Controlling plant pathogens with CRISPR.
Published in Bioengineering & Biotechnology
Plant diseases are a major constraint on global food security across the world. The disease-causing agents come in all shapes and sizes, and from across the kingdoms of life. It is generally accepted that plants which are inherently (i.e. genetically) resistant to a disease are an optimal solution in agriculture from practical, financial, and sustainability/equity-oriented points of view because no additional work/cost/inputs are required to prevent disease and thereby yield loss.
How you get to a resistant plant is important. There are, in essence, two ways to do this: the first, and most intuitive, is to add a gene to the plant that has some negative impact on the pathogen (often so called “resistance genes” or “R genes”); the second, and somewhat unintuitive, is to remove a gene from the plant that the pathogen was relying on manipulating in some way in order to cause disease (so called “susceptibility genes” or “S genes”).
The evolutionary biologist might favour the S gene because of a fundamental rule of nature: breaking things is easy - making things is hard. Intuitively, we all know this to be true without the proof. It is easier to break your phone, than it is to make yourself a new phone. The same is true with the interactions between proteins: mutations that change the shape of a protein are much more likely to break its function than they are to make a new function. This is important because many R genes added to plants work by recognising pathogen proteins in order to mount an immune response. Breaking this recognition is “easy”, and so pathogens readily evolve to avoid this type of resistance. On the flip side, resistance derived by removing an S gene from the plant, that the pathogen was manipulating and relying on, requires the pathogen to regain this ability in some way (i.e. making something new), and making things is “hard”. As a result, it is generally harder for pathogen evolution to overcome a resistant plant derived from the loss of an S gene, than it is for pathogen evolution to overcome a resistant plant derived from the addition of an R gene.
The biotechnologist and the politician might agree with the evolutionary biologist because of CRISPR. CRIPSR/Cas genome editing has revolutionised many fields of study. CRISPR/Cas is extremely good at making targeted deletions in crops genomes, and its approval for commercial use in many geographies is streamlined compared to other biotechnological approaches to crop improvement (e.g. transgenesis/genetic modification). There is a clear opportunity to use CRISPR/Cas to develop disease resistant crops, provided the resistant plant is derived by making a targeted deletion (i.e. removing a gene). It is for this reason, that the biotechnologist, the politician, and the evolutionary biologist might favour the S gene.
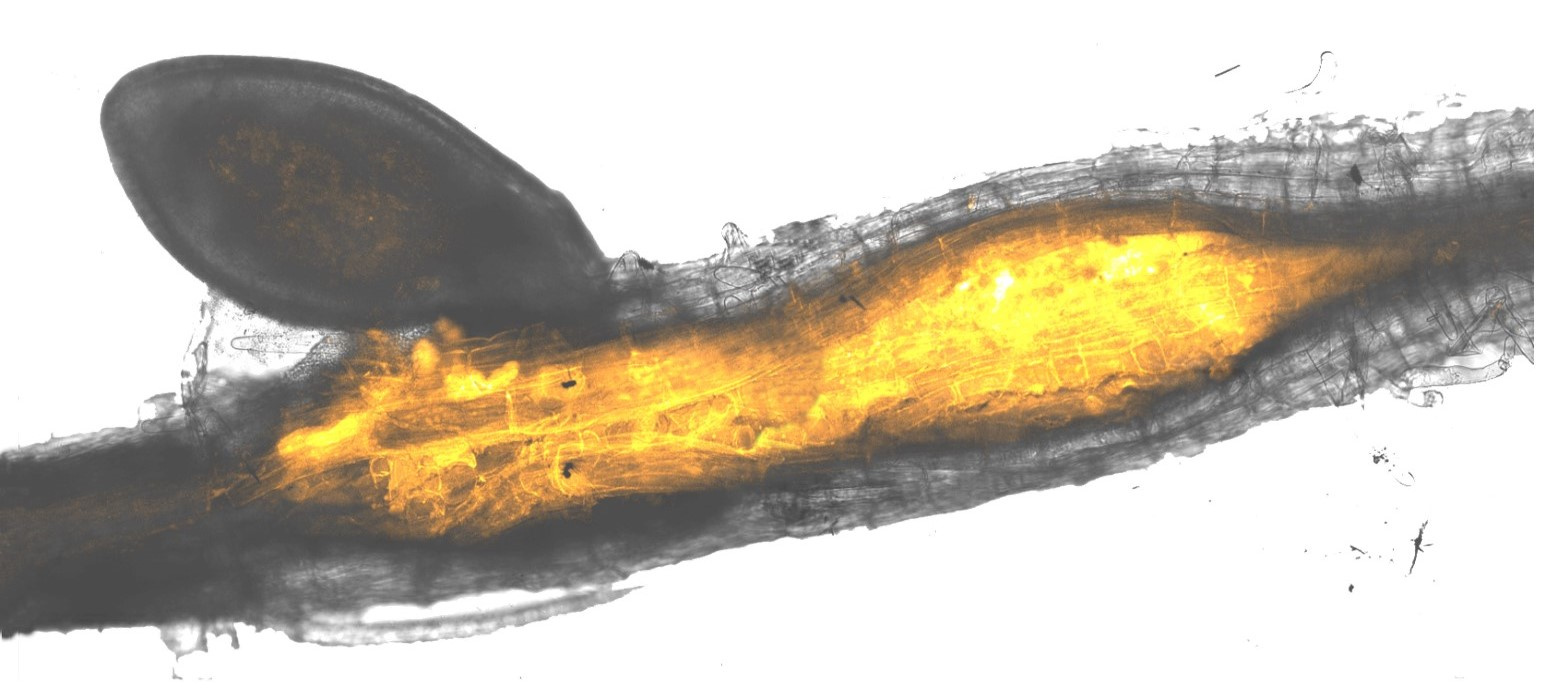
Discovering more S genes is therefore paramount. This paper describes a new way to do this – termed here, “the hologenome theory of S gene discovery”. The term hologenome, coined for use in the hologenome theory of evolution, argues that we should consider the host plus all of its associated symbiotic microbes as a single entity. We used this definition to consider the host and the plant-parasitic nematode – a plant-pathogen of global agricultural importance – as a single unit. Conceptualising them in this way, we could ask, “what is the metabolic capacity of the plant-nematode hologenome?”. Identifying metabolic pathways that were in part contributed by the host genome and in part contributed by the parasite’s genome would highlight genes in the plant that the nematode relies on – and thereby new S genes for CRISPR-mediated crop improvement.
To achieve this, we sequenced the genome of the plant-parasitic nematode Heterodera schachtii (so we know what the genes are), and the transcriptome of the plant-nematode hologenome (so we know when they are deployed by each organism). We could then map these data to metabolic pathways that were partially overlapping between the two organisms and focus on pathways that, for example, start in the host and end in the parasite. We identified several such pathways, the most notable being the Vitamin B5 biosynthesis pathway: the beginning of this pathway is switched on in the plant, and the end of the pathway is switched on in the nematode. These data therefore suggest that the parasitic nematode relies on the beginning of the pathway in the plant in order to cause disease. We tested this idea by removing the early steps in the plant, and found that in these plants we could increase the resistance to nematode infection. Taken together, this demonstrates not only a new S gene, but more importantly a new way to identify S genes: the hologenome theory of S gene discovery.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in