Discovering dominant viral variants and their implications for host-protein binding.
Published in Healthcare & Nursing
Protein flexibility is very crucial for the virus to “negotiate” with the host
Proteins are known to be highly flexible yet efficient workers. For example, a protein can be a strict professional recruiter for a specific molecule like water (aquaporins) or could be a highly adaptable one as well, especially when it comes to immune response (antibodies).
Mechanically, proteins are very specialised in their ability to adopt differential flexibility across their 3D structures. Most often, the interim flexibility in a part of the protein 3D structure governs critical biological functions, like in the case of virus invasion. Here the structural proteins on the virus’s outer surface try to interact with the host cell. At the core level, this process is protein-protein interactions, where a viral protein tries to negotiate with the host cell for a safer passage inside a cell. Both a viral protein and the respective host protein are new to each other; hence they extensively use their flexible arms to interact with each other. Like in the case of SARS-CoV-2, the virus has actively deployed its spike protein to negotiate with the human angiotensin-converting enzyme 2 (hACE2) receptor proteins, primarily present on the alveolar cell lines of the lungs.
Digging out the most flexible spot in SARS-CoV-2’s “arms”
During the onset of the SARS-CoV-2 pandemic, we constantly monitored the emerging amino-acid changes in spike protein sequences. We aimed to discover the most frequent amino-acid exchanges in spike protein, especially inside the receptor binding domain (RBD), which is spike protein’s most exposed part to the hACE2 receptor. Analyzing spike sequences available on the GISAID database, we tracked down S477, the most variable position in RBD then. Driven by curiosity, we filtered the SARS-CoV-2 variants spreading in Austria. To our surprise, S477G (an abbreviation for substituting serine residue at the 477 position with a much smaller glycine residue) was among Austria’s leading SARS-CoV-2 sequences collected from infected individuals. On the contrary, S477N was the most circulated variant of native S477 across the globe. To understand these two particular S477 variants of SARS-CoV-2, we first performed a flexibility analysis, showing that S477 structurally constitutes the most flexible part of RBD, which could directly influence hACE2:RBD crosstalk during viral membrane fusion with the human cell membrane.
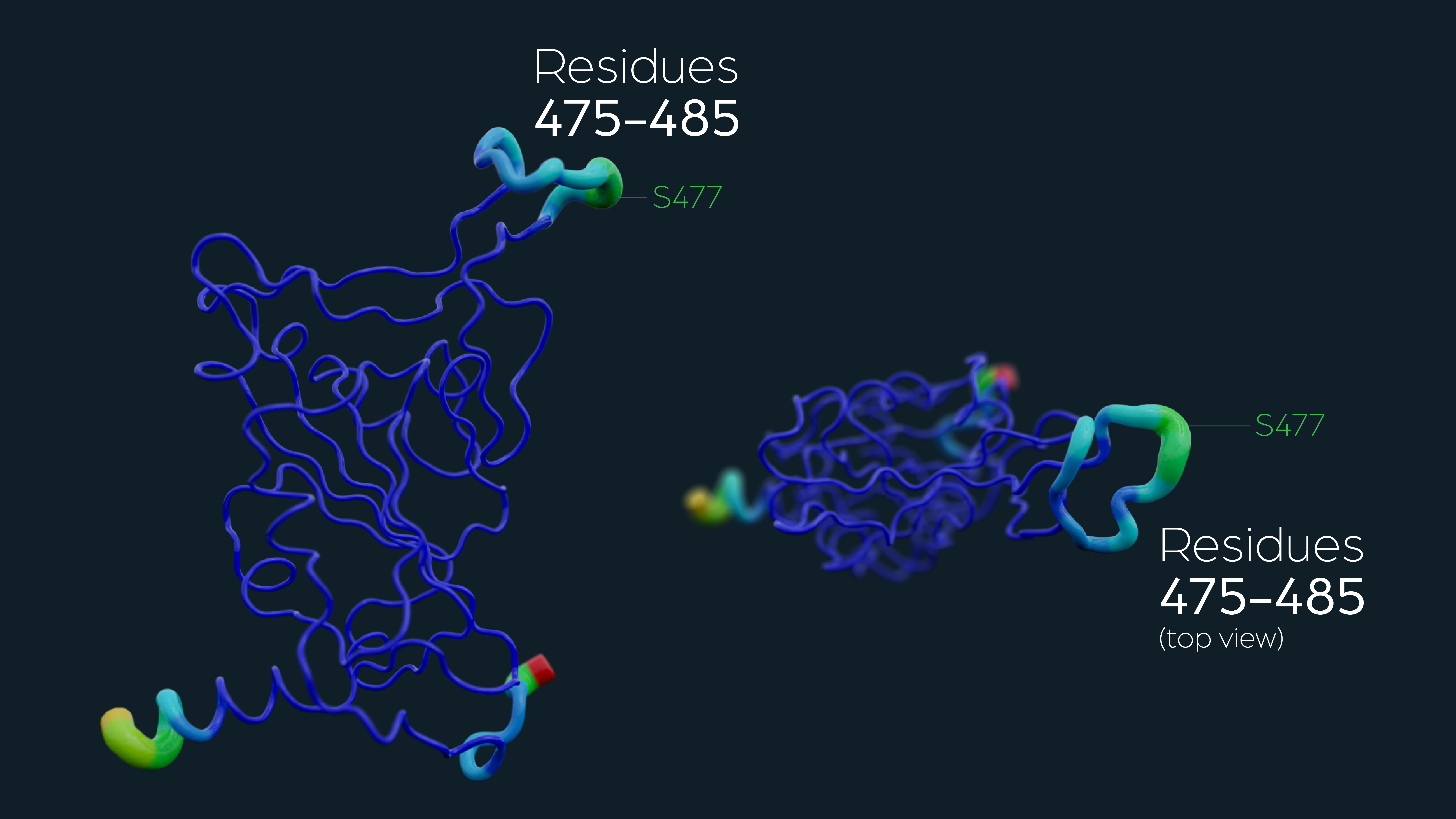
Molecular dynamics simulations are indispensable
The onset course of the SARS-CoV-2 pandemic was a novel and highly demanding situation for structural biologists. In such challenging times, computational techniques have become indispensable. Therefore, we opt for molecular dynamics (MD) simulations to understand the significance of S477 variants in the context of the hACE2:RBD interactions. Various other factors lead to preferring MD simulations, including but not limited to its ability to run purely on chips, reproducibility and sharability, minimal labor requirements, its ability to adopt simplistic modular solutions to complex biomolecular problems, and most importantly, the minimum safety and ethical constraints. By nature, MD simulations are physics-based techniques often employed to comprehend biomolecular dynamics at the atomic level. For example, the ingrained crosstalks between the atomic species at protein-protein or protein-drug interaction interfaces can be characterized by MD simulations. It’s worth mentioning that for the first US FDA-approved HIV integrase inhibitor, MD simulations reveal a hidden druggable site [1] on HIV integrase that was further utilized by Merck & Co. to develop the antiretroviral drug Raltegravir [2], which is among one of the examples that signify the importance of protein flexibility characterized by MD simulations.
Leveraging the power of MD simulations to reveal the significance of S477 and its “pro” variants for host protein interactions
With limited access to laboratories during the pandemic, we were unconditionally allowed to utilize our high-performance supercomputer facility at Innophore and Vienna Scientific Clusters (VSC) to gain more insight into S477 variants using MD simulations. We analyzed the impact of S477G and S477N substitutions on hACE2:RBD interactions using “footprint” analysis of MD simulations; both variants clearly stand out, as depicted in the figure below. S477N has shown the highest contact with hACE2, followed by S477G and the native S477.
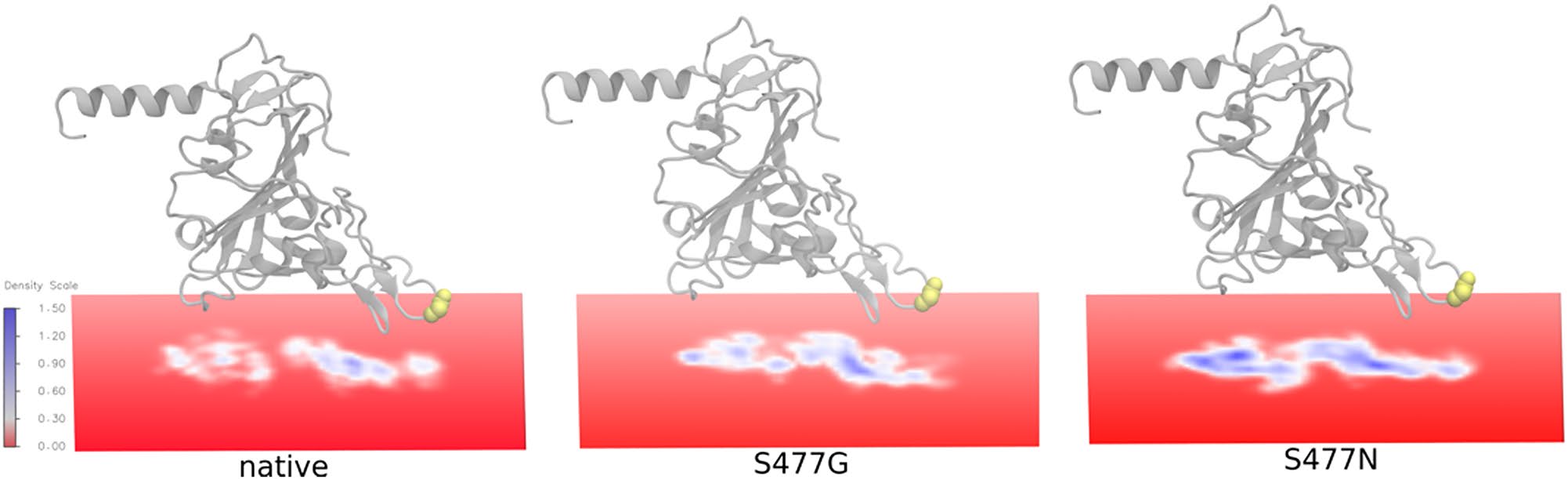
Furthermore, we have employed umbrella sampling, an advanced MD simulation technique often used to predict a particular protein-protein interaction’s strength. Our detailed structural analysis demonstrated that the compared to native S477, S447G, and S477N variants would strengthen the SARS-CoV-2’s spike protein binding with hACE2. Our findings have been followed by experimental validation [3,4].
Our structural advice became highly “transmissible”
After that, S477N emerged as an integral part of various SARS-CoV-2 lineages. Among them, the B.1.526 lineage has a 35% more transmissible rate than the non-variant virus, resulted in a sharp increase in infections from late 2020, and remained dominant from March to May 2021 [5]. Particularly in New York, the S477N variant was spreading at an alarming rate [6] and subsequently became part of the “super-spreader” Omicron variant [7]. Both of our simulated variants are also reported to be highly resistant to antibodies [4]. Till today (08.03.2023), out of 15,138,488 sequences deposited on GISAID, S477N (7,234,065) and S477G (1337) combinedly appears in around half of them.
Our suggested first line of approach for tackling the emergence of virus variants
- Sequence Collection: Collect deposited sequences from authorized and legitimate viral databases like GISAID.
- Filtering the dominating variants: Perform frequency-based sequence analysis like Shannon Entory or Surprisal Index to monitor the position-specific changes across viral proteins.
- Flexibility Mapping: Perform Normal Mode Analysis (NMA) of the target viral protein, followed by the structural mapping of dominating variants. This step would reveal the most potent and flexible variants for host protein interactions.
-
Binding Energy Prediction: Using advanced MD simulations like Umbrella Sampling, assess the effect of dominating variants on host protein binding.
About the companies behind the paper
Based in Austria and San Francisco, Innophore is a high-tech spin-off, specializing in the fields of digital drug discovery and enzyme search [8] using 3D point clouds - Catalophores, AI and Deep Learning. Innophore’s vision is to identify and develop high-value industrial and therapeutic enzymes and more efficient, environmentally friendly ‘green’ chemical production processes and novel biosimilars for medical treatments, including contributions to drug repurposing [9], analyses of virus mutational dynamics [10], finding new inhibitors [11], and side-effect prediction using our 3D point-cloud technology.
[1] Schames, J. R. et al. Discovery of a Novel Binding Trench in HIV Integrase. J. Med. Chem. 47, 1879–1881 (2004).
Follow the Topic
-
Scientific Reports
An open access journal publishing original research from across all areas of the natural sciences, psychology, medicine and engineering.
Related Collections
With Collections, you can get published faster and increase your visibility.
Computational biology and mathematical modelling of biological systems
Publishing Model: Open Access
Deadline: Jul 18, 2026
Water pollution and advanced treatment processes
Publishing Model: Hybrid
Deadline: May 31, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in