Transcriptomes, fossils and singing insects
Published in Ecology & Evolution
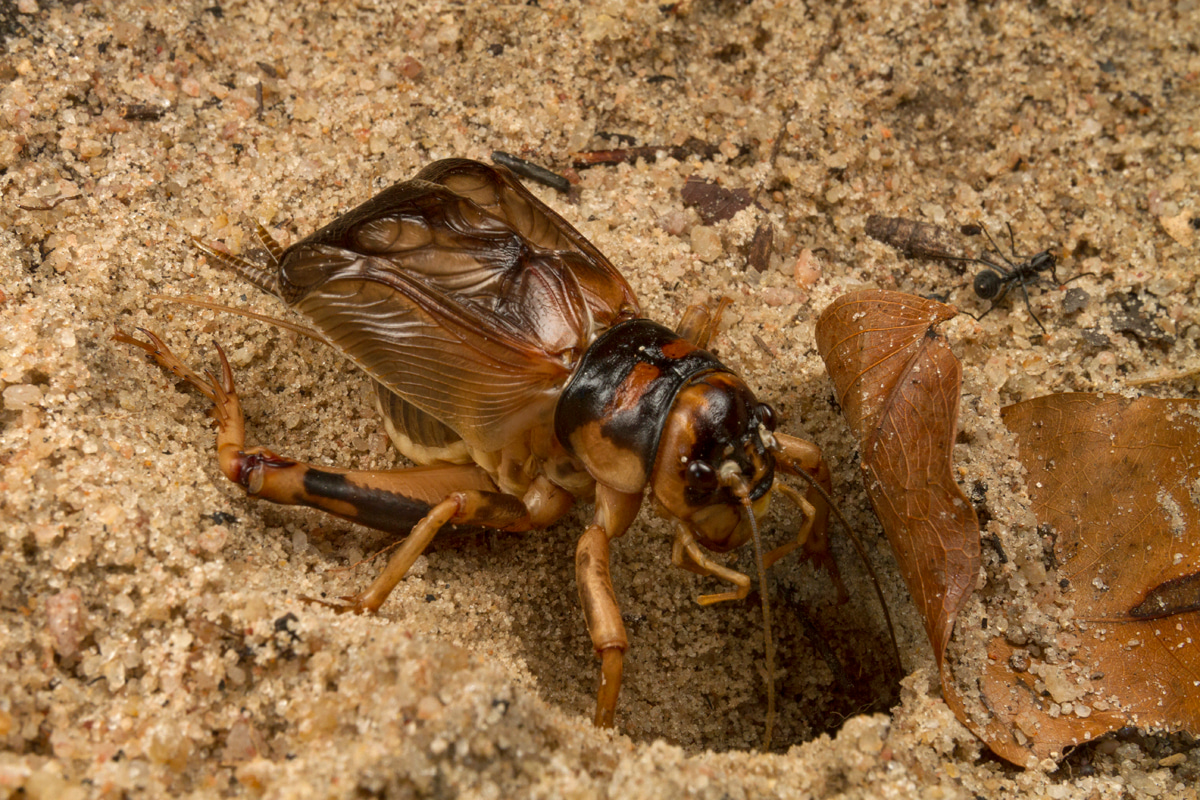

Take a walk at summer night and you will undoubtedly hear crickets chirping. In fact, the soundscape at night around the world is filled with the songs of orthopteran insects. Orthoptera is an insect order that contains familiar insects, such as crickets, katydids, and grasshoppers, that are characterized by the ability to jump. Out of ~28,000 described species, more than 16,000 species of Orthoptera primarily use acoustic signalling as the main mode of sexual communication, outnumbering frogs, birds, mammals, or any of the known acoustically-active lineages. Moreover, Orthoptera is the earliest known lineage of animals to have evolved complex acoustic communication, as evidenced by fossil forewings with stridulatory apparatus, known from as early as the Triassic. Therefore, by studying how acoustic communication evolved in Orthoptera, we can gain valuable insights into the evolution of animal communication.
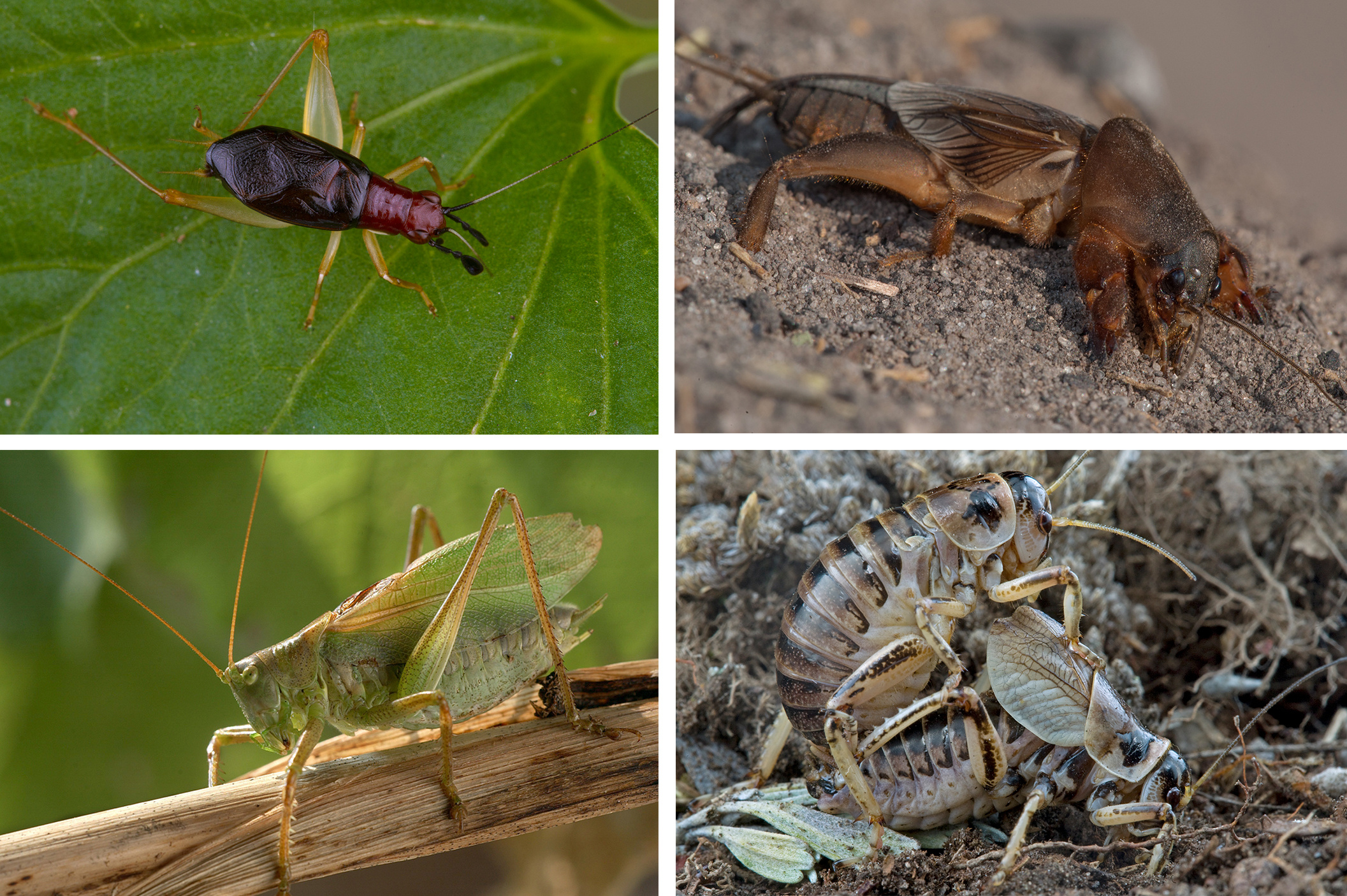

Our new study published in Nature Communications takes a phylogenomic approach to address how hearing organs and sound-producing organs originated and diversified within this amazing group of insects. Here is the take-home message of our paper: hearing and sound production organs co-evolved via a Fisherian mechanism in Ensifera (katydids, crickets, and allies), but in Caelifera (grasshoppers), hearing evolved first probably in the context of flight modulation, and later co-opted for acoustic signalling when some groups evolved stridulatory apparatus. Also, the evolution of acoustic communication did not necessarily increase the diversification rates of singing lineages within Orthoptera.
In our study, we partnered with the 1KITE project to generate phylogenomic data to recover the most robust phylogenetic relationships of Orthoptera to date. Our partnership was rather serendipitous, and something that was not originally planned. But before going further, let us briefly introduce what the 1KITE project is. The “1K Insect Transcriptome Evolution (1KITE) project” is an international research initiative that started in 2011 led by the Beijing Genomics Institute in Shenzen in China and the Museum Koenig in Bonn in Germany to study the insect evolution by generating and analyzing 1,000+ de novo transcriptomes from across the phylogeny of insects. This project brought together internationally recognized experts from around the globe in molecular biology, morphology, paleontology, embryology, bioinformatics, and scientific computing in a yet unparalleled way. Since then, the 1KITE team organized numerous international field trips sampling key insect species, organized numerous meetings e.g. in Germany, Austria or the USA, spent thousands of computation hours, and established several workflows and collaborations in order to sequence and evaluate the transcriptomes of more than 1,400 insect species representing all major insect lineages. The results of these endeavors greatly enhanced our understanding of insect evolution by numerous published studies across the Insect Tree of Life, including the landmark study on the phylogeny of the class Insecta, as well as lineage-specific studies on Hymenoptera (bees, ants, wasps), Coleoptera (beetles), Lepidoptera (butterflies, moths), Polyneoptera (cockroaches, stoneflies, stick insects, grasshoppers, etc.), Phasmatodea (stick insects), Blattodea (cockroaches, termites), and Dermaptera (earwigs).
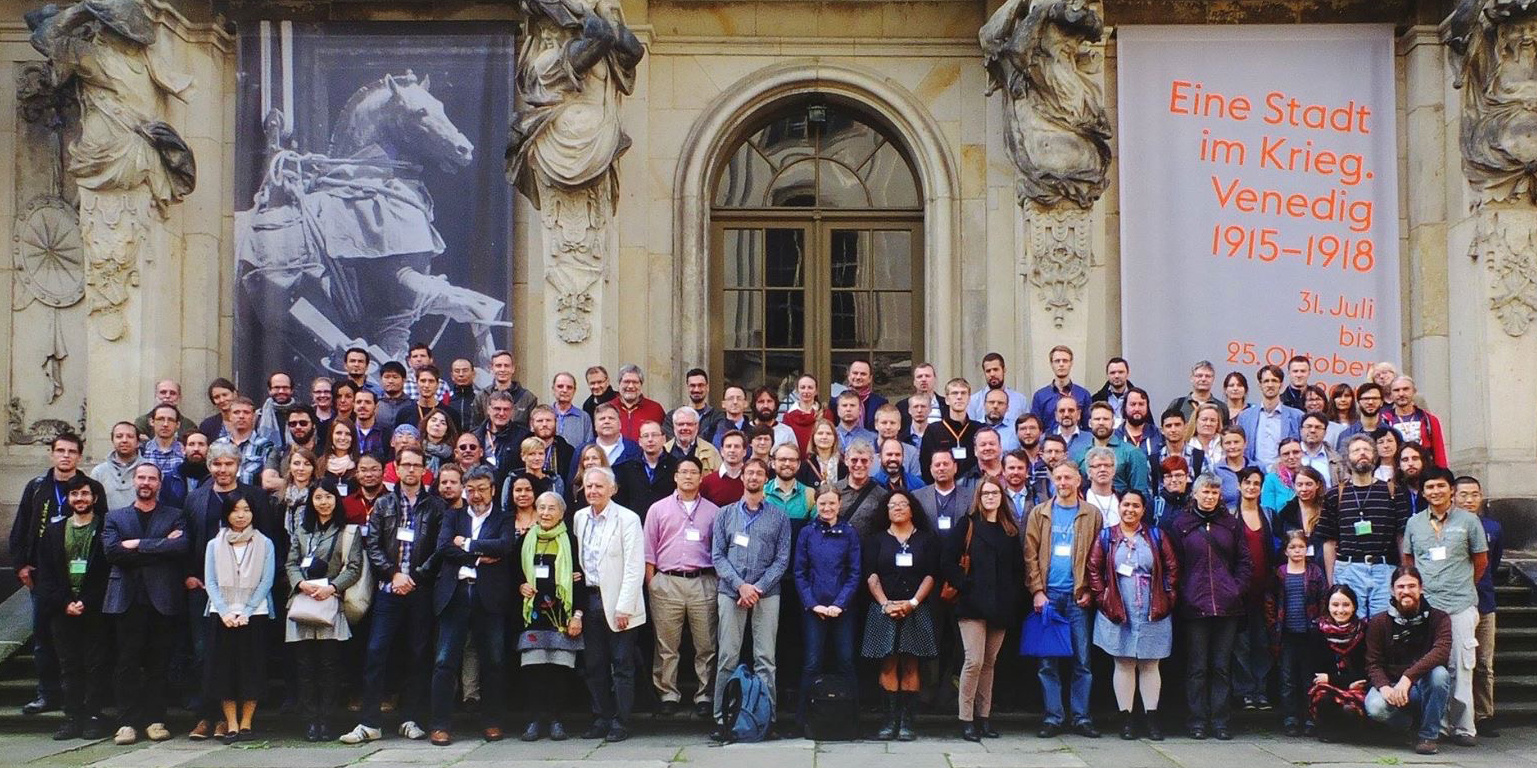

In September 2015, there was a small conference in Dresden, Germany called the “Dresden Meeting on Insect Phylogeny”, a biennial meeting on insect phylogenetics. This particular meeting had many presentations from the 1KITE project, particularly because the project’s first major output was just published in 2014 as a cover of the journal Science. When the 1KITE project was launched, it included several subprojects focusing on different insect lineages, but a project specifically focusing on the phylogeny of Orthoptera was not in the original plan. During the Dresden meeting, we agreed that a new project on Orthoptera would be very important, and discussed a possibility of collaborating by bringing together expertise in orthopteran systematics, phylogenomics, and paleontology. This was the beginning of our project, which came to fruition five years later.
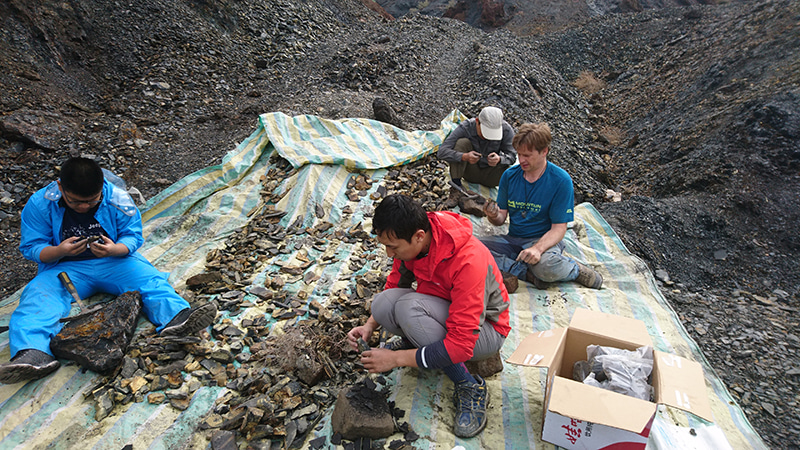

One of the major findings of our study is the antiquity of the Orthoptera, which has major implications for the origin of acoustic communication. The earliest known fossil Ensifera is Raphogla rubra found in a Permian (approx. 270 million years old) locality in Southern France near by the Salagou Lake. Interestingly, our dating analysis provided much more ancient ages for this group and other major taxa within Orthoptera. On the other hand, many groups of fossil Orthoptera are difficult to relate to extant forms. For example, it is yet unclear whether Permian species possessing a long ovipositor, the name 'Ensifera' refers to, are either stem-Orthoptera, stem-Ensifera, or stem-Caelifera. Bearing this possibility in mind, the ages delivered by the analysis became credible. Our study proposes that Orthoptera originated about 355 million years ago, and that both suborders originated in the Late Carboniferous.
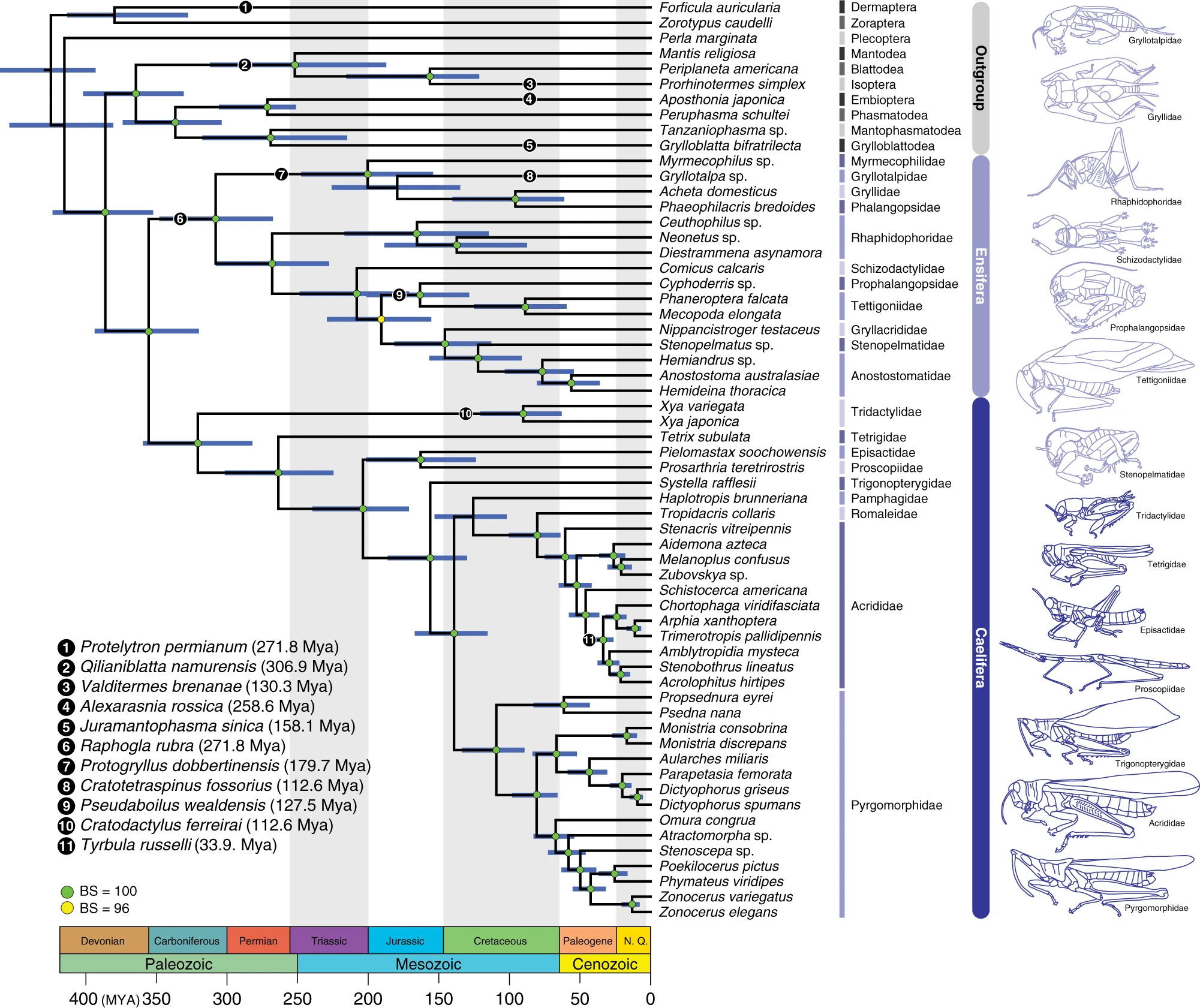
Dated phylogeny of Orthoptera based on the phylogenomic data. (Figure 1 from our new study).
If we could go back to the early Permian period (300 million years ago), we would see large synapsids (proto-mammals) roaming around, but if you pay attention at night, there would be ancient crickets chirping by rubbing their wings to attract females that hear through tiny ears on their forelegs. It is amazing to think that the basic mechanisms of hearing and sound production in Ensifera we see today remained fundamentally unchanged over 300 million years. Something we don’t mention too much in the paper is the evolution of acoustic communication in katydids. About 70% of known katydid species produce ultrasound (20-150 kHz) for acoustic signalling, and all katydids have specialized hearing sensory organ known as crista acustica inside the tibial ears that are tuned to different frequencies tonotopically (the spatial mapping of frequency). Katydids originated and diversified during the Cretaceous, and this elaborate form of acoustic communication probably evolved because the soundscape at night was saturated with other singing animals (such as crickets and frogs). The evolution of ultrasound communication probably has nothing to do with bats because bats evolved much later in the Eocene.
One thing that has become very evident when we put together this study is how little we know about the mode of sound production and hearing, other than what’s known in crickets, katydids, and some grasshoppers. Particularly, our finding that the Old World family Pamphagidae, for which we know very little about its acoustic communication skills and habits, had an increased diversification rate compared to other lineages, illustrates that there are still many groups worth investigating in more depth.

So what’s next? Our study paints with a broad brush on the evolution of acoustic communication in Orthoptera, but we still have a long way to go to understand how hearing and sound producing evolved in individual lineages. For example, within crickets, there is a large range of variation in hearing and singing, and the same is true for katydids. Thus, the next step would be to apply phylogenomics to resolve the relationships within these singing lineages and to get into the nitty-gritty of how acoustic communication has been shaped through time. But for now, we celebrate the end of our long collaboration and cherish all of the memories of putting together this project.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in