Combined whole-mount fluorescent in situ hybridization and antibody staining in zebrafish embryos and larvae
Published in Protocols & Methods
Written by Jianbo He, Dashuang Mo, Jingying Chen & Lingfei Luo
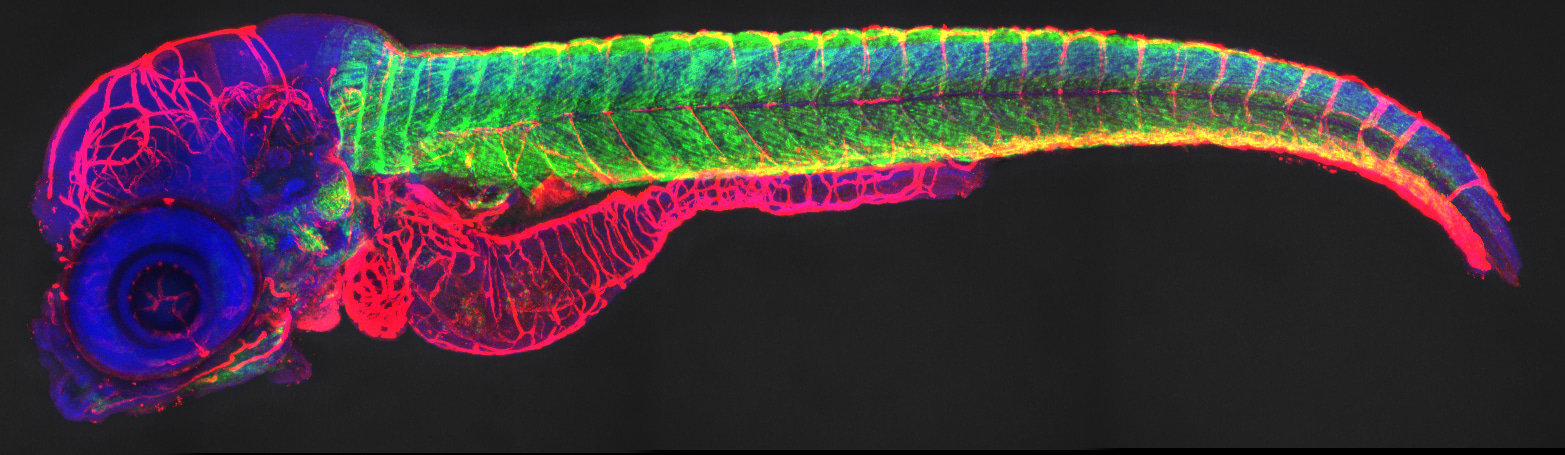
Zebrafish are widely used as a vertebrate model to study embryonic development and tissue regeneration. The spatiotemporal expression of specific genes can be detected through RNA fluorescent in situ hybridization (FISH). Whereas ISH probes penetrate well into zebrafish embryos before three days post-fertilization (dpf), the permeability of late-stage embryos (three dpf and four dpf) and larvae dramatically decreases. In contrast, whole-mount antibody staining/immunofluorescence (IF) is used to detect the localization of antigens using fluorescent labelled primary or secondary antibodies. In comparison to ISH, IF results in higher optical resolution and can, therefore, be used to detect precise protein localization even at the subcellular level, but the reliable antibodies that work in zebrafish are limited.
FISH and antibody staining/immunofluorescence (IF) are widely used to detect distributions of mRNAs and proteins. When there is no antibody available, therefore, it is necessary to combine FISH and IF to study expression patterns of several genes. Combination of FISH and IF has been successfully applied in Drosophila. However, in this approach, IF is performed before FISH and proteinase K treatment is used to enhance probe penetration. It is at the expense of antigen integrity, and this strategy has proven to be unsuccessful for zebrafish larvae. Our approach is based on robust protocols previously described to detect mRNA in early zebrafish embryos. However, at larval stages, the skin and the extracellular matrix components are dense and thick so that penetration of RNA probes limited. To overcome this difficulty, we describe procedures for removal of the skin with tweezers and use washing with PBS + Triton X-100 instead of proteinase K treatment to increase the permeability and preserve the antigens. We want to get more accurate gene expression patterns at single-cell resolution and hope to improve the sensitivity and efficiency of genetic testing.
To prevent RNA degradation and fluorescence quenching, we performed fluorescence in situ hybridization before antibody staining. Instead of proteinase K, zebrafish skin is removed with tweezers in the larvae stage (Fig. 1a), and the larvae are washed with PBS+tritonx-100 to increase tissue permeability and preserve embryonic and larval epitopes. With these optimizations, we get precise results, just as we are interested in the insulin gene, so we use this optimization methold to get the exact location of the gene (Fig. 1b).
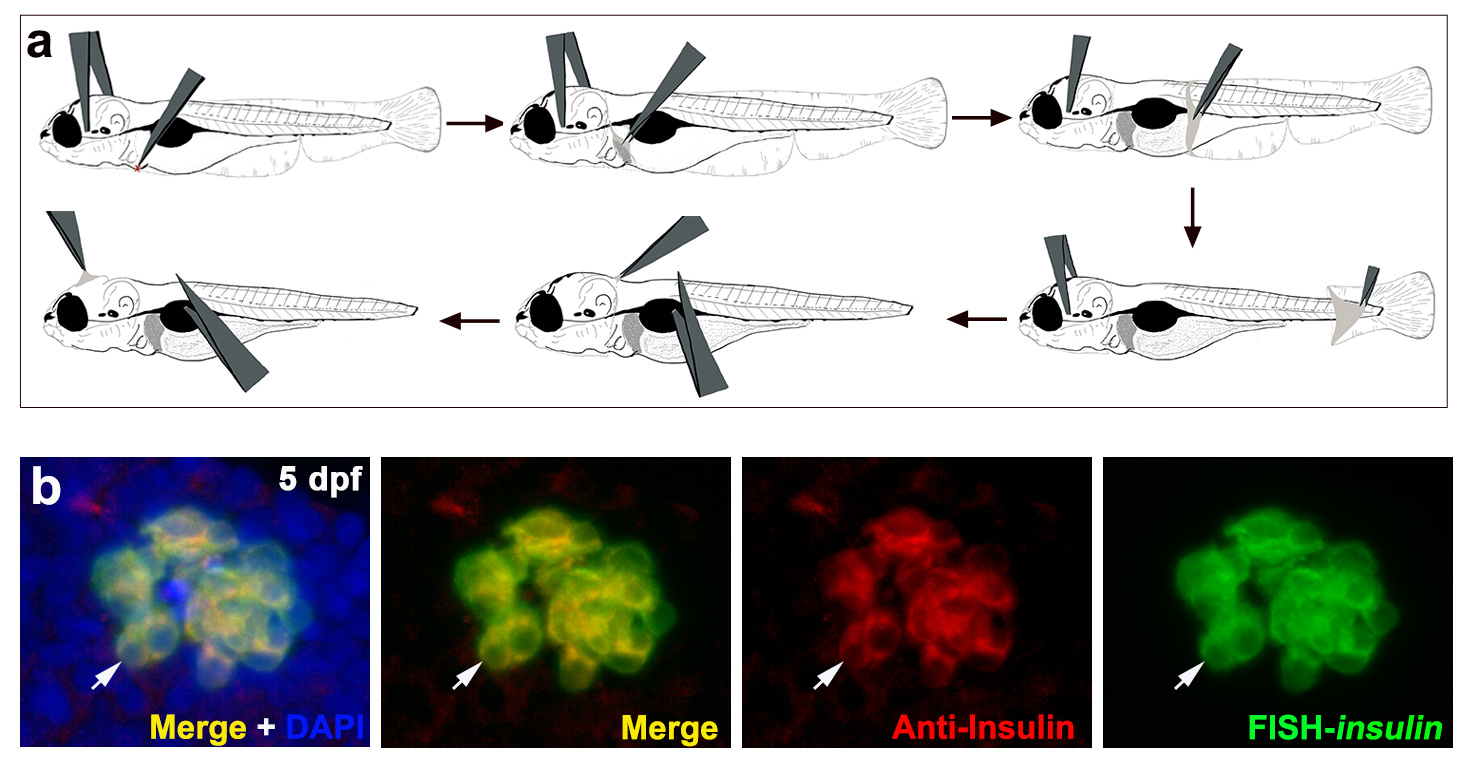
Figure 1. The schematic of skin removed and the application of this protocol to pancreatic beta cells in zebrafish larvae. a, the schematic of the skin removed. b, the application of the FISH-IF to detect the expression patterns of insulin at transcriptional and translational levels at larval stage of zebrafish.
This protocol can be used to visualize gene expression patterns in whole-mount in different tissues at different embryonic and larval stages up to 20 dpf. The high-resolution expression patterns of genes in tissues and even the location of target genes at the single-cell level can be observed. Therefore, this protocol can be applied in many fields of life sciences.
Please link to full text article: https://doi.org/10.1038/s41596-020-0376-7.
Follow the Topic
-
Nature Protocols
This journal publishes secondary research articles and covers new techniques and technologies, as well as established methods, used in all fields of the biological, chemical and clinical sciences.






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in