Behind FISHnCHIPs
Published in Protocols & Methods
The idea: clarity in numbers
Imagine you are trying to identify someone in a crowded city without being able to see them up close. One approach would be to look for their distinguishing features, like haircolor, hairstyle, or accessories they are wearing. However, from a distance, these individual features might not be clear enough. One way to overcome this challenge is to look for combinations of multiple features. Even if you can't make out an individual feature, like the hair color, from a distance, the combination of features (hair color plus clothing, plus height, and etc.) can form a distinct pattern that would allow you to more confidently recognize that person.
In the world of biology, cells are the fundamental units of life, and their complex organization in a tissue is truly remarkable. From the specialized neurons that transmit signals in our brains to the immune cells that protect us from diseases, each cell type, in its unique position, plays a crucial role in the intricate symphony of life. The traditional goal of spatial transcriptomics, which is to identify individual cell types within a tissue, can be as challenging as picking out specific individuals in a crowded city from a distance. Just as people might be difficult to identify when viewed from afar, the molecular marker for specific cell types can become too miniscule for us to notice the difference. Furthermore, molecular crowding can also make it difficult to discern the overlapping signals in traditional gene marker based spatial transcriptomics methods.
FISHnCHIPs elegantly overcomes these challenges by fusing multiple co-expressed genes to form powerful yet coherent multidimensional signals. The key idea behind FISHnCHIPs is the recognition that genes involved in the same biological programs or pathways are also spatially co-localized within the same cells. Therefore, instead of measuring individual marker genes, FISHnCHIPs targets gene programs, the fusion of many marker genes. This allows FISHnCHIPs to rely on the combined unique molecular fingerprints to detect and distinguish the different cell types within a densely packed tissue.
The process: How does FISHnCHIPs work
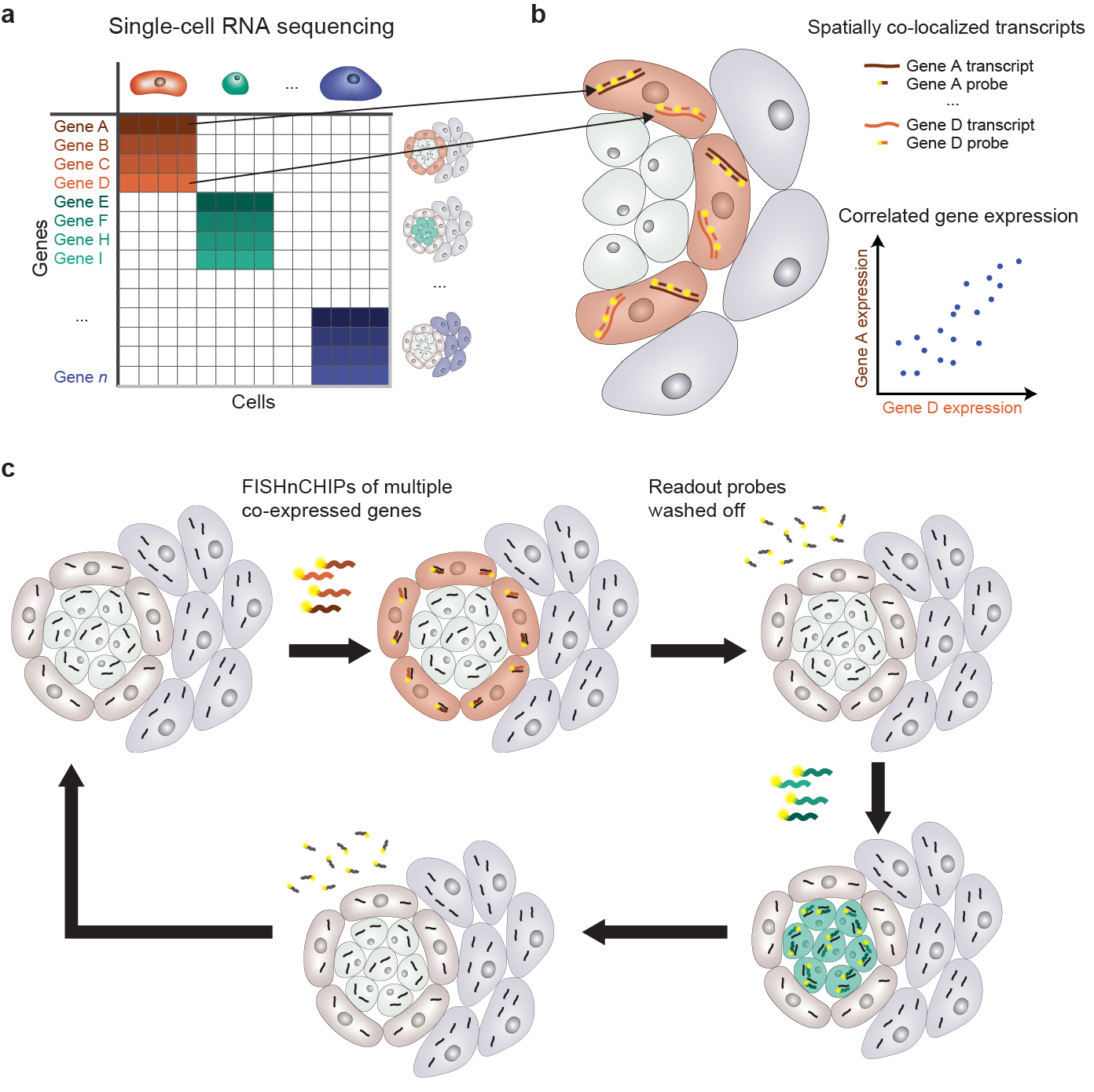
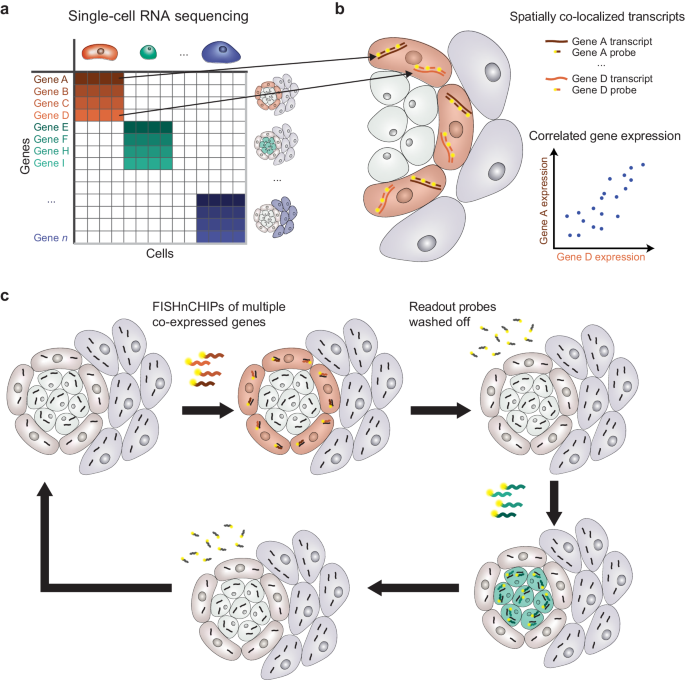
FISHnCHIPs introduces a strategic approach in designing the gene panel. The process begins by analyzing single-cell RNA sequencing (scRNA-seq) data to identify groups of highly correlated genes, known as gene modules or expression programs (Figure 1a). These gene modules act as molecular signatures for specific cell types or cellular states, which also provides a comprehensive view of gene expression pattern across different cell types. Those co-expressed genes are not only correlated in their expression level, but also spatially colocalized within the tissue (Figure 1b). This colocalization allows us to design a pool of fluorescently labeled oligonucleotide probes to target the transcripts of these co-expressed genes in repeated rounds of hybridization and washing (Figure 1c). Importantly, because multiple co-expressed genes are targeted simultaneously, the resulting signal is amplified and can be detected under a microscope with low magnification.
We first performed a validation experiment using mouse kidney tissue. Five kidney cell types – renal macrophages, endothelial cells, loop of Henle cells, collecting duct cells, and the podocytes – were selected for comparison with single gene FISH using the top differentially expressed (DE) gene. For FISHnCHIPs, each cell type was labelled with 14-23 genes. A striking co-localization and increased signal was observed. Moreover, we quantified the cell signals and found a remarkable 6 to 39-fold increase in fluorescence intensity per cell, closely matching our computational predictions. The improved signal made it easier to capture intricate details of the kidney tissue architecture, such as the arrangement of podocytes wrapping around the glomerular endothelial cells in the highly fenestrated Bowman’s capsule.
The cellular landscape: power of FISHnCHIPs in action
Building on the initial success with the mouse kidney, we expanded FISHnCHIPs to profile more modules, in more complex tissues, and in a shorter time.
Mouse cortex layer-specific structure
In the mouse cortex, we accurately recapitulated the expected frequency and spatial distribution of major brain cell types, including glutamatergic neurons, GABAergic neurons, astrocytes, oligodendrocytes, endothelial cells, microglia, and more. The technique can also be used to unveil the layer-specific localization of neuronal subtypes stratified across the canonical laminar structure of the cortex.
Whole brain, rapid profiling
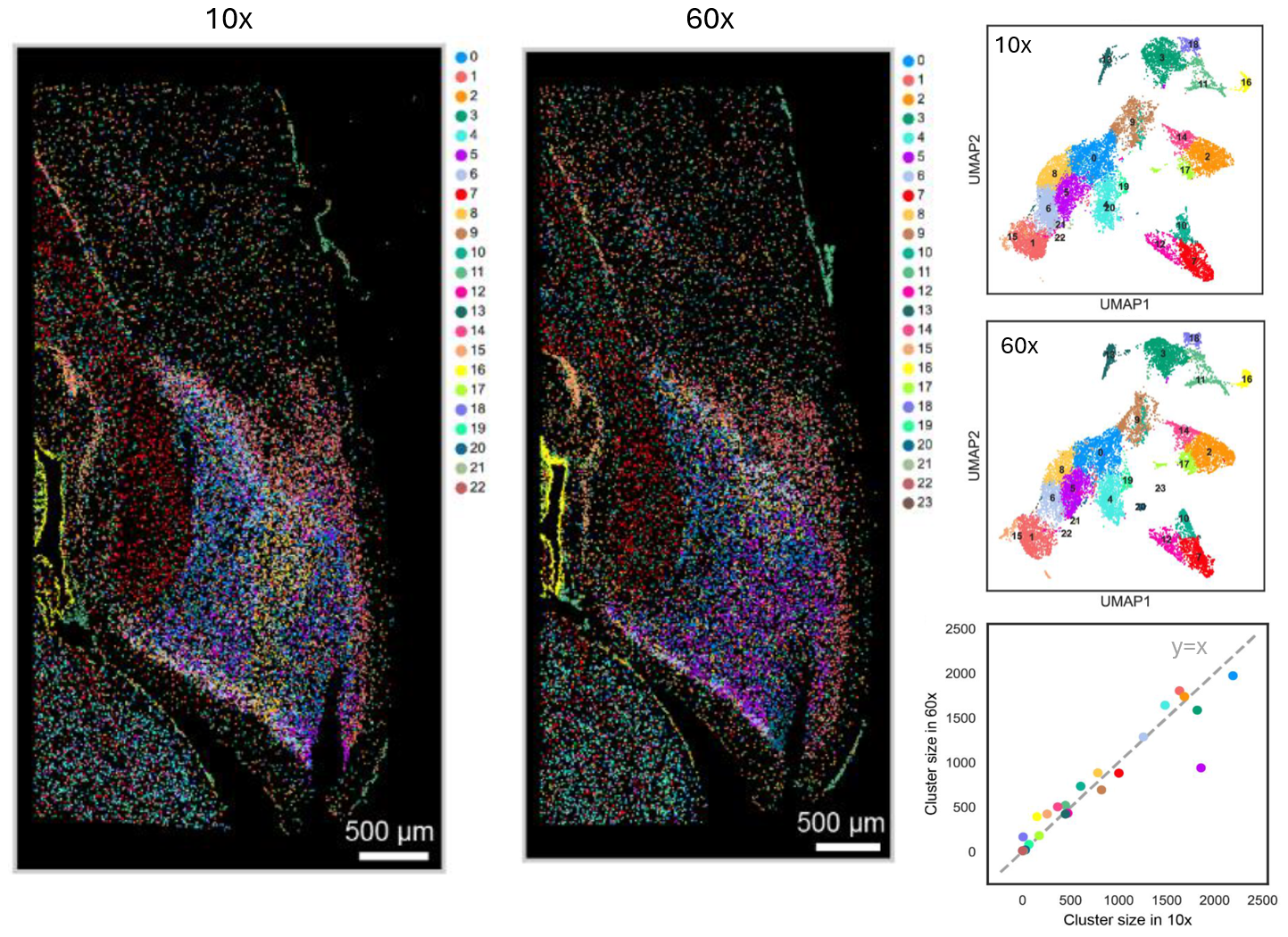
Encouraged by these results, we harnessed the high-throughput capabilities of FISHnCHIPs to profile 53 gene modules across a whole sagittal section of the mouse brain. The profiling for this large field of view using a 10x objective lens was completed in only 21 hours – a task that would typically take almost a week for ~700 genes under a 60x objective. Remarkably, despite the lower magnification, FISHnCHIPs enabled reliable detection and clustering of cells, where the cluster sizes of 10x and 60x imaging in adjacent sections were highly correlated (Figure 2). It suggested that there was no observable degradation of data quality despite the lower magnification and higher throughput.
Robust performance in clinical samples
Extending beyond basic research, we also demonstrated FISHnCHIPs robustness by profiling a frozen clinical sample, which often suffers from compromised RNA quality, making them a particularly difficult tissue type to profile. In this experiment, we successfully used FISHnCHIPs to unveil the distinct spatial distribution of two cancer-associated fibroblast (CAF) subtypes. One of these subtypes was previously found to be highly relevant in establishing an immune suppressive tumor microenvironment.
Through these examples, we showed that FISHnCHIPs is remarkably sensitive, scalable, and robust, and can be harnessed for molecular analyses of clinical tissue samples. Just as a sophisticated facial recognition system incorporates multiple identifying features to pinpoint an individual, FISHnCHIPs can precisely elucidate the spatial origin of rare cell types in complex diseases. Looking ahead, the potential applications of FISHnCHIPs are vast and exciting. Given the wealth of transcriptomic data generated by many large-scale single-cell RNA sequencing efforts worldwide, we hope to see more researchers and clinicians designing FISHnCHIPs assay to gain unprecedented insights into the cellular basis of human health and diseases.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in