Acyls lost, dimers arose
Published in Chemistry
The paper in Nature Communications is here: https://doi.org/10.1038/s41467-018-04487-z
Many natural products are isolated as dimers. Dimerization appears to be important to amplify the therapeutic potential of these compounds. But how these dimers are biosynthesized in most cases are not known. Limited examples have been shown that the C–C dimerizations are commonly catalyzed by cytochrome P450 enzymes as in the cases of himastatin, ditryptophenaline, julichrome Q, desertorins, orlandin, and cladofulvin, etc. Other modes of enzyme-mediated C–C dimerization are also known, such as an alkylating enzyme CylK-catalyzed formation of dimeric cylindrocyclophanes, and a putative enzymatic [2 + 2] cycloaddition reaction in forming diasteltoxins. In addition, the NmrA family of regulatory proteins have also implicated as the C−C dimerization enzymes in actinorhodins and lomaiviticins.
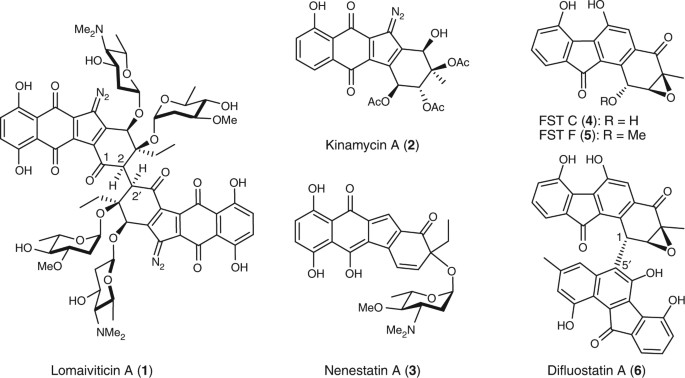
Lomaiviticin A and difluostatin A are benzofluorene-containing atypical angucycline dimers. Lomaiviticin A is highly toxic DNA cleavage agent and is currently undergoing preclinical evaluation for antitumor treatment. However, little is known about the dimerization mechanism to form the carbon–carbon (C–C) bonds in lomaiviticin A (C2/C2′, symmetric) and difluostatin A (C1/C5′, asymmetric). In a sharp contrast to a previous speculation that the dimerization of difluostatin A might be catalyzed by the NmrA family regulatory protein FlsQ1, this work demonstrates a non-enzymatic way to form difluostatin A.
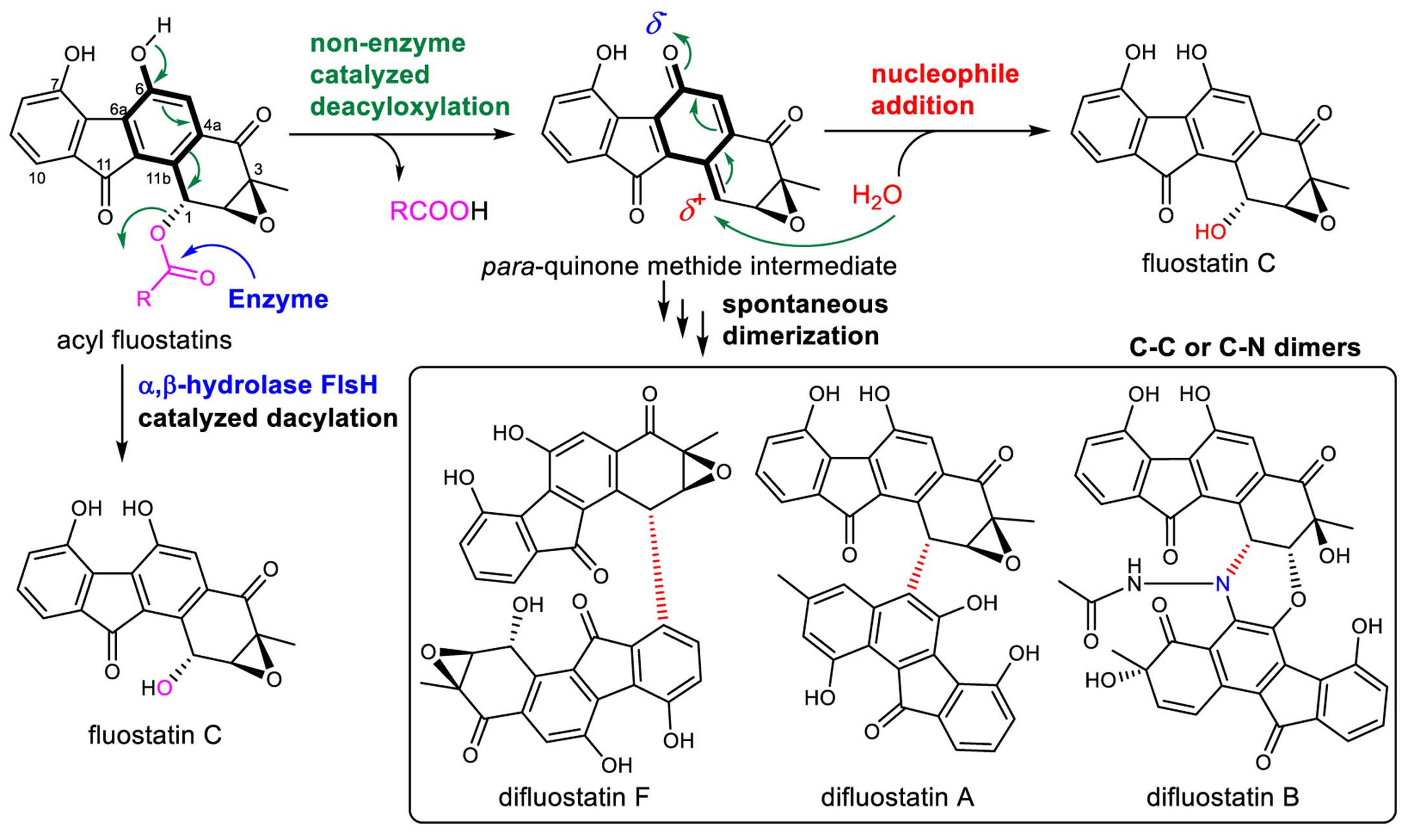
The heterologous expression of the fluostatin biosynthetic gene cluster from Micromonospora rosaria SCSIO N160 in Streptomyces albus J1074 afforded several new fluostatin heterodimers/trimers with diverse C–C and C–N-couplings. Serendipitously, acyl fluostatins were found to undergo spontaneous deacylation with simultaneous dimerization to form C–C coupled homodimers. The reaction is illustrated as a chemical process mediated by a reactive para-quinone methide (p-QM) species, which is formed via an autocatalytic 1,6-elimination of the acyl group in fluostatins. The p-QM species can serve as a promiscuous acceptor for 1,6-addition to form diverse C–C and C–N-coupled dimers. These findings have brought an efficient strategy for the facile synthesis of novel dimeric compounds as well as hybrid antibiotic conjugants. Intriguingly, a deacylase FlsH was characterized which may be evolved in the fluostatin pathway to prevent the accumulation of toxic p-QMs by enzymatic hydrolysis of the acyl group.
The results present in this paper highlight that many natural products could be artifacts if their assembly is non-enzyme catalyzed. Also, this work brings an interesting question to rethink about the proposed dimerization strategy for actinohordin and lomaiviticin A: enzymatic or non-enzymatic reaction? It would be highly interesting to unravel the complex biosynthetic enzymology of these pathways.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in