DIA metaproteomics is ready to take off
Published in Microbiology

Metaproteomics has shown the capability to reveal not only taxonomic composition of complex microbial communities, such as gut microbiota, but also deep function information regarding the dynamic host-microbiota interactions. Due to the extreme complexity and heterogeneity of gut metaproteome, accurate protein identification and quantification in metaproteome is a severe challenge compared to the conventional proteomics of a single organism.
During the past years, while most proteomic and metaproteomic studies adopt traditional mass spectrometry (MS) approaches based on data-dependent acquisition (DDA), data-independent acquisition (DIA) techniques have shown outstanding performance in conventional proteomics with increased proteome coverage, reproducibility, and accuracy in quantification1. In 2020, Aakko et al. pioneered the use of DIA in metaproteomics and reported a proof-of-concept study to demonstrate its technical feasibility2. In the same year, our team also reported one of the earliest DIA-based metaproteomic studies, in which more than 30,000 proteins were identified and quantified from gut microbiota in a real clinical cohort3. As an emerging technique, there may be matters of concern about the trustworthiness of the DIA-based metaproteomic methods, which is exactly the main message this paper is expected to convey.
DIA changes how metaproteomic data are generated
In a traditional DDA-based proteomic analysis, MS2 acquisition is triggered by precursor peaks in an MS1 scan (in other words, “dependent” on the MS1 scan). Protein identification is achieved via database searching of peptide fragments in MS2 spectra and thus information of a peptide precursor would be lost if it had not been selected for MS2 fragmentation. What is worse, the stochastic precursor selection can result in the “missing value” problem, which is more significant in metaproteomics compared to conventional single organism proteomics due to the significantly enhanced sample heterogeneity, complexity and dynamic range.
In contrast, DIA performs a sequence of MS2 scans in an acquisition cycle, where precursor ions within defined isolation windows are reproducibly fragmented. This feature allows systematical and unbiased recording of fragmentation information of theoretically all peptides present in a sample. It is even somewhat overdrawn to say that the sample has been digitized for long-term storage. Reanalysis using more advanced informatic solutions in the future may contribute to biological findings that are undetectable currently.
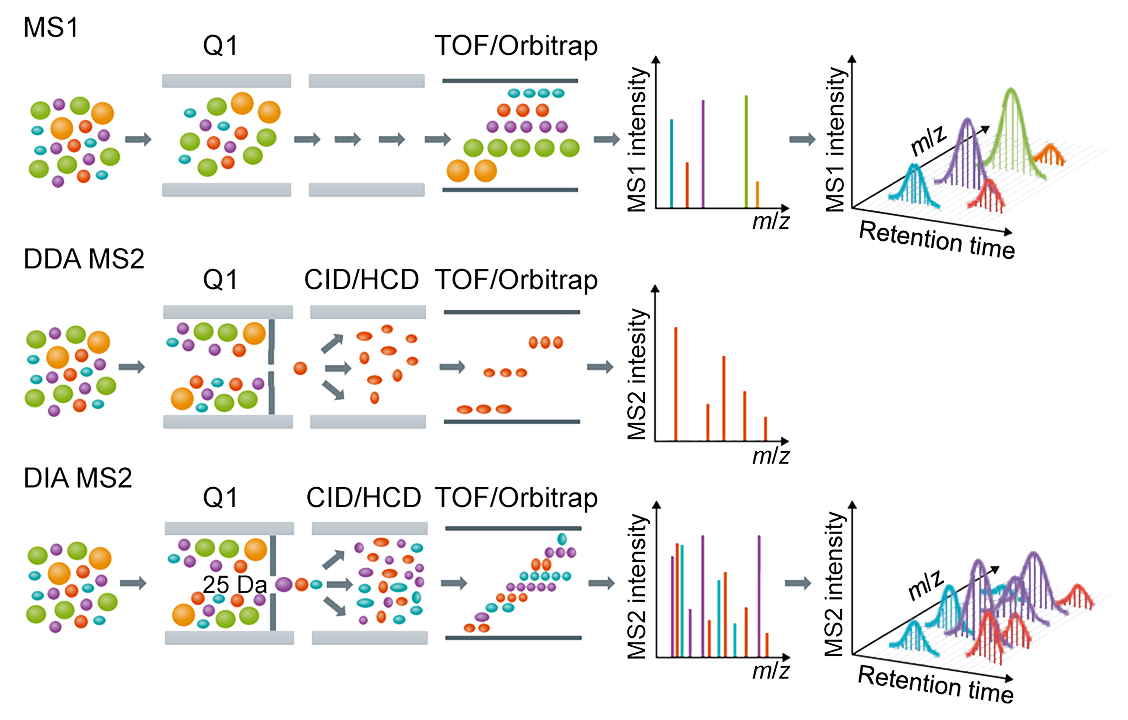
Adapted from ref. 1, under CC BY 4.0 license.
DIA deepens metaproteome coverage and improves quantitative accuracy
Popular quantification strategies other than DIA include DDA-based label-free quantification (LFQ) and tandem mass tag (TMT) isobaric labeling quantification. In a typical LFQ-DDA analysis, protein quantification is based on precursor ion intensities. In a TMT analysis, protein quantification is based on the reporter ions generated by the isobaric labels after fragmentation.
In this paper, we have benchmarked DIA and the other two quantification strategies on human fecal metaproteome samples spiked with 6 known bacterial species. DIA outperformed LFQ-DDA with superior proteome coverage and data completeness. Compared with LFQ-DDA, DIA yielded approximately 80% more proteins and 55% more peptides that were validly identified and quantified (with no more than 1/3 missing values).
From the quantified proteins, those uniquely belonging to the 6 species were selected to evaluate the quantitative accuracy of the methods. We demonstrated that DIA outperformed TMT with higher quantitative accuracy. This result is different from those on simpler mixture samples4 and simulated microbial communities of up to 12 species, where DIA showed quantitative accuracy slight worse than or close to TMT. (We greatly appreciate the reviewers’ suggestion on mimicking the complexity of real metaproteome samples.) Due to the bias came from superposition of reporter ion intensities of co-isolated and co-fragmented peptides in complex metaproteome samples, TMT suffered from ratio compression that measured fold changes deviated from theoretical values toward a 1:1 ratio. In DIA, protein quantification can be performed based on peptide-specific fragment chromatogram profiles to alleviate interference from other peptides, improving the quantification accuracy.
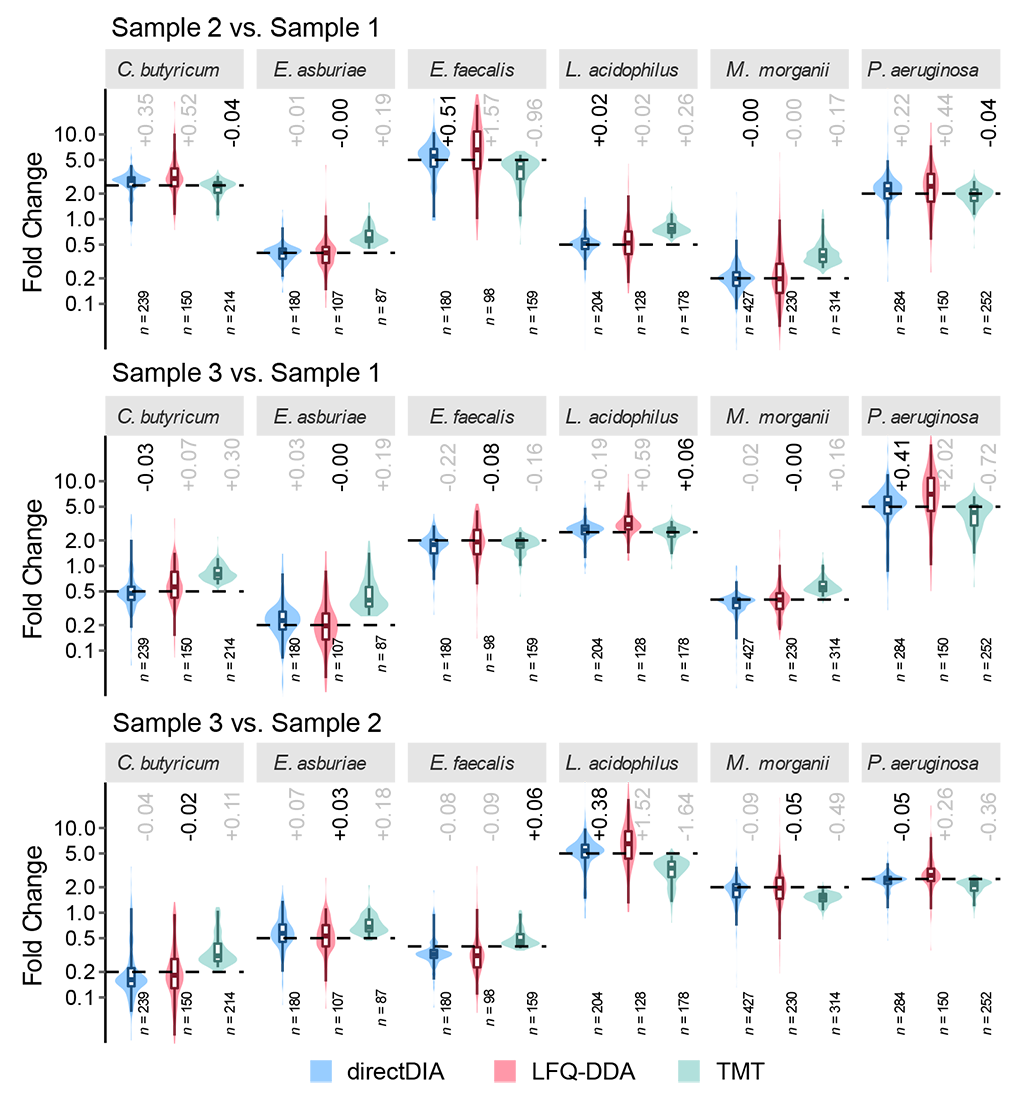
DIA no longer requires external spectral libraries
In the past, a main drawback of DIA was that its data analysis required a spectral library, which is usually built from fractionated samples by DDA. More recently, Pietilä et al. introduced library-free DIA for metaproteomic analysis5. With improvements of informatic solutions by incorporating deep learning6, experimental spectral libraries are no longer necessary for DIA analysis.
In this paper, we compared the library-free workflow in Spectronaut, directDIA, with alternative DIA data analysis methods based on experimental and predicted spectral libraries. Our results demonstrated that directDIA outperformed library-based approaches on complex proteome samples of microbial communities.
The only prior knowledge for directDIA data analysis is the protein sequence database. We applied directDIA to characterize human gut microbiota in two real clinical cohorts. Our results demonstrated that directDIA cooperated well with the mainstream metaproteome sequence database choices7, i.e., an experimental metagenome of the sample and a “pseudo-metagenome” assembled from publicly available sequences.
This study indicates a promising prospect of DIA in deep characterization of gut microbiota. With further innovations in instrumentation and informatics, DIA metaproteomics is bound to shine forth with all its strength. We hope our work can promote broader adoption of DIA metaproteomics in microbiome research.
References
- Ludwig, C. et al. Data-independent acquisition-based SWATH-MS for quantitative proteomics: a tutorial. Mol. Syst. Biol. 14, e8126 (2018).
- Aakko, J. et al. Data-independent acquisition mass spectrometry in metaproteomics of gut microbiota—implementation and computational analysis. J. Proteome Res. 19, 432-436 (2020).
- Long, S. et al. Metaproteomics characterizes human gut microbiome function in colorectal cancer. npj Biofilms Microbiomes 6, 14 (2020).
- Muntel, J. et al. Comparison of protein quantification in a complex background by DIA and TMT workflows with fixed instrument time. J. Proteome Res. 18, 1340-1351 (2019).
- Pietilä, S., Suomi, T. & Elo, L. L. Introducing untargeted data-independent acquisition for metaproteomics of complex microbial samples. ISME Commun. 2, 51 (2022).
- Demichev, V., Messner, C. B., Vernardis, S. I., Lilley, K. S. & Ralser, M. DIA-NN: neural networks and interference correction enable deep proteome coverage in high throughput. Nat. Methods 17, 41-44 (2020).
- Tanca, A. et al. The impact of sequence database choice on metaproteomic results in gut microbiota studies. Microbiome 4, 51 (2016).
Follow the Topic
-
npj Biofilms and Microbiomes
The aim of this journal is to serve as a comprehensive platform to promote biofilms and microbiomes research across a wide spectrum of scientific disciplines.
Related Collections
With Collections, you can get published faster and increase your visibility.
Microbial endocrinology
Publishing Model: Open Access
Deadline: Oct 21, 2026
Microbiome and energy metabolism
Publishing Model: Hybrid
Deadline: Dec 06, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in