By Pallavi Mathur & Kristine Schauer, illustrations: Apolline Boureau@EPSAA
Background: Cancer is a disease characterized by large scale genomic alterations that lead to a wide array of cancer phenotypes resulting from unwired cellular pathways normally under strict surveillance. Sequencing and molecular analysis have revealed a plethora of genes deregulated in cancers. The amount of deregulated genes presents a significant challenge to the understanding how they participate in cancer progression. Notably, genes do not function as an independent unit but are part of integrated, complex networks often referred as cellular pathways. These pathways can convene to the smallest functional unit of the cells i.e., the organelle. Organelles are self-organized subcellular compartments that carry out specific functions in the cells. They can be visualized and studied on their own. Therefore, studying organelles allows us to zoom out of the singular gene to the functional unit to which several genes belong. Widescale efforts have been made to establish atlases of genomic and proteomic alteration in cancer, however, no atlas of organelle-level changes has been profiled till date.
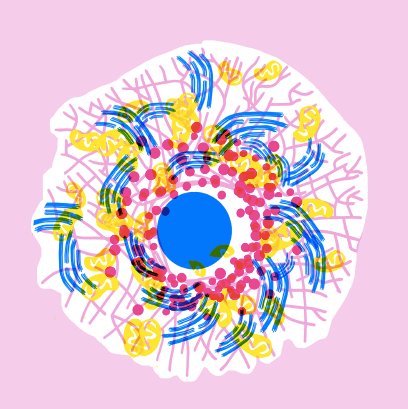

The image is an artistic illustration of a cell with its organelles. Organelles are the smallest functional units of cells found in our body. The nucleus and Golgi stacks are shown in blue, the lysosomes are in red, the mitochondria in yellow and the endoplasmic reticulum in rose. Although several atlases of genomic and proteomic alteration of cancer cells have been established, no atlas of organelle-level changes has been profiled till date. Illustration: Apolline Boureau@EPSAA
Hypothesis: In this study we tested the hypothesis that organelle organization is perturbed in cancer cells focusing on bladder cancer as our model. We have studied lysosomes as the organelle of interest. Indeed, lysosomes have recently emerged as important players in cancer development1,2, because they can support the high energetic demands of increased proliferation and/or motility of cancer cells. Lysosomes are dynamic organelles that act as the cellular hub for metabolism and signaling: in addition to their classical role of cell clearance through degradation and recycling of molecules, lysosomes act as nutrient sensors that adjust cellular metabolism depending on availability of the nutrients. They do this via important serine/threonine kinase complex called mammalian target of rapamycin complex 1 (mTORC1). mTORC1 is activated upon assembly on the surface of lysosomes in response to growth factors and amino acids to promote protein synthesis. Conversely, absence of nutrients inactivates and dissociates mTORC1 from lysosomes subsequently triggering catabolic pathways in cells. Since lysosomes are dynamic organelles that alter their organization depending on cellular needs, we decided to study the landscape of this organelle in bladder cell lines representing low-grade and high-grade bladder cancers in comparison to normal human urothelium (NHU) cells.
Findings: Employing the technique of normalized cell cultures on identical microscale adhesive surfaces that we call micropatterns3, we compared lysosomes of bladder cancer cells to those found in NHU. We observed that while in NHU cells lysosomes were positioned centrally, they were peripherally dispersed in bladder cancer cells with a stronger phenotype in high-grade cell lines. Lysosomal positioning has been described to regulate mTORC1 signaling4. Indeed, we found that although the fraction of mTORC1 localized on lysosomes was comparable between different cell lines, the mTORC1 substrate p70-S6 Kinase 1 (S6K1) was stronger phosphorylated in low-grade cancer cells whereas another mTORC1 substrate, eIF4E Binding Protein (4EBP1), was stronger phosphorylated in high-grade cells. Surprisingly, we found nuclear translocation, and thus activation, of transcription factor EB (TFEB), another important substrate of mTORC1, in high-grade cells besides activated mTORC1 signaling. In healthy cells, mTORC1 phosphorylates TFEB that leads to its cytosolic retention or inactivation. Thus, nuclear translocation of TFEB under conditions of active mTORC1 indicates a dysfunction of metabolic signaling in bladder cancer. TFEB acts as a master regulator of lysosomal biogenesis and autophagy and is a family member of MiT/TFE transcription factors that are implicated in several cancers such as renal cell carcinoma, pancreatic adenocarcinoma or sarcoma5.
Since TFEB hyperactivation correlated with peripheral lysosomes, we next questioned whether TFEB regulates lysosome positioning in high-grade cells. Indeed, we found that depletion of TFEB reversed the lysosomal dispersion phenotype, thus confirming that peripheral lysosomes are a result of TFEB hyperactivation.
Finally, we explored the mechanism through which TFEB regulates lysosomal dispersion in high-grade cells. Phosphoinositides and their associated enzymes are often implicated in cancers, for instance the phosphoinositide-3-kinase (PI3K class I) pathway is one of the most frequently activated signaling in tumorigenesis e.g. glioblastomas6. Since phosphatidylinositol-3-phosphate (PtdIns3P) is a regulator of lysosome positioning7, we studied whether TFEB controls the levels of PtdIns3P. We found that TFEB transcriptionally regulates class III PI3K (PIK3C3/ VPS34), and thus, PtdIns3P production on lysosomes. This facilitates binding of PtdIns3P-specific proteins to lysosomes, such as protrudin. Protrudin activates peripheral lysosome movement and its overexpression has been associated with peripheral lysosomal dispersion8.
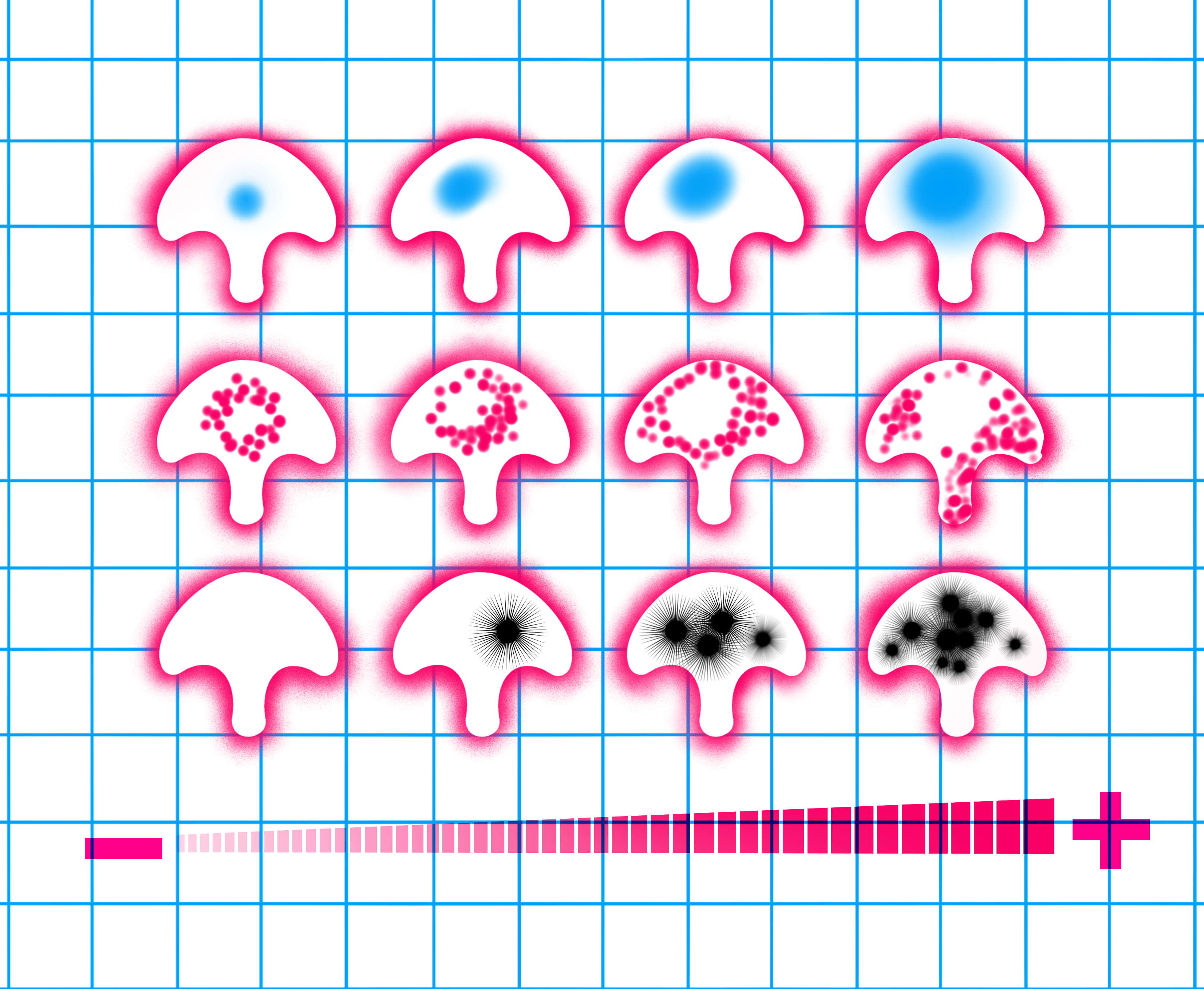

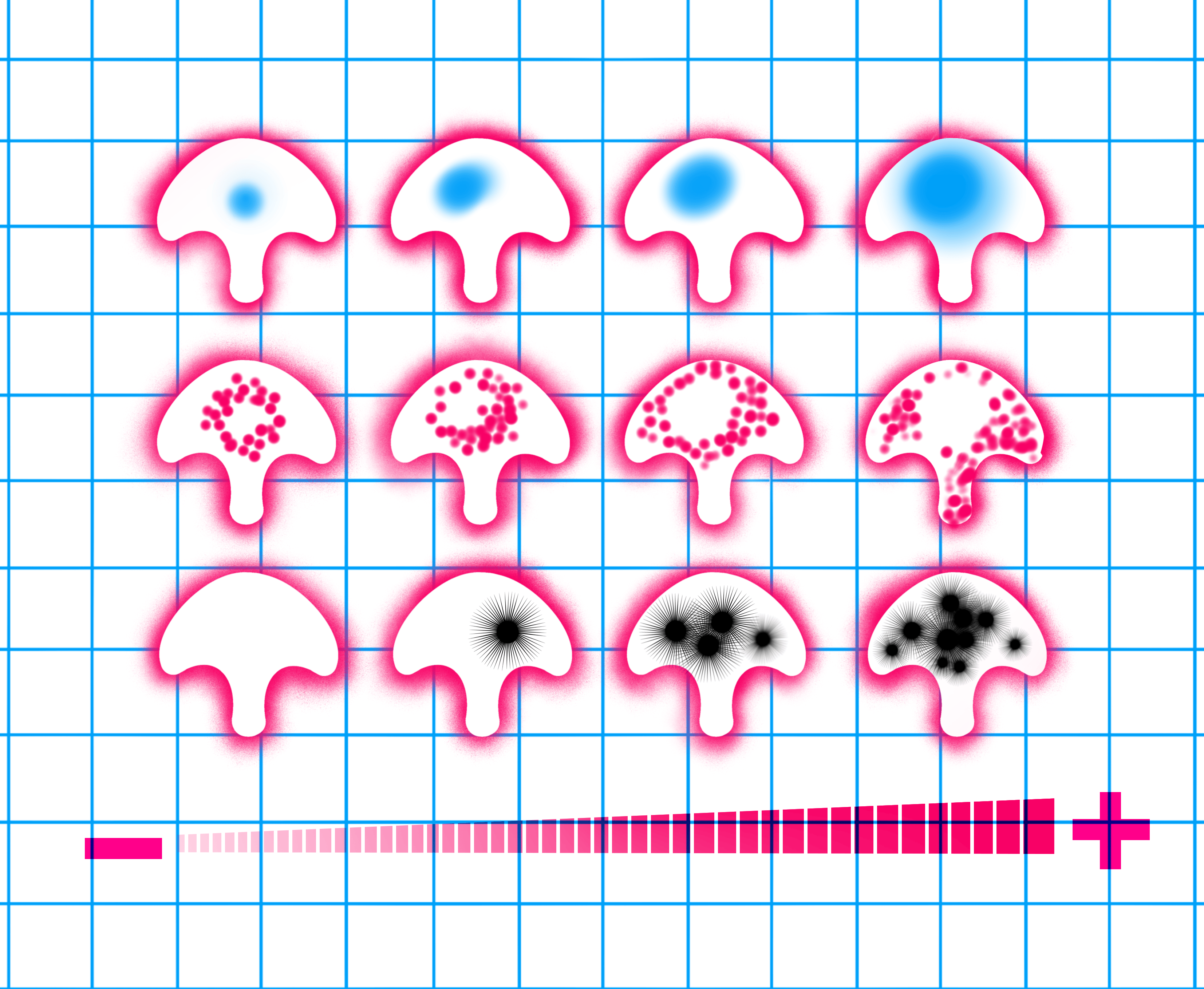
The image illustrates the comparison of normal human urothelium (NHU) cells, on the left, to bladder cancer cells of different grades, on the right, employing culturing techniques on identical microscale adhesive surfaces that we call micropatterns4. The nucleus is shown in blue, the lysosomes are in red and the black stars represent cancer progression. We observe a correlation between organelle changes and cancer progression indicated by – to +. Illustration: Apolline Boureau@EPSAA
What do we learn? We provide the first atlas for the landscape of the lysosomal compartment in the bladder cancer model. We reveal the mechanistic role of TFEB in regulating endosomal PtdIns3P levels and subsequent lysosomal dispersion. Thus, we unveiled lysosomal positioning as a potential biomarker for malignant bladder cancer with activated TFEB which might arise as an actionable target for cancer therapy. Pioneering studies, such as those from the lab of Dr. Marja Jäättelä and Dr. Rushika M. Perera, have focused on lysosomal pathway disruptions in cancers, which inspires the approach of targeting selected organelles to understand cancer progression. Our study presents a novel area of studying organelles and associated perturbations as a biomarker of cancer.
The image illustrates our approach to zoom on organelles to understand cancer progression. Lysosomal changes unveiled dysregulation of metabolic signaling in bladder cancer which might arise as an actionable target for cancer therapy. The endoplasmic reticulum is shown in blue, the lysosomes are in red and the mitochondria are in yellow. Illustration: Apolline Boureau@EPSAA
References:
- Perera, R. M. & Zoncu, R. The Lysosome as a Regulatory Hub. Annu. Rev. Cell Dev. Biol. 32, 223–253 (2016).
- Hämälistö, S. & Jäättelä, M. Lysosomes in cancer-living on the edge (of the cell). Curr. Opin. Cell Biol. 39, 69–76 (2016).
- Schauer, K. et al. Probabilistic density maps to study global endomembrane organization. Nat. Methods 7, 560–566 (2010).
- Korolchuk, V. I. et al. Lysosomal positioning coordinates cellular nutrient responses. Nat. Cell Biol. 13, 453–460 (2011).
- Perera, R. M., Di Malta, C. & Ballabio, A. MiT/TFE Family of Transcription Factors, Lysosomes, and Cancer. Annu. Rev. Cancer Biol. 3, 203–222 (2019).
- Liu, P., Cheng, H., Roberts, T. M. & Zhao, J. J. Targeting the phosphoinositide 3-kinase pathway in cancer. Nat. Rev. Drug Discov. 8, 627–644 (2009).
- Raiborg, C. et al. Repeated ER–endosome contacts promote endosome translocation and neurite outgrowth. Nature 520, 234–238 (2015).
- Hong, Z. et al. PtdIns3P controls mTORC1 signaling through lysosomal positioning. J. Cell Biol. 216, 4217–4233 (2017).
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
DNA repair and human disease
Publishing Model: Hybrid
Deadline: Oct 31, 2026
Cell death and inflammatory signalling
Publishing Model: Hybrid
Deadline: Oct 28, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in