Claim your spot on particles with secondary metabolites
Published in Microbiology
Oceans cover 70% of Earth’s surface and produce 50-80% of the oxygen introduced to the atmosphere. Loss of oxygen from the global ocean has been documented over the last few decades, and is predicted to accelerate into the future as a consequence of global warming and coastal eutrophication. Deoxygenated regions of the world oceans, including oxygen-deficient water columns (ODWCs) are expected to expand geographically and intensify. ODWCs are usually areas with highly productive surface waters that are important for fisheries, and they host active prokaryotic communities (bacteria and archaea) that dominate carbon, nitrogen and sulfur cycling. Because of their high productivity, ODWCs are replete with sinking particles that provide colonizable, nutrient-rich substrates for microbes.
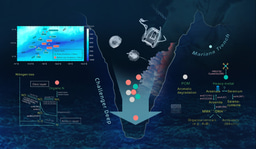
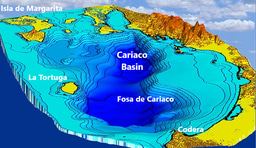
The Cariaco Basin, lying off the northern coast of Venezuela, is an end-member of global ODWCs with a pronounced vertical redox gradient (redoxcline) that reflects oxic, suboxic (O2 < 1 µM) and anoxic waters, including a euxinic (anoxic and sulfidic) core. The relatively stable redoxcline makes Cariaco Basin an ideal natural laboratory for studying how microbes organize and function under different O2 regimes that may mimic those unfolding as our oceans deoxygenate.
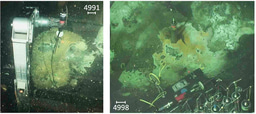
We addressed a critical knowledge gap regarding whether free-living (FL) and particle-associated (PA) prokaryotic taxon composition along oxygen and sulfide gradients in the Cariaco Basin is shaped by strategic secondary metabolite production. To experimentally segregate free-living and particle-associated prokaryote taxa, water samples were size fractionated using 2.7 and 0.2 µm filters (FL: 0.2-2.7 µm vs. PA: > 2.7 μm). Water samples were collected in three discrete water masses: within the oxycline (O2 declining from 110 μΜ to ~1μM), the anoxic layer (undetectable O2 and H2S) and the euxinic (no O2, and 80μΜ H2S) bottom layer. We reconstructed 565 metagenome-assembled genomes (MAGs) from FL and PA fraction DNA extractions. Putative biosynthetic gene clusters (BGCs) in these MAGS involved in secondary metabolite production were identified using antiSMASH. Expression of these BGCs in PA and FL fractions was addressed through metatranscriptomic analyses.
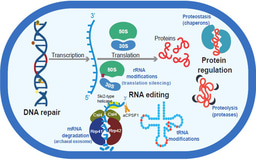
Secondary metabolites are not essential for growth but play essential roles in numerous ecological interactions, such as nutrient acquisition, microbial defense (e.g., antibiotics), and competitive exclusion. Our data revealed recognizable BGCs encoding for antimicrobial compounds (non-ribosomal peptide synthetases: NRPS, polyketides, post-translationally modified peptides: RiPPs). Furthermore, BGCs encoding for compounds that potentially have broader impacts on microbial survival in ODWCs, such as pigments and toxins (e.g., aryl polyenes, terpenes) or those facilitating biofilm formation (e.g., RiPPs and phenazines) were common in our MAGs.
Overall, our data provide evidence for secondary metabolite synthesis by a broad diversity of bacterial and archaeal phyla inhabiting the Cariaco Basin. We find that BGCs involved in antibiotic synthesis (e.g., NRPSs, polyketides, RiPPs, and terpenes) recovered from particle-associated samples were more highly expressed than in the free-living samples. We postulate that production of a suite of secondary metabolites by PA-associated taxa reflects a competitive strategy for nutrients, space and oxygen acquisition on particles. Particles are known to provide nutrients and protection for oxygen- or sulfide-sensitive microbiota. The BGCs identified in our study indicate a complex network of ecological interactions related to a competitive, yet communicative lifestyle within PA microbes, and to a lesser extent within FL communities. These findings pave the way for future laboratory characterizations of genes for novel bioactive metabolites with potential ecological and pharmaceutical importance.
For details, please see our manuscript https://www.nature.com/articles/s41467-023-36026-w.
*Geller-McGrath, D., *Mara, P., Taylor, G.T., Elizabeth Suter, Edgcomb V., Pachiadaki, M. Diverse secondary metabolites are expressed in particle-associated and free-living microorganisms of the permanently anoxic Cariaco Basin. Nat Commun 14, 656 (2023). https://doi.org/10.1038/s41467-023-36026-w
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in