Deceiving viruses: a blueprint for fighting future outbreaks and pandemics?
Published in Healthcare & Nursing
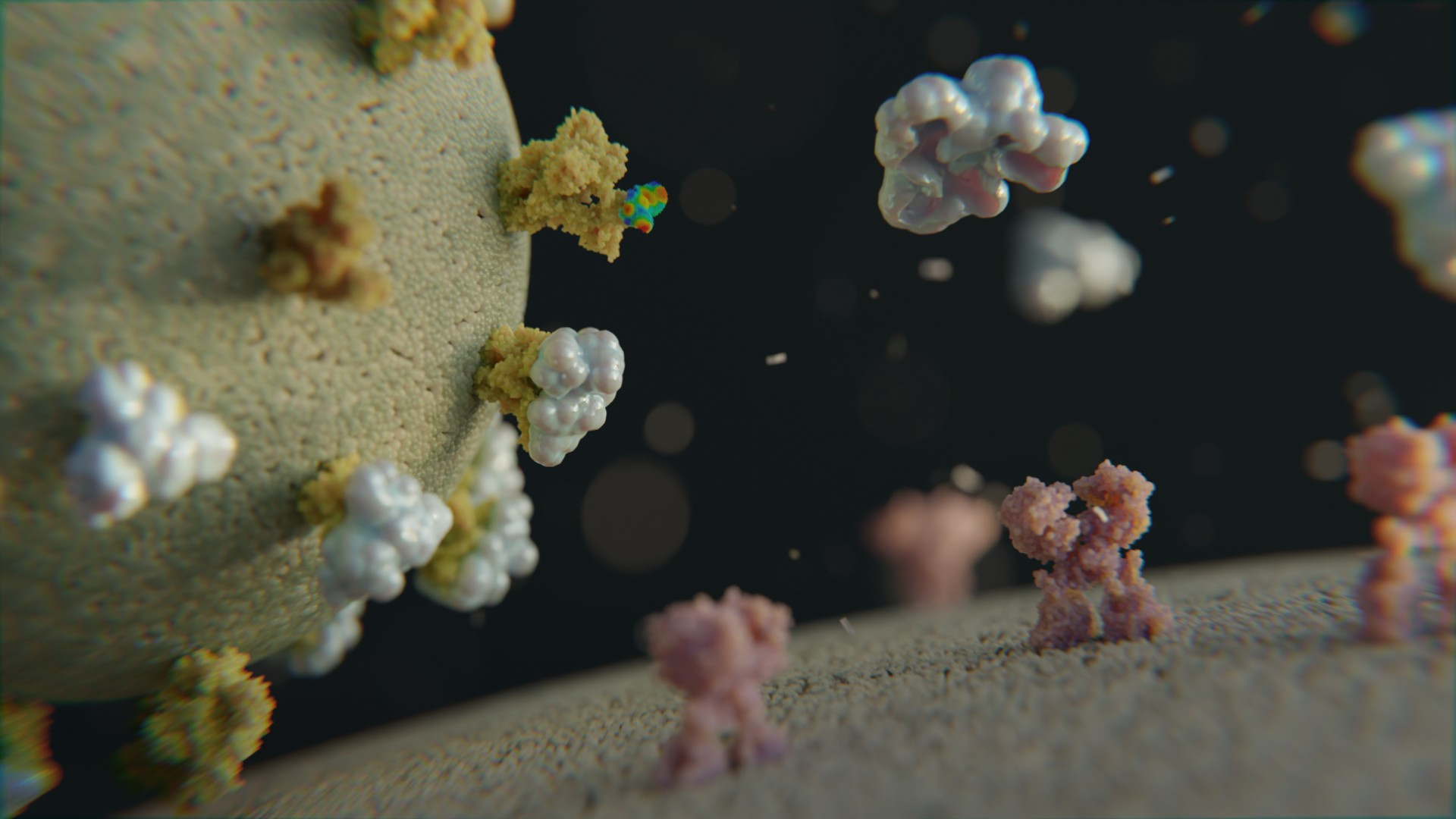

Biologic decoys (looking like human cell receptors) float around in the bloodstream and neutralize the SARS-CoV-2 virus in the body.
But wouldn’t it be awesome if the introduced angiotensin-converting enzyme 2 (ACE2) decoys were engineered to bind to SARS-CoV-2 virus particles much more strongly than the original human cell receptor? To find such candidates, we have modeled the spike-ACE2 complex, trained an artificial neural network with the model results, and thus can make very fast predictions for improved ACE2 variants (and various spike variants as well) that help to drastically speed up the design process.
How exactly? Let’s start at the beginning:
Biologics: the new old megatrend in pharma
The usage of biologics is as old as humanity itself — human breast milk is also considered a biologic. The development of biologics started a bit more recently, but is still an old story. An example of a significant milestone was set in the 19th century: Kitasato and Behring injected diphtheria and tetanus antigens into animals in order to produce therapeutic antibodies in their serum.
Several groundbreaking technologies such as tissue culture and recombinant DNA technology combined with an in-depth understanding of disease mechanisms have fundamentally pushed biologics’ development ever since.
In recent years, biologic drugs have been the fastest growing section of the pharmaceutical market, accounting for 22% of newly approved drugs between 2008 and 2017 in the US. And not to forget: around 40% of the total US pharmaceutical expenditures in 2017 was on biologics [1].
Design and potential of biologics
Biologics revolutionized the available options for disease diagnosis, prevention and treatment. They benefit from high specificity and small side effects, and make it possible to target previously “untreatable diseases”. Indeed, they tend not to harm the patient, because the active ingredient may well be a substance native to the patient’s body.
Biologics can be designed for numerous purposes. Among the most prominent examples are hormones, vaccines, and monoclonal antibodies, to name just a few. This concept even reaches into the realm of personalized medicine, although that is not our emphasis here.
In short, as the International Alliance of Patients’ Organisations (IAPO) and the International Federation of Pharmaceutical Manufacturers and Associations (IFPMA) write in the conclusion of their joint research report on biologic medicines:
Biologics have the potential to benefit millions of patients worldwide – across a large number of conditions.
A booming drug-design paradigm: decoys
Inspired by natural regulatory mechanisms in the human body, therapeutic decoys seem predestined as potent drugs without severe side effects. The remarkable concept of using the binding domains of cellular receptors to trap ligands didn’t get as much attention as monoclonal-antibody therapies in the last few years. Still, its full potential is just beginning to be realized.
Therapeutic-decoy receptors are the second largest branch among approved ligand-targeting agents nowadays. They have already been successfully used to treat several inflammatory diseases, certain forms of anemia and colorectal cancer in a combination therapy [2]. During the COVID-19 pandemic, another use case for decoys has intensified research in the field.
ACE2 decoys against the SARS-CoV-2 virus
Using recombinant ACE2 protein as a treatment was initially considered during the spread of SARS-CoV-1 in the early 2000s. The research was resumed when SARS-CoV-2 emerged in 2019 [3].
The idea is strikingly simple: after infection, the virus binds to cellular receptors on human cells to initiate entry into the cell and to be able to reproduce inside the cell. In order to avoid this, a soluble form of the cellular receptor ACE2 is administered to trap virus particles before they infect human cells.
Copyright © Innophore 2023
However, as is often the case for decoy receptors, modifications are needed to enhance the binding affinity and to optimize pharmacokinetic properties.
Unlocking the potential of biologics: lead optimization
So, how do we unlock the potential of biologics mentioned above in our concrete example? First, we want to optimize a human-ACE2 decoy for binding to the SARS-CoV-2 spike protein. However, we must not forget that the virus is evolving, and new variants are emerging that can have differences in the spike receptor-binding domain (RBD). In general, these differences influence the binding strength and mechanism.
As a result, the idea is to optimize our ACE2 decoy for binding to the spike protein, but for that purpose use a mix of several spike variants instead of just the wild type. More precisely, the goal is to optimize the binding to all major spike variants simultaneously. On top of that, we should be ready to add newly emerging spike variants and adapt optimization to include them quickly.
Molecular-dynamics-simulation models for protein-protein interactions
A standard tool for studying protein-protein interactions (PPI) in silico (i.e. on the computer) is the use of molecular-dynamics (MD) simulations. These MD simulations produce potent models for the prediction of the interaction strength between two protein structures that are encoded for the computational model to understand and use.
This is a more detailed picture of such a simulation: imagine two protein structures, with atoms stored as points in three-dimensional space, linked by molecular bonds. Then, the simulation assumes physical force fields to act on these atoms over time. Time itself is a key element in the simulation: the two structures are input at some point in time, then time is “switched on”, and everything starts moving, simulated by the computer.
Information is taken from the simulation by computing interesting quantities for several timesteps along the way, from which other quantities can be extracted, like a binding affinity.
A special case of PPI: the SARS-CoV-2 spike and the human ACE2 receptor
Since the SARS-CoV-2 spike and the human ACE2 receptor are both proteins interacting with each other, an MD simulation can study their behavior and binding properties. We have looked at this before, and published a paper about the binding affinity of several major SARS-CoV-2 variant spikes to the human ACE2 receptor [4]. You can see an actual real example of this in the video above.
But we have also written a blog post about that paper here on the Nature Portfolio Health Community in the “Behind the Paper” series [5]. So if you are interested in this aspect of our research, we encourage you to read that article next.
Conventional techniques: experiment and computation
It is one thing to compute a binding affinity, but it is also necessary to compare the computational predictions to experimental data. This usually creates a feedback loop, where computational results are tested in costly, time-consuming lab experiments. The experimental results are then used to refine the predictions, new (better) predictions are made, and tested again in the lab, and so on.
We have created a workflow that can help reduce the computational effort or, more precisely, make it much more efficient. This also helps on the lab-testing side: more promising candidates are created more quickly.
CatalophoreTM Halos: more than just imagination
The starting point in our computation that goes beyond the standard protein structure is what we call a Halo: The idea is to encode the function of a protein in fields of physicochemical properties, which basically means that we make biology (easily) machine-readable. These fields are sampled on a discrete grid of points in 3D space. The result of this sampling is a CatalophoreTM point cloud. Here you can see this process visualized for an example structure:
Enter Deep Learning: encoding PPI via opposite Halos
So, we create two Halos for the opposite binding regions in the spike-ACE2 complex and get a pair of point clouds. Why is that so helpful? The idea is that these point clouds encode the PPI in all necessary complexity in order to generate a reliable prediction for the corresponding binding affinity. Let's visualize this setup for the actual spike and ACE2 structures and Halos:
In fact, these Halos are ready to be fed into advanced deep-learning models. So we trained an artificial neural network (ANN) that can use the information contained in the Halos on a lot of binding-affinity data generated from MD simulations. This works to the effect that the ANN is then able to predict the MD result.
So, just take opposite Halos, feed them into an ANN, train, predict, and that's it. If that sounds too easy to be true to you, your intuition serves you well. Actually, this is only the first step: the ANN is able to very, very quickly screen a lot of possible candidates for mutations/variants in both the ACE2 and the spike RBD. As a second step, however, we validate the most promising predictions by running MD simulations. Sometimes, we include further rounds of even more involved simulations [6]. Even further validation is possible in lab experiments.
Multi-step decoy design: a blueprint for the future?
Now imagine this would be an automated workflow that can be executed for any virus or target in the human body (be it during a pandemic [7] or for another use case). That could make the design of analog biologics very efficient. The steps are:
- Get structures for the relevant virus-receptor complex
- Run a lot of MD simulations to generate enough training data, possibly with international partners [8]
- Train an ANN to predict the MD-simulation results
- Generate a lot of interesting (or random) mutations in the virus and/or receptor sequences
- Let the ANN predict them all, almost in an instant
- Validate the most interesting (strongly binding) ANN results via MD simulations
- Take further steps
- Repeat with more data
- ...
No prediction without experimental check!
Apropos further steps: we already mentioned experiments. We also did some. Here you can catch a sneak peek:

© Innophore, March 2021
Our team specialists were trained to work in our partner BSL3 lab at the Medical University of Graz conducting measurements with live samples of SARS-CoV-2 variants in cell models. And the results are promising:
On the one hand, ACE2 decoys effectively prevented the virus from infecting cells. On the other hand, the designed ACE2 variants were indeed more potent in this process than the native ACE2 receptor.
Further reading
The paper in front of this blog post
Read our latest Scientific reports paper Optimizing variant-specific therapeutic SARS-CoV-2 decoys using deep-learning-guided molecular dynamics simulations [9] on nature.com to learn how we leverage the power of our CatalophoreTM Halos combined with deep learning based on molecular-dynamics simulation data to run lead-optimization campaigns for biologics.
About the companies behind the paper
Based in Austria and San Francisco, Innophore is a high-tech spin-off, specializing in the fields of digital drug discovery and enzyme search [10] using 3D point clouds - Catalophores, AI and Deep Learning. Innophore’s vision is to identify and develop high-value industrial and therapeutic enzymes and more efficient, environmentally friendly ‘green’ chemical production processes and novel biosimilars for medical treatments, including contributions to drug repurposing [11], analyses of virus mutational dynamics [12], finding new inhibitors [13], and side-effect prediction using our 3D point-cloud technology.
SignalChem Lifesciences Corporation, headquartered in British Columbia, Canada, is a clinical-stage company developing novel targeted therapies for oncology. Its unique business model has been built upon four important pillars: Drug Discovery and Development, the Bioreagents and Research Services business, In-Vitro Diagnostic Development and Plant Biosynthesis System. A group of scientists with extraordinary expertise and experience in protein engineering and drug discovery are working together cohesively to provide the best products and services to its customers around the world and to maximize the efficiency of its own drug discovery efforts.
References
[1] A. Sarpatwari et al.: The US Biosimilar Market: Stunted Growth and Possible Reforms, Therapeutic Innovations 105 (2019) 92-100
[2] M. M. Attwood et al.: Soluble ligands as drug targets, Nature Reviews Drug Discovery volume 19, pages 695–710 (2020)
[3] K. Kuba et al.: A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus–induced lung injury, Nat Med. 2005; 11(8): 875–879
[4] V. Durmaz et al.: Structural bioinformatics analysis of SARS-CoV-2 variants reveals higher hACE2 receptor binding affinity for Omicron B.1.1.529 spike RBD compared to wild type reference, Scientific Reports volume 12, Article number: 14534 (2022)
[5] C. C. Gruber and V. Resch: Innophore's 3D point clouds provide a head start in monitoring emerging SARS-CoV-2 variants, Nature Portfolio Health community, 2022
[6] A. Singh et al.: Serine 477 plays a crucial role in the interaction of the SARS-CoV-2 spike protein with the human receptor ACE2, Scientific Reports volume 11, Article number: 4320 (2021)
[7] J. Hodgson: The pandemic pipeline, Nature Biotechnology 38, 523-532 (2020)
[8] Open for outbreaks. Nat Biotechnol 38, 377 (2020)
[9] K. Köchl et al.: Optimizing variant-specific therapeutic SARS-CoV-2 decoys using deep-learning-guided molecular dynamics simulations, Scientific Reports volume 13, Article number: 774 (2023)
[10] G. Steinkellner et al.: Identification of promiscuous ene-reductase activity by mining structural databases using active site constellations, Nature Communications volume 5, Article number: 4150 (2014)
[11] C. Gorgulla et al.: A multi-pronged approach targeting SARS-CoV-2 proteins using ultra-large virtual screening, iScience 24, 102021 (2021)
[12] L. Parigger et al.: Recent changes in the mutational dynamics of the SARS-CoV-2 main protease substantiate the danger of emerging resistance to antiviral drugs, Front. Med. 9:1061142 (2022)
[13] M. Prattes et al.: Structural basis for inhibition of the AAA-ATPase Drg1 by diazaborine, Nature Communications volume 12, Article number: 3483 (2021)
Follow the Topic
-
Scientific Reports
An open access journal publishing original research from across all areas of the natural sciences, psychology, medicine and engineering.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026
Health Policy and Systems Research
Publishing Model: Hybrid
Deadline: Jul 14, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in