DrugSolver CavitomiX: A GPU-accelerated tool for drug repurposing and off-target analysis using cavity property point clouds on NVIDIA DGX
Published in Healthcare & Nursing, Social Sciences, and Chemistry
The importance of drug repurposing
Drug repurposing refers to the process of identifying new therapeutic uses for existing drugs that were originally developed and approved for different indications. Instead of starting from scratch and developing a new drug, researchers explore the potential of existing drugs to treat different diseases or conditions. By repurposing drugs, we can bypass many of the time-consuming and costly stages of traditional drug development, such as safety testing and initial clinical trials. This approach offers several advantages, including faster timelines, reduced costs, and the ability to leverage existing knowledge about the drug's safety and pharmacokinetics. Consequently, a lot of bioinformatic approaches focus on this topic. In our paper, we present a new approach: Using cavity point clouds for drug repurposing, which has the potential to detect candidates, which might have been overlooked using traditional structure and sequence-based methods.
Understanding Cavity point clouds
Cavity point clouds provide a unique glimpse into the inner workings of protein cavities. They capture the physico-chemical properties that define these pockets, offering a machine-readable perspective on biological structures and pave the way for leveraging artificial intelligence (AI) algorithms. Comparing cavity point clouds enables us to explore enzymes with similar binding sites, irrespective of structural and sequence similarities.
A cavity represented as a cavity point cloud representing its physio-chemical properties
Embracing New Possibilities: Exploring Drug Repurposing
Cavity point clouds offer a promising avenue for drug repurposing. By utilizing these advanced three-dimensional representations, we can uncover new opportunities for existing drugs and investigate their therapeutic potential in previously unexplored contexts. This approach allows us to identify compounds that may have been overlooked using conventional methods, which are based on structure and/or sequence comparisons. Through the power of cavity point clouds, we aim to unlock untapped potential within known molecules and advance novel treatment strategies for a wide range of diseases.
To test the ability of our DrugSolver CavitomiX for the application of drug repurposing we focused on druggable targets of SARS-CoV-2. We modeled the structures of those enzymes (e.g. for the main protease (Mpro) of SARS-CoV-2, which is crucial for the replication of the virus[1]) and after calculation of the enzyme's structure and modeling its dynamics using molecular dynamics simulations, we calculated the cavity property point clouds. By comparing the SARS-CoV-2 cavities with cavities from databases, we could identify cavities with high similarity to SARS-CoV-2 cavities and a known binding partner. We selected the compounds bound in the most similar cavities and tested them in cell-based viral infection assays.
We were able to identify two compounds, both approved drugs, which showed inhibition of viral replication of SARS-CoV-2 in cell culture. Despite their high active-site cavity similarity, the enzymes which these two compounds (fusidic acid and flufenamic acid) originate from, show very little structural and very low sequence identity to Mpro and TMPRSS2, respectively, and therefore would not have been identified using sequence- and structure-based comparisons.
Cavity comparison of the Mpro with the transcription factor TEAD, from which the inhibitor for the Mpro originated from
Unveiling Potential Side Effects: Off-target Identification using Cavity Point Clouds
Cavity point clouds extend beyond their utility in drug repurposing. They empower us to delve into off-target identification, shedding light on potential interactions outside the primary target. By analyzing similarities in binding-site cavities, we gain valuable insights into possible side effects and unintended interactions, enhancing drug safety and precision. To test this, we studied the enzyme AChE, an essential enzyme in the central nervous system of animals and the target for drugs against Alzheimer’s disease (AD) [2]. The active-site cavity of AChe was compared to dataset cavities. Amongst the enzymes with the most similar cavities, we identified human carboxylesterase 1 (CES1). CES1 is already described to be a target of AChE inhibitors. The enzyme was even suggested to be applied as a bio-scavenger to protect against organophosphorus nerve agents, which are known to bind to AChE [3]. Finding the CES1 among the best hits is a promising result, showing the potential of the DrugSolver CavitomiX to be used for off-target identification.
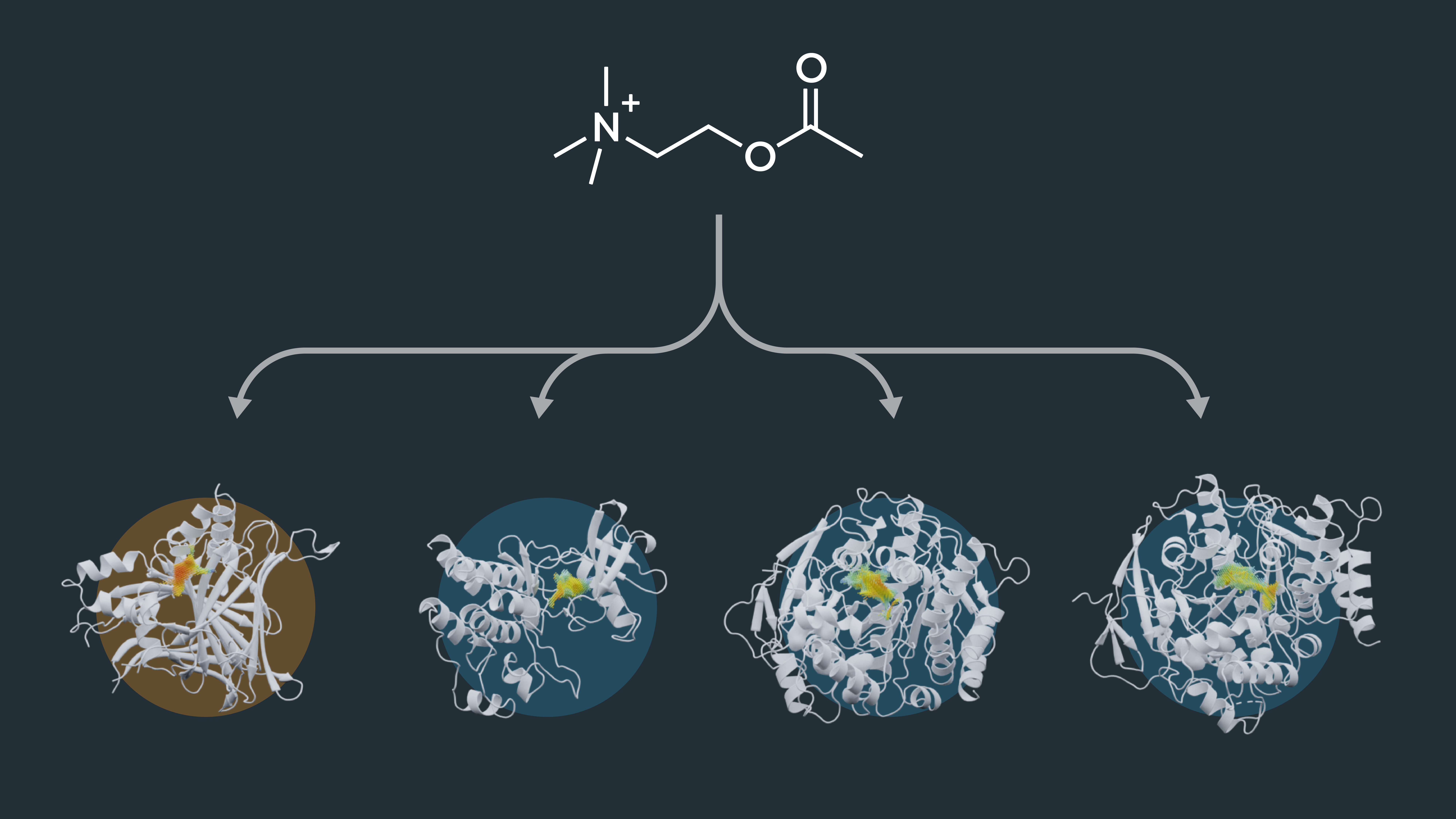
Off-target identification: Enzymes with a similar binding-cavity are potential candidates for binding the drug molecule making them off-targets. After computational screening, a refined list of selected enzymes is tested in further wet-lab experiments.
Speeding up cavity comparisons using GPUs
Efficiently comparing large datasets is crucial in bioinformatics, as such comparisons are computationally expensive. To tackle this computational challenge, we are continuously exploring ways to enhance processing speed. Utilizing GPUs is a powerful strategy to boost processing speed and accelerate various computational methods, such as cavity comparisons, molecular dynamics simulations, or machine learning algorithms. The use of GPUs significantly enhances the efficiency of bioinformatics analyses, enabling us to run our tools with a dramatic speedup, especially on NVIDIA DGX systems. Innophore and our partners at NVIDIA work closely together to optimize GPU utilization, striving for the most efficient acceleration of all processes within the DrugSolver Cavitomix. Utilizing the advanced capabilities of the NVIDIA DGX platform, the computational efficiency for matching Innophore's point cloud datasets has been amplified by a factor exceeding 100. Processes that formerly took approximately 625 seconds to complete can now be executed in a mere 5 seconds.
What’s next?
To further expand the capabilities of the DrugSolver CavitomiX Innophore is currently creating the “Human Cavitome Atlas”. Also in this endeavor, Innophore is partnering with NVIDIA, leveraging the currently most advanced GPUs and NVIDIAs BioNeMo platform for protein structure generation. This atlas will contain all druggable human cavities. This is going to allow us to perform a cavity-based off-target search in all human targets. The integration of the “Human Cavitome Atlas” will further enhance the tool’s ability to identify potential interactions and broaden its applications in drug discovery and off-target analysis.
The Human Cavitome Atlas will include all druggable human cavities, making it a promising tool for drug repurposing and drug discovery
Companies behind the paper
Innophore
Based in Graz, Austria, and San Francisco, California, Innophore is a high-tech spin-off, specializing in the fields of digital drug discovery and enzyme search [4] using 3D point clouds - Catalophores, AI and Deep Learning. Innophore’s vision is to identify and develop high-value industrial and therapeutic enzymes and more efficient, environmentally friendly ‘green’ chemical production processes and novel biosimilars for medical treatments, including contributions to drug repurposing [5], analyses of virus mutational dynamics [6], finding new inhibitors [7], and side-effect prediction using our 3D point-cloud technology.
NVIDIA
NVIDIA is a global technology company that specializes in artificial intelligence (AI) and accelerated computing. The company's GPUs are used in a wide variety of applications, including gaming, machine learning, and data science. NVIDIA has been a pioneer in the field of GPU-accelerated computing, and its products have helped to accelerate the development of AI and machine learning, and also bioinformatics analysis including drug development and drug repurposing. Also Innophore CavitomiX platform uses NVIDIA hardware for accelerating bioinformatics calculation and for machine learning.
Try a method behind this paper yourself
If you regularly work with protein structures, download Innophore's latest Schrödinger PyMOL plugin CavitOmiX for free and generate your own AI-predicted protein structures, using NVIDIA's BioNeMo, DeepMind's AlphaFold or Meta's ESMfold. Furthermore, Catalophore™ cavities can be calculated directly for molecules loaded in PyMOL. More information is also available at the official PyMOL wiki.
Literature
1. Ullrich, S. & Nitsche, C. The SARS-CoV-2 main protease as drug target. Bioorg. Med. Chem. Lett. 30, 127377 (2020).
2. McGleenon, Dynan, & Passmore. Acetylcholinesterase inhibitors in Alzheimer’s disease. Br. J. Clin. Pharmacol. 48, 471–480 (1999).
3. Hemmert, A. C. et al. Human carboxylesterase 1 stereoselectively binds the nerve agent cyclosarin and spontaneously hydrolyzes the nerve agent sarin. Mol. Pharmacol. 77, 508–516 (2010).
4. Steinkellner, G. et al. Identification of promiscuous ene-reductase activity by mining structural databases using active site constellations. Nat. Commun. 5, 4150 (2014).
5. Gorgulla, C. et al. A multi-pronged approach targeting SARS-CoV-2 proteins using ultra-large virtual screening. iScience 24, 102021 (2021).
6. Parigger, L. et al. Recent changes in the mutational dynamics of the SARS-CoV-2 main protease substantiate the danger of emerging resistance to antiviral drugs. Front. Med. 9, 1061142 (2022).
7. Prattes, M. et al. Structural basis for inhibition of the AAA-ATPase Drg1 by diazaborine. Nat. Commun. 12, 3483 (2021).
Follow the Topic
-
Scientific Reports
An open access journal publishing original research from across all areas of the natural sciences, psychology, medicine and engineering.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026
Health Policy and Systems Research
Publishing Model: Hybrid
Deadline: Jul 14, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in