Behind the Paper: "Phase-Separating Peptides for Direct Cytosolic Delivery and Redox-Activated Release of Macromolecular Therapeutics"
Published in Chemistry

Our paper “Phase-Separating Peptides for Direct Cytosolic Delivery and Redox-Activated Release of Macromolecular Therapeutics” is an illuminating example of how a fundamental, curiosity-driven exploration unexpectedly evolved into a translational research project that has now promising therapeutic applications, which we never imagined when this research started.
The genesis of this study can be traced back to 2004 when I started my post-doc at UC Santa Barbara. Having been trained in traditional Materials Science (composite and mechanics of materials), I sought to expand my research skills towards the Life Sciences. The lab of Herb Waite, who explores fundamental links between the biochemistry of biological materials (such as mussel adhesive fibers) and their biophysical and mechanical properties, ticked all the boxes of acquiring interdisciplinary knowledge. As soon as I arrived in the lab, Herb handed me a parrot-like beak of a Jumbo squid (Fig. 1) and suggested to analyze its biochemical composition and measure its mechanical properties. The squid beak turned out to be an intriguing material. Hard tissues with biting and chewing functionalities (such as human teeth) are normally calcified to impart them with hardness and strength [1]. However, the squid beak proved to be non-mineralized and fully organic [2], made of chitin and highly cross-linked proteins arranged in a graded fashion [3]: soft and flexible at its proximal part embedded within the muscular buccal mass, and very hard at its distal region called the rostrum that is used to crush and chew on preys. Interestingly, we found that the flexible region mostly contained hydrated chitin whereas the hard rostrum was protein-dominated, which is non-intuitive since chitin is usually viewed as a load-bearing biopolymer and would thus be expected to be more abundant at the hard rostrum.

Because of the highly cross-linked nature of squid beaks, isolating intact proteins for subsequent sequencing proved exceedingly challenging. The cross-link chemistry was identified [4] and evidence that squid beak proteins were bound to polysaccharides could be established [5] but obtaining any sequence information remained an intractable task. A breakthrough came a few years later. Upon starting an independent Academic position at Nanyang Technological University (NTU) in Singapore, my lab initiated a collaboration with Dr. Shawn Hoon, an expert in next-generation RNA sequencing (RNA-seq). Together with a previous Senior Research Fellow in my lab, Dr. Paul Guerette, we quickly recognized the amazing potential of RNA-seq. to sequence structural proteins from extra-cellular biological materials. Traditionally, the highly insoluble nature of such strong protein-based materials made sequencing of their protein building blocks a tedious and lengthy endeavor that took years using classical proteomic and molecular biology methods. And for large MW proteins with highly repetitive motifs, such as spider silks [6] or mussel adhesives [7], only partial sequences were usually obtained, and complete protein sequences remained inaccessible. By probing the transcriptome assembled by RNA seq. of secreting glands with traditional and high-throughput proteomic technologies [8] (such as tandem mass spectroscopy, tandem MS), dozens of structural proteins comprising mussel adhesive proteins, hard proteinaceous teeth from squids’ arms and tentacles [9], and egg case capsules of oviparous animals could be sequenced [10] within just a few months.
This approach proved to be a game-changer for the squid beak. A transcriptome library of the “beccublast cells” that secretes the beak proteins was generated. In parallel, even though only a few peptide fragments could be chemically fragmented and de novo sequenced by tandem MS, this was sufficient to probe the transcriptome and successfully obtain the complete sequences of proteins making up the squid beak [11]. Two main family of proteins were identified, namely proteins containing chitin-binding domains as well as proteins enriched in glycine (G), alanine (A), and histidine (H) that were called “histidine reach beak proteins” (HBPs). A noticeable feature in HBPs’ primary sequences was the presence of multiple copies of the pentapeptide repeat GHGXY, where X is usually proline (P), valine (V), or leucine (L). This sequence design was reminiscent of tropoelastin, which is well-known to undergo self-coacervation [12], a process during which a homogenous protein solution phase-separates into protein-rich droplets and a protein-depleted phase when subjected to external stimuli (typically temperature of pH change). Coacervation was demonstrated for both full-length HBPs as well as shorter peptides designed from the pentapeptide repeats. However, a subtle difference between HBPs’ repeats and those from tropoelastin is the high histidine content in the former, making the coacervation pH-dependent.
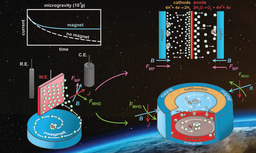
Since coacervation had previously been demonstrated to be able to readily recruit biomacromolecules in aqueous conditions [13] to form microdroplets loaded with therapeutic cargos, we sought to exploit this property and hypothesized that polypeptides derived from HBPs may also be able to recruit macromolecular cargos. Thus, a first proof-of-principle study consisted in designing a short peptide made of a few GHGXY repeats that could simultaneously recruit insulin and glucose oxidase (GOx) [14]. Upon exposure of the cargo-loaded coacervates to glucose, the latter reacted with GOx to form gluconic acid, lowering the pH in the microenvironment of the coacervates, leading to their disassembly and eventually allowing glucose-responsive release of insulin. In a subsequent study [15], the low MW anti-cancer compound Doxorubicin (Dox) and iron oxide magnetic nanoparticles were recruited in the peptide coacervates. By applying an alternate magnetic field, localized heating by the magnetic hyperthermia effect was achieved within the microdroplets, leading to heat-induced disassembly and release of Dox.
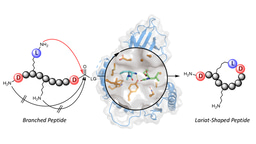
At this stage, a serendipitous finding prompted the current article. Despite their size in the 1-2 µm range, we realized that coacervates were uptaken by cells in a manner that did not follow classical endocytic pathways. In fact, once inside the cells, the team of our collaborator, Dr. Farid Ghadessy from Singapore’s Agency for Science Technology and Research (A*Star), realized that the coacervates remained stable for at least a week, without being degraded as would be expected if they were within endosomic compartments. In other words, our peptide coacervates can be viewed as membraneless organelles formed outside cells but that can cross the cell membrane and remains stable in the cytoplasm. This behavior is both a blessing and a curse: a blessing because the stability would preclude premature degradation of cargos to occur. But it is also a curse, because a therapeutic cargo that remained trapped within an membraneless organelle is unhelpful for its ultimate task of reaching its intracellular target. It thus appeared evident that the peptide coacervates had to be modified such that they could disassemble once in the cytoplasm. A new PhD student in the lab, Sun Yue, decided to tackle this challenge and came up with a clever chemical design. First, he introduced one basic residue lysine to the peptide, shifting its phase separation pH above pH 9 instead of 6.5 for the original peptide. Then, he grafted to the lysine a self-immolative and hydrophobic moiety containing a disulfide bond. Since the positive charge of lysine was neutralized, the peptide phase-separated again at pH 6.5. But once inside the cell, the endogenous antioxidant glutathione reduced the disulfide bond. This activated the self-immolative chemistry, such that the attached moiety was cleaved off from the peptide backbone through a series of auto-catalytic reactions. As a result, the peptide disassembled in the cytoplasm, in turn releasing its cargo.
The design worked beyond our expectations. As long as the cargo was recruited within the coacervate microdroplets (see Fig. 2 for GFP-loaded coacervates), we found that they could cross the cell membrane and release their payload directly in the cytoplasm (Fig. 2). This included proteins with a MW ranging from 14 kDa (lysozyme) up to the 470 kDa enzyme beta-galactosidase as well as proteins with both acidic and basic isoelectric points. The delivery of anti-cancer stapled peptides from our collaborator Fahrid Ghadessy, which allow the activation of the tumor suppressor protein p53 but have limited cellular permeabilities, could also be mediated by the peptide coacervates. Finally, intrigued by our delivery system, Dr. Anthony Partridge from MSD International suggested to try delivering mRNAs, given some of the drawbacks associated with lipid nanoparticles delivery (it should be highlighted that these experiments were conducted before the COVID-19 pandemic brought mRNA therapeutics to the central stage). Here again, a high transfection efficiency was achieved.
.jpg)
There remain many unanswered questions from this study. First and foremost, how do the coacervates cross the cell membrane? We know it does not happen through classical endocytic pathways, and we also know it is cholesterol-dependent. But beyond this, it is still a bit of a mystery. Towards therapeutic applications, in vivo stability and biodistribution will need to be established and such studies are currently underway in our lab. But the key message I would like to convey here is that curiosity-driven explorations must be pursued and funded, even when an obvious translational application is not immediately apparent. When I was first handed this strange-looking squid beak by my post-doc supervisor back in 2004, I would have never imagined that initial research into its nanomechanical properties would one day lead to a new intracellular drug delivery platform. As a matter of fact, with my then background in metallurgy and fracture mechanics, I did not even know what drug delivery consisted of !
References
- Wegst, U. & Ashby, M. F. The Mechanical Efficiency of Natural Materials. Philosophical Magazine 84, 2167-2181, (2004).
- Miserez, A., Li, Y., Waite, J. H. & Zok, F. Jumbo Squid Beaks: Inspiration for the Design of Robust Organic Composites. Acta Biomateriala 3, 139-149, (2007).
- Miserez, A., Schneberk, T., Sun, C., Zok, F. W. & Waite, J. H. The Transition from Stiff to Compliant Materials in Squid Beak. Science 319, 1816-1819, (2008).
- Miserez, A., Rubin, D. & Waite, J. H. Cross-linking Chemistry of Squid Beak. Journal of Biological Chemistry 285, 38115-38124, (2010).
- Rubin, D., Miserez, A. & Waite, J. H. in Advances in Insect Physiology Vol. 38 (ed J. Casas) 75-133 (Elsevier, 2010).
- Xu, M. & Lewis, R. V. Structure of a Protein Superfiber: Spider Dragline Silk. Proceedings of the National Academy of Sciences of the United States of America 87, 7120-7124, (1990).
- Waite, J. H., Housley, T. J. & Tanzer, M. L. Peptide Repeats in a Mussel Glue Protein - Theme and Variations. Biochemistry 24, 5010-5014, (1985).
- Guerette, P. A. et al. Accelerating the Design of Biomimetic Materials by Integrating RNA-Seq. with Proteomics and Materials Science. Nature Biotechnology 31, 908–915, (2013).
- Guerette, P. A. et al. Nanoconfined β-sheets Mechanically Reinforce the Supra-Biomolecular Network of Robust Squid Sucker Ring Teeth. ACS Nano 8, 7170–7179, (2014).
- Guerette, P. A. et al. Integrative and Comparative Analysis of Coiled-Coil Based Marine Snail Egg Cases - A Model for Biomimetic Elastomers. Biomaterials Science 2, 710-722, (2014).
- Tan, Y. P. et al. Infiltration of a Chitin Scaffold by Protein Coacervates Defines the Squid Beak Mechanical Gradient. Nature Chemical Biology 11, 488–495, (2015).
- Yeo, G. C., Keeley, F. W. & Weiss, A. S. Coacervation of Tropoelastin. Advances in Colloid and Interface Science 167, 94-103, (2011).
- Johnson, N. R. & Wang, Y. Coacervate delivery systems for proteins and small molecule drugs. Expert Opinion on Drug Delivery 11, 1829-1832, (2014).
- Lim, Z. W., Ping, Y. & Miserez, A. Glucose-Responsive Peptide Coacervates with High Encapsulation Efficiency for Controlled Release of Insulin. Bioconjugate Chemistry 29, 2176-2180, (2018).
- Lim, Z. W., Varma, V., Ramanujan, R. V. & Miserez, A. Magnetically Responsive Peptide Coacervates for Dual Hyperthermia and Chemotherapy Treatments of Liver Cancer. Acta Biomaterialia 110, 221-230, (2020).
Follow the Topic
-
Nature Chemistry

A monthly journal dedicated to publishing high-quality papers that describe the most significant and cutting-edge research in all areas of chemistry, reflecting the traditional core subjects of analytical, inorganic, organic and physical chemistry.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in