An unconventional vaccination strategy against influenza viruses from humans and swine
Published in Microbiology, Biomedical Research, and Immunology
The quest for universal influenza vaccines
The standard influenza vaccine contains at least three killed influenza virus strains, in professional terms known as A subtype H1N1, A subtype H3N2, and B. These vaccines induce antibodies against the major viral surface protein, the hemagglutinin (HA). Only these antibodies can “neutralize” the virus and prevent it from binding and entering cells of the host. One weak point is that they mainly bind to the most variable sites in the head or outer portion of the HA (Fig. 1). For this reason, the strains in the seasonal flu vaccines need to be updated every few years, because influenza viruses constantly evolve. They will also fail to protect against influenza viruses of animal origin, which may spark the next pandemic.
The lower stalk portion of the HA is an attractive target for broadly protective influenza vaccines, as it is conserved between all strains within a subtype, and even between some subtypes. Unfortunately, anti-stalk antibodies work in a different way than those against the HA head and they can only attack the virus after it has entered the host cell. They are also more difficult to induce, as the stalk is less easily seen by cells of the immune system.
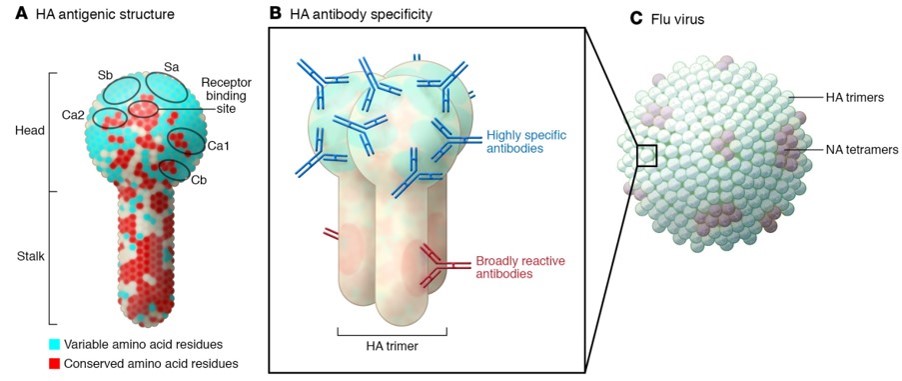
Fig. 1. The hemagglutinin (HA) protein of H1 influenza viruses. A The most variable antigenic determinants or “epitopes” are in five antigenic sites (Sa, Sb, Ca1, Ca2, Cb) in the head domain, these are “immunodominant”. Conserved epitopes are in the stalk and some parts of the head. B Strain-specific and broadly reactive antibodies bind to different regions. C HA and NA (neuraminidase) protrude from the surface of the virus. Their combination determines the subtype, such as H1N1, H3N2, H5N1, etc. Illustration from J Clin Invest. 2018;128(11):4751-4754.
The pig as an influenza model and source of zoonotic influenza
Pigs are naturally susceptible to the same influenza subtypes as humans, H1N1 and H3N2, and they are an underused influenza model. Pigs are also a source of novel influenza viruses for humans, and the 2009 H1N1 pandemic virus was of swine origin. The other way round, most swine-adapted viruses originate from those that once circulated in people. The evolution of influenza in swine is complex and unique, and there is simultaneous circulation of multiple H1 and H3 lineages, and many different strains within each lineage (Fig. 2). The three H1 lineages – 1A, 1B and 1C – show significant differences in the HA, between 27% and 30% of the amino acid sequence. This extraordinary diversity greatly hampers the control of swine influenza by vaccination. Human seasonal H1N1 viruses, in contrast, undergo smaller and stepwise changes over time, generally less than 6% over a decade.
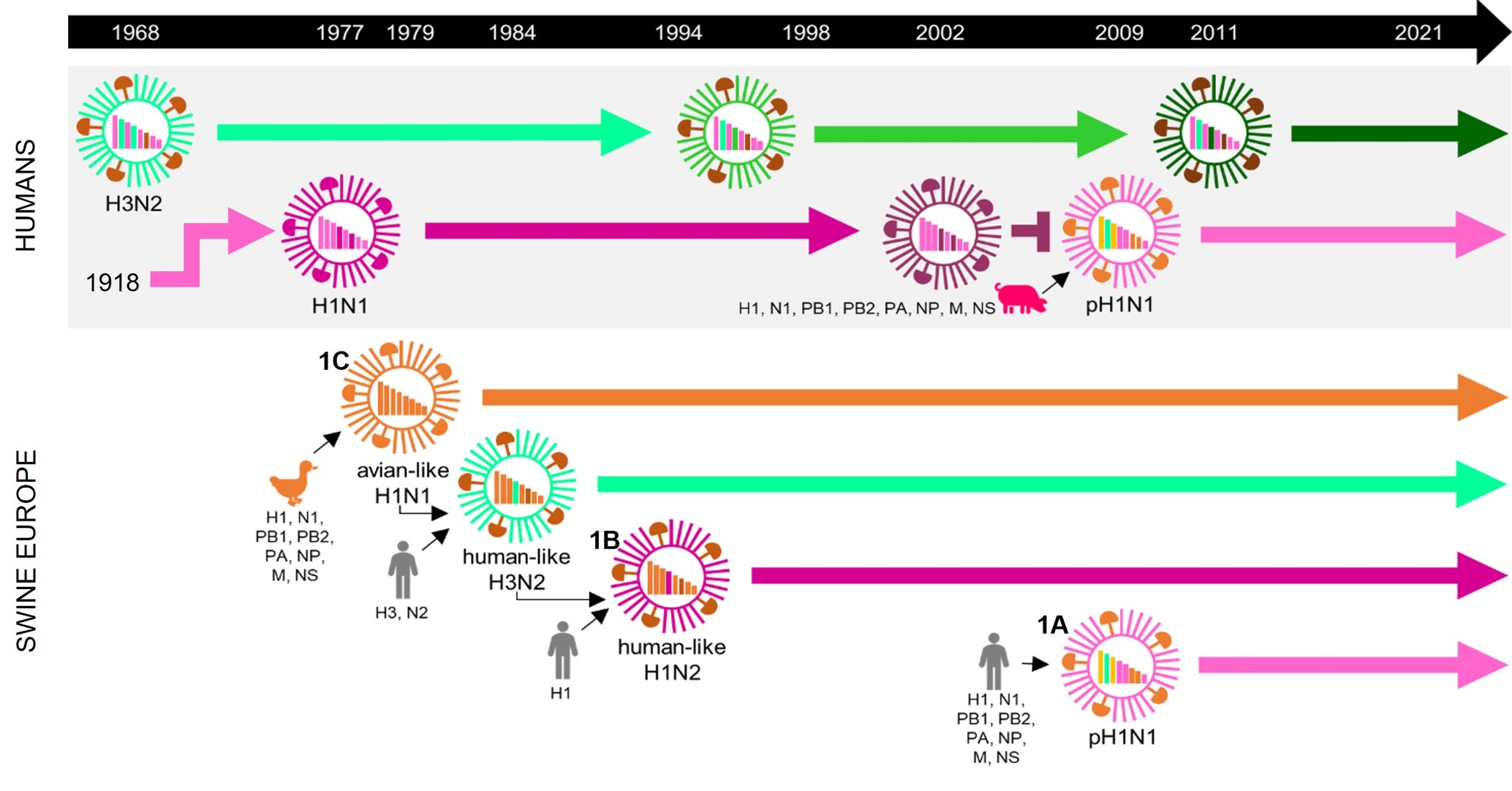
Fig. 2. Evolution of H1 influenza in humans and swine in Europe. In humans, a single virus strain (pH1N1) undergoes gradual changes. In swine, three different lineages of the H1 subtype circulate simultaneously: 1A, 1B and 1C. The prevailing lineages and strains differ between geographic regions. Image credit: Elien Vandoorn.
Exploiting the diversity of H1N1 swine influenza viruses for heterologous prime-boost vaccination
People and pigs need multiple doses of influenza vaccine: two injections one month apart for the initial immunization and yearly repeat vaccinations thereafter. A promising approach to broaden the immune response is to use divergent virus strains for the first (primary) and subsequent (booster) vaccinations. If such strains show little overlap in the most variable, most immunogenic epitopes of the HA, antibody-producing cells will be redirected to conserved epitopes of lower immunogenicity. This “heterologous prime-boost” strategy is not new, but protection against all existing H1 strains has never been established.
We therefore tried to induce antibodies against any H1 strain by sequential vaccination of swine with strains from different H1 lineages in different orders and combinations. We prepared traditional killed vaccines containing a single strain and an adjuvant. Over 100 pigs were vaccinated in the neck and bled every few weeks to measure antibodies against the vaccine strains and a broad set of other strains and subtypes. One month after the last vaccination, they were exposed to a high dose of a live H1N1 strain that did not match with the strains in the vaccines.
We started out with simple two-dose schedules. Pigs were vaccinated first with a strain of the 1A lineage, followed 4 weeks later by a 1B or 1C strain, or vice versa. Control pigs received the same strain for both vaccinations. This traditional method resulted in a narrow anti-HA antibody response against a single vaccine strain. The combinations of two different strains usually induced antibodies against both vaccine strains, which was a plus. Still, the antibodies were unable to neutralize H1 strains that were substantially different from those in the vaccines.
We also looked for antibodies against the second and somewhat more conserved surface protein, the neuraminidase (NA). These antibodies were clearly more cross-reactive, but strains with a very distant N1 could still escape recognition.
“Pan-H1N1” antibodies after three sequential vaccinations with three different H1N1 strains
We then tried to broaden the antibody response by using three doses of three different strains. A principal group of pigs was vaccinated sequentially with 1C, 1B and 1A strains, 4 and 6 weeks apart. Control groups received three doses of 1B, or three doses of “trivalent” vaccine combining three strains in a single shot (Fig. 3).
This time, we also tested blood samples for antibody secreting cells (ASC), the B cells responsible for large-scale production of antibodies. These cells became detectable one week after the second vaccination, to have gone by the time of the third vaccination and reappear one week later. Most remarkable was the very robust expansion of ASC responding to the vaccine strains after the third dose of a different strain. This was followed by an up to 100-fold increase in antibody titers. A third dose of the same vaccine, in contrast, mobilized fewer ASC than the second dose, and had little effect on antibody titers. Finally, the 3-dose 3-strain regimen stood out on all fronts. Post-vaccination sera neutralized 88% of a diverse panel of human and swine H1 strains, and they reacted with any N1 strain, including H5N1 and H7N1 from birds. Protection against challenge infection was as robust as after three doses of trivalent vaccine. The mismatched H1N1 strains could not even infect pigs or cause lung lesions.
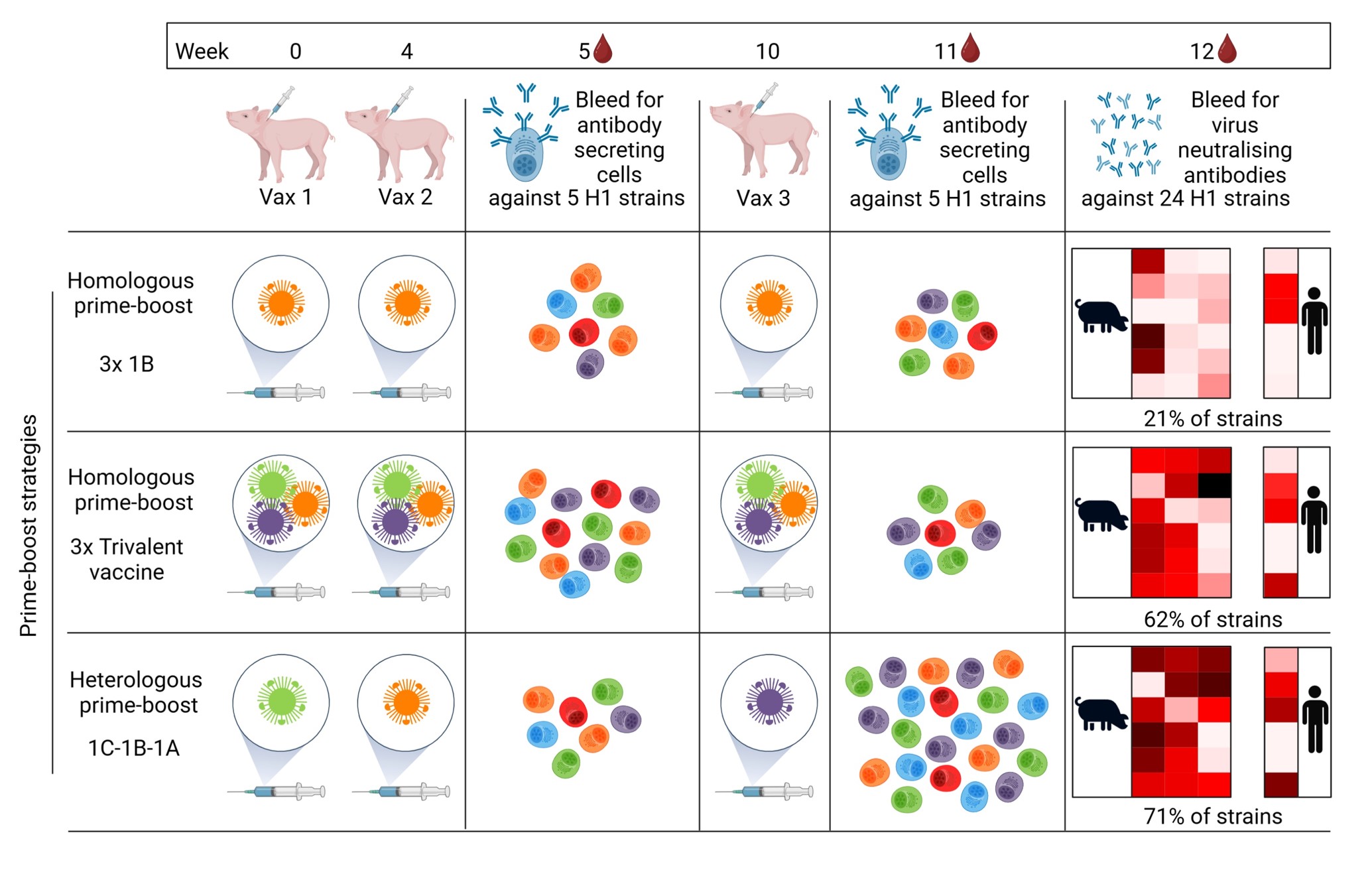
Fig. 3. Schematic of the results of various 3-dose vaccination schedules. The heterologous prime-boost strategy outperforms other regimens in terms of numbers of antibody-secreting cells and breadth of anti-H1 antibodies. Image credit: Ruth Mumo.
These findings suggest that boosting with diverse strains may recruit broadly reactive memory B cells more efficiently than using similar strains. They may also explain the blunted immune response that has been reported in people who got flu shots for several years in a row.
Unraveling the targets on the HA
But why are these antibodies so good at neutralizing diverse H1 strains? To find out, we had to unravel their targets on the HA. Florian Krammer and Peter Palese at the Icahn School of Medicine at Mount Sinai, New York, have built their career around this viral protein, and they were willing to collaborate. In the Krammer lab, Phil Meade tested hundreds of swine sera for antibodies against the HA stalk. The antibodies were there, but the levels remained below those required for cross-protection between different HA subtypes. Not surprisingly, the sera also failed to neutralize non-H1 subtype influenza viruses, such as H3, H5 and H7.
We then focused on the HA head using a series of H1 virus mutants constructed by Sean Liu. These were perfect tools to measure antibodies against each of the five individual antigenic sites. Another mutant allowed to uncover antibodies against the residual, more conserved part of the head. These rare antibodies were only found after three vaccinations with three different strains, which was a key finding. Thus, the broad anti-H1 antibodies likely target conserved HA head epitopes. Apparently, we could bypass the immune system’s tendency to focus on the variable antigenic site epitopes by stepwise exposure to strains that are largely different in these epitopes (Fig. 4). A similar phenomenon seems to occur naturally in a handful of people.
Fig. 4. A model contrasting repeated vaccinations with homologous or heterologous strains. Shown are the antigenic determinants of the HA (left) and the resulting antibodies (right). Orange, green, purple = strain-specific epitopes and antibodies; red = conserved epitopes and broadly reactive antibodies. Traditional, homologous prime-boost vaccination induces strain-specific antibodies. By sequential vaccinations with H1 strains that are substantially different in the most variable epitopes, the immune system can be trained to make antibodies against conserved HA head epitopes (in red), which tend to be overlooked. Image credit: Ruth Mumo.
Still a long way to go
We did not come up with the perfect universal influenza vaccine. Our approach protects only against the H1 subtype and requires at least three injections. However, it combines protection against human drift variants and highly diverse swine flu strains with pandemic potential. The licensed swine flu vaccines contain differing strains, and mixed vaccination schedules are gaining wider acceptance in pig farming. It would be much more difficult to use our approach in humans, and it does not fit into the current influenza vaccination policy. A major complication is pre-existing immunity, which may require different approaches in different age categories. Yet other high-profile viruses – COVID-19, HIV, respiratory syncytial virus, Ebola – can also benefit from heterologous prime-boost vaccination, and personalized vaccines are lurking around the corner. This makes me believe that “What stands in the way becomes the way” (Marcus Aurelius, 121-180).
Postscript
The pig experiments and laboratory analyses took several years and have been a real “tour de force”. My sincere thanks go to everyone involved and many who helped in another way.
References
- Krammer, F. The human antibody response to influenza A virus infection and vaccination. Rev. Immunol. 19, 383-397 (2019).
- Thompson, M. G. & Cowling, B. J. How repeated influenza vaccination effects might apply to COVID-19 vaccines. Lancet Respir. Med. 10, 636-638 (2022).
- Liu, S. T. H., et al. Antigenic sites in influenza H1 hemagglutinin display species-specific immunodominance. Clin. Invest. 128, 4992-4996 (2018).
- Krause, J. C. & Crowe, J. E. Committing the oldest sins in the newest kind of ways-antibodies targeting the influenza virus type A hemagglutinin globular head. Spectr. 2, AID-0021-2014 (2014).
- Thompson, C.P., et al. A naturally protective epitope of limited variability as an influenza vaccine target. Nature Communications 9, 6696 (2018).
- Li, C., et al. Exploring heterologous prime-boost vaccination approaches to enhance influenza control in pigs. Res. 51, 89 (2020).
- Zhang, A., Stacey, H.D., Mullarkey, C.E. & Miller, M.S. Original Antigenic Sin: How first exposure shapes lifelong anti-influenza virus immune responses. Immunol. 202, 335-340 (2019).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in