Zika Virus: Master Membrane Manipulator
Published in Microbiology
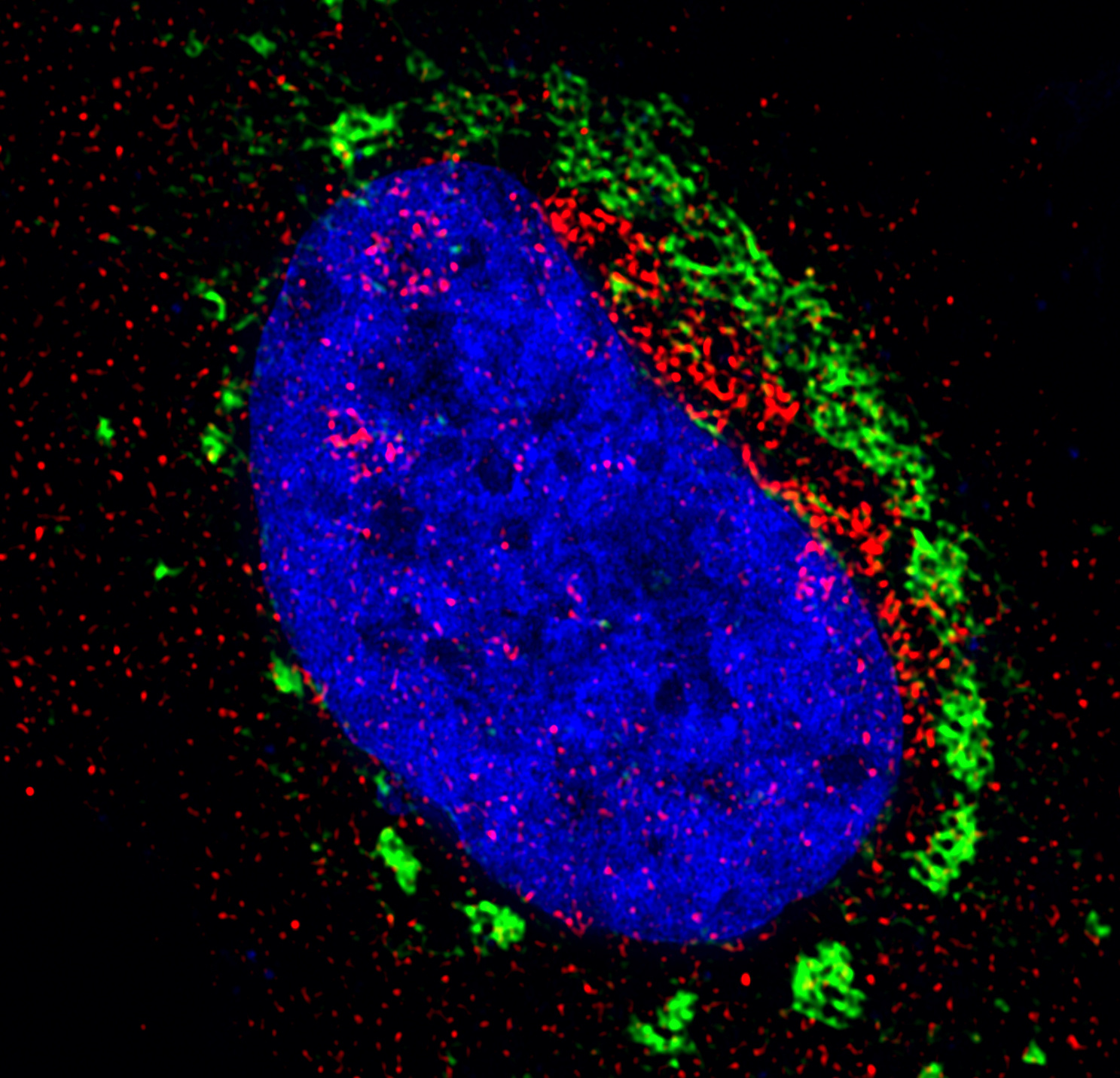
Our initial question was straightforward: Zika virus (ZIKV) churns out copies of itself within intricate organelle-like structures formed from the lipid membranes of its host. By depriving infected cells of key lipid types, could viral replication be disrupted? We were particularly interested in a family of bioactive membrane components called sphingolipids, both because we had a set of knockout cell lines and other tools on hand from previous studies, and because sphingolipids are highly enriched in the neuronal cells target by ZIKV. Sure enough, we found that treating cultured cells with inhibitors of critical sphingolipid biosynthesis enzymes reduced the shedding of infectious particles and decreased intracellular levels of viral RNA.
The first twist in this story came when we followed our inhibitor experiments by infecting a sphingomyelin synthase-1 (SGMS1) knockout cell line. SGMS1 converts ceramide, a key precursor to other sphingolipids, to sphingomyelin, the most abundant sphingolipid in mammalian cells. Unexpectedly, the knockout cells produced 100-200 times more infectious ZIKV than their wild-type control! After repeating the experiment a half-dozen times or so, we were forced to confront the possibility that our two main findings were in conflict with each other. Fortunately, a graph buried in the supplementary information of an earlier paper featuring our SGMS1 knockout cells hinted at a solution: in a mass spectrometry experiment to validate the gene deletion, the expected decrease in levels of sphingomyelin was accompanied by a noticeable increase in levels of ceramide. Could a buildup of ceramide in cells lacking the enzyme to metabolize it be having a pro-viral effect? Gratifyingly, follow-up experiments showed this to be the case.
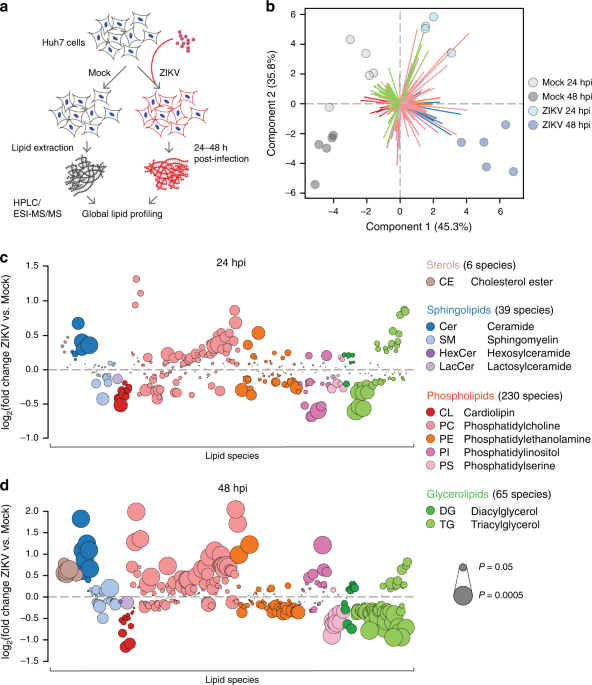
A second change in direction came with a lipidomics experiment we performed in 2018 with Edward Dennis’s group at the University of California, San Diego (of LIPID MAPS fame). This turned into a remarkably productive (and ongoing) collaboration with the multi-disciplinary omics team at Pacific Northwest National Laboratory. Up until this point, our project had been a fairly reductionist study of the relationship between ZIKV infection and sphingolipid metabolism; seeking the bigger picture, we carried out global surveys of the lipidomes of infected cells. Our previously virology-minded manuscript was now squarely in systems biology territory. What was eye-opening, though, beyond the trove of new data and potential directions for future studies, was how well the two approaches could mesh: our virology experiments were confirmed by omics data, and that data deepened our understanding of the mechanisms underlying our results. We concluded that ZIKV globally upregulates ceramide biosynthesis, then enriches its replication structures with the newly synthesized sphingolipids to its ultimate advantage.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in