BA.2 and BA.5 omicron differ immunologically from both BA.1 omicron and pre-omicron variants
Published in Healthcare & Nursing
The emergence of SARS-CoV-2 presented an unprecedented challenge to the general public, policy makers and the scientific community, but at the same time offered the unique opportunity of studying the epidemiology of a virus in a naïve human population. Researchers worldwide rose to the occasion and global cooperation networks were quickly established to share data and knowledge about the development of SARS-CoV-2. One of these initiatives is the SARS-CoV-2 Assessment of Viral Evolution (SAVE) Program1 by the National Institute of Allergy and Infectious Diseases at the National Institutes of Health (NIH), which facilitated the collaboration between the Medical University Innsbruck and the University of Cambridge that resulted in our study of the antigenic relationships among pre-omicron and omicron variants.
Given the rapid evolution of SARS-CoV-2 during it’s nearly three years of circulation within humans, a key question was whether infection/vaccination with one variant would protect against the others or more general how antigenically related the individual variants are to each other. This very question, but with respect to Influenza virus, has been a main research focus in the Smith’s group (Centre for Pathogen Evolution, Cambridge University) for years. Naturally, we thought that antigenic cartography, which proved to represent the antigenic relationships between influenza strains well2, could be an useful tool to understand the relationships of evolving SARS-CoV-2 strains. In brief, antigenic cartography is a method to visualize antibody neutralization titers against multiple variants in different serum cohorts in 2D, in a so-called antigenic map (see figure scheme). More antigenically similar variants will be close to each other on the map, while variants that induce no or only low levels of cross-neutralizing antibodies will be further apart. Crucial for a reliable map, however, is that it is created from single exposure sera, as cross-reactivity after exposure to multiple variants overshadows the underlying antigenic relationships.

At the time of omicron emergence such single exposure sera for BA.1 or BA.2 were very rare as most people had already been vaccinated or infected with a pre-omicron variant. Thus, the Kimpel Lab’s (Institute of Virology, Medical University of Innsbruck) sample collection is unique in having both BA.1 and BA.2 first infection sera from non-vaccinated individuals. To map the antigenic relationship of different omicron variants among each other, the resolution of the map should be improved by analyzing neutralization of omicron variants in both BA.1 and BA.2 first exposure sera. Additionally, neutralization assays were performed with live SARS-CoV-2 isolates, which is advantageous since results obtained by pseudovirus neutralization assays might differ to some extent. In our study, for example, we found good neutralization of BA.2 by double vaccinated sera (equivalent to a single infection), while BA.2 omicron mostly escapes these sera in pseudovirus assays. Given that by now the majority of people have had multiple vaccinations or a combination of vaccination and infection, it is of central importance to compare animal models for single exposure sera with human sera which will allow us to accurately characterize the relationship of future emerging variants.
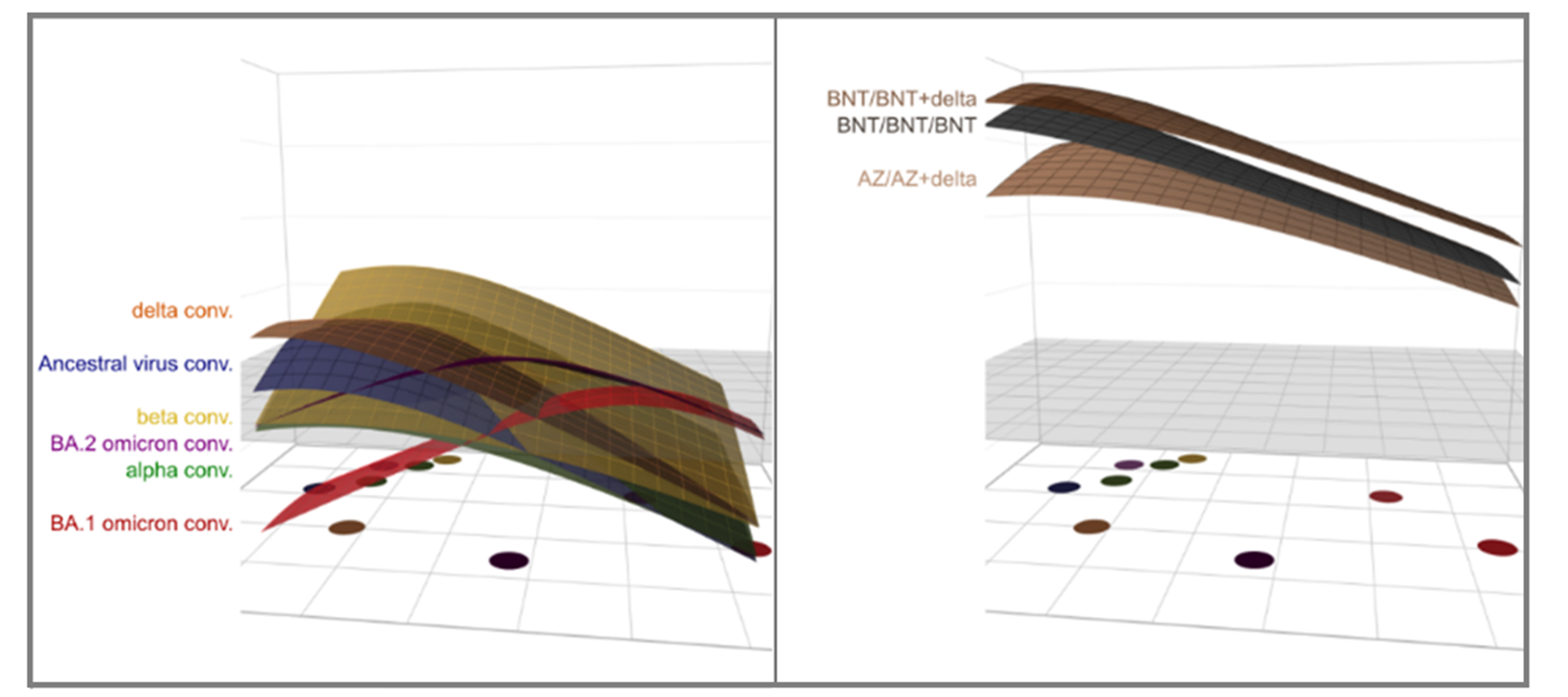
While all three analyzed omicron variants are located distant from pre-omicron variants on the antigenic map, our data revealed that they also differ from each other. We also found that hybrid immunity (a combination of vaccination and infection) conferred high antibody levels and hence good protection against all variants using the method of antibody landscapes. While we can make inferences about immunity after single exposure in an antigenic map, antibody landscapes allow us to look at cross-reactive immunity after multiple exposures. Antibody landscapes are an illustration of an individual’s antibody neutralization profile against virus variants on an antigenic map, where the neutralization titers are plotted in a third dimension above the map and a surface is fitted to these titers to create a continuous immune landscape. The broadening of the neutralization profile was particularly evident in the combination of pre-omicron exposure (vaccination or infection) and subsequent infection with an omicron strain. From this, we conjectured that an update of the vaccine to an omicron strain would be beneficial in inducing protective antibody levels against all analyzed variants, and better against future emerging variants given the increased breadth compared to three exposures to the same strain.
Our finding that the analyzed omicron variants are distinct from each other is highly relevant for public health and future vaccine updates. For vaccine strain updates, it is important to consider how similar the vaccine strain is to currently and likely future circulating variants. At the time of our analysis, BA.5 had replaced BA.1 and BA.2 in many countries. However, meanwhile the virus has diversified substantially and a number of BA.2 and BA.5 variants with additional mutations associated to immune escape co-circulate. Therefore these and future variants might be distant to original BA.1, BA.2 and BA.5 omicron variants on the antigenic map and a vaccine update to BA.5 will likely not confer 100% protection against these future variants.
The better performance of variant vaccines compared to the standard vaccine has now been confirmed in clinical trials.3,4 At the time of data collection and analysis for our study, these trials were still in the planning phase. It is promising to see that results from small, regionally restricted datasets align with these big trials, and highlights the potential of studies such as ours in the guidance of clinical trials and vaccine design.
References
1 DeGrace, M. M. et al. Defining the risk of SARS-CoV-2 variants on immune protection. Nature 605, 640-652 (2022). https://doi.org:10.1038/s41586-022-04690-5
2 Smith, D. J. et al. Mapping the antigenic and genetic evolution of influenza virus. Science 305, 371-376 (2004). https://doi.org:10.1126/science.1097211
3 Pfizer & BioNTech. Pfizer and BioNTech Announce Updated Clinical Data for Omicron BA.4/BA.5-Adapted Bivalent Booster Demonstrating Substantially Higher Immune Response in Adults Compared to the Original COVID-19 Vaccine <https://www.pfizer.com/news/press-release/press-release-detail/pfizer-and-biontech-announce-updated-clinical-data-omicron> (04.11.2022).
4 moderna. Moderna's BA.4/BA.5 Targeting Bivalent Booster, mRNA-1273.222, Meets Primary Endpoint of Superiority Against Omicron Variants Compared to Booster Dose of mRNA-1273 in Phase 2/3 Clinical Trial, <https://investors.modernatx.com/news/news-details/2022/Modernas-BA.4BA.5-Targeting-Bivalent-Booster-mRNA-1273.222-Meets-Primary-Endpoint-of-Superiority-Against-Omicron-Variants-Compared-to-Booster-Dose-of-mRNA-1273-in-Phase-23-Clinical-Trial/default.aspx> (14.11.2022).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in