021;12:1979.
- Yamaguchi T, Hosono Y, Yanagisawa K, Takahashi T. NKX2-1/TTF-1: an enigmatic oncogene that functions as a double-edged sword for cancer cell survival and progression. Cancer Cell. 2013;23:718–23.
- Geeraerts S, Kampen K, Rinaldi G, Gupta P, Planque M, de Cremer K, et al. Repurposing the antidepressant sertraline as SHMT inhibitor to suppress serine/glycine synthesis addicted breast tumor growth. Mol Cancer Ther. 2020;20:50–63. https://aacrjournals.org/mct/article/20/1/50/93024/Repurposing-the-Antidepressant-Sertraline-as-SHMT
Follow the Topic
-
British Journal of Cancer
This journal is devoted to publishing cutting edge discovery, translational and clinical cancer research across the broad spectrum of oncology.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in