Extracellular RNA is directly involved in building Pseudomonas aeruginosa biofilm matrix
Published in Bioengineering & Biotechnology, Microbiology, and Protocols & Methods

The established tenet for biofilm extracellular polymeric substance (EPS) matrix formation is that RNA is too unstable to be anything more than a short-lived messenger molecule. This, however, has been challenged by the existence of extracellular DNA (eDNA) as a key biofilm network component and the subsequent discovery that extracellular RNA (eRNA) bound to eDNA contributes to biofilm viscoelastic network formation, and hence assembly of the EPS matrix. The biofilm matrix structure and function are not black and white.
The first indication the biofilm matrix is not clear-cut, as with everything else in the world of biomolecules, was the observation that extracellular DNA (eDNA) is a key biofilm network component. This was described firstly in biofilms of the pathogen and widely accepted model biofilm former Pseudomonas aeruginosa, and then in various other important clinical and environmental biofilms. Once established that eDNA was not so different from chromosomal DNA, it then became a question of how the bacterial DNA manages completely different functions, inside and outside cells, and importantly, how the DNA undergoes phase transitions into a viscoelastic network. For a research community more accustomed to assigning specific molecules for particular extracellular functions, resolving how DNA achieves this was a challenge that was not dissimilar to that faced earlier by researchers studying intracellular processes. The latter group identified phase transitions as playing an important role in organising cellular functions, often coordinated by molecules with either no clear function owing to lack of structural order, or with alternative primary functions. One such molecule is RNA.
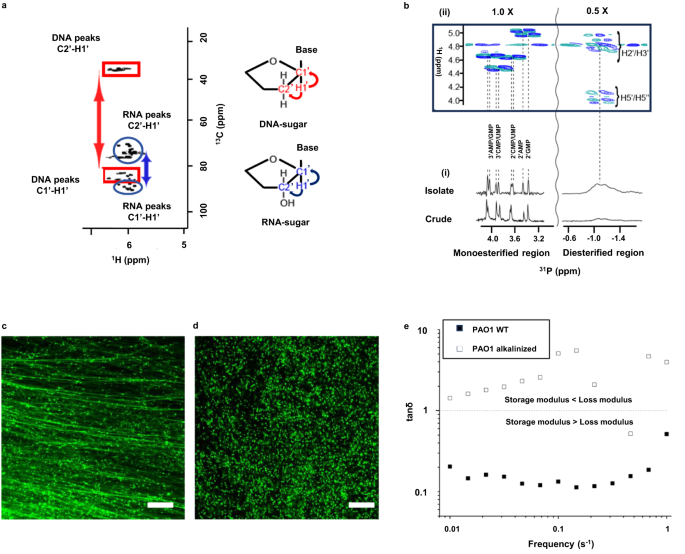
RNA is often detected in the extracellular matrix of biofilms, however this has always been attributed to cell lysis and of no direct consequence to biofilm properties. In a previous study by our research team, we isolated a viscoelastic extracellular nucleic acid gel from a model Pseudomonas aeruginosa biofilm that had the same phase transition behaviours as the biofilm: they dissolved at the same temperature and after caustic soda addition (i.e., alkalinisation). Here, we observed that the alkalinised extracellular gel contained eDNA in chain form, and RNA as monoribonucleotides that only appeared after alkalinisation. RNA was therefore transesterified, or broken down into its subunits, as the biofilm dissolved. This led us to ponder whether extracellular RNA directly contributes to the assembly of the viscoelastic extracellular matrix. It also suggests that the common method for Pseudomonas aeruginosa biofilm dissolution, involving caustic soda addition, might work because it in fact transesterifies the RNA.
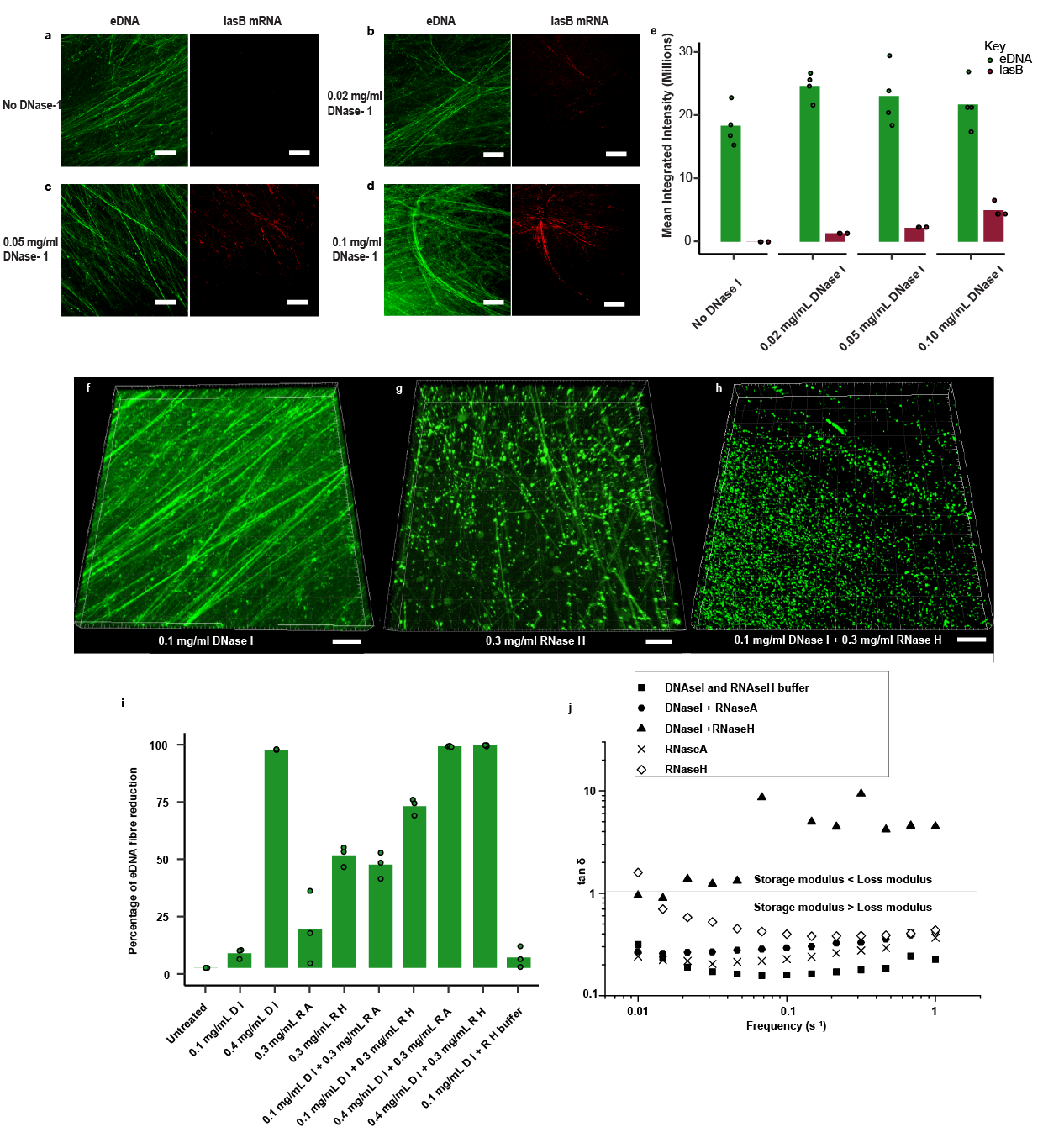
While digesting the biofilm with an enzyme targeting RNA didn’t have the same effect of complete dissolution that was observed for caustic soda exposure, it did slightly reduce biofilm viscoelasticity. We identified several putative eRNA targets that were enriched in the biofilm matrix by comparing RNA levels from different extraction methods favouring extracellular and intracellular RNA. Our challenge was then to observe these in the biofilm matrix, specifically, to assess their contribution to the viscoelastic network. Discerning individual molecules, particularly RNA, in biological samples can be like finding a needle in a haystack owing to their small size, susceptibility to degradation, and the possibility that there are thousands of similar molecules crowded into the same space. Hence, we amplified the detection signal intensity using oligoribonucleotide probes complementary to our putative eRNA, allowing multiple fluorophores to bind. While an initial outcome was not evident, we knew from previous studies that eDNA contributed to matrix structure, and suspected a similar role for eRNA. A viscoelastic eDNA network is clearly visible throughout biofilms of Pseudomonas aeruginosa and many other biofilms, and we hypothesised that the oligoribonucleotide probes were blocked from accessing the eRNA as it was bound to eDNA in these networks. This was confirmed following addition of a mild DNase concentration that exposed the eRNA but did not otherwise degrade the eDNA fibres or dissolve the biofilm. This resulted in fluorescent signals associated with eDNA fibres.
One particularly abundant transcript was lasB, which elicited very strong signals in our model biofilm as well as in sputum samples from patients with high P. aeruginosa microbial burden lung infections. lasB knockouts still formed biofilms with eDNA networks, and we did not observe a lasB oligoribonucleotide signal in lasB knockout negative control biofilms. So, while lasB is not necessarily an essential component of the biofilms, it does serve as a biomarker for the presence of eRNA in clinical and model Pseudomonas aeruginosa viscoelastic eDNA networks. Furthermore, while its expression in the matrix relative to total cell expression was the greatest, lasB, along with several genes with high relative expression, were evident in matrix fibres, while others with an overall higher total cell expression were only observed within cells. The appearance of eRNA in the network is decoupled from total cell expression and could therefore be a regulated process. Finally, further evidence that eRNA bound to eDNA contributes to biofilm viscoelastic network formation was provided when the P. aeruginosa biofilm undergoing the same mild DNase pretreatment completely dissolved following subsequent RNAse digestion. Hence, we could conclude that specific mRNA associated with the biofilm matrix enabled eDNA to form viscoelastic networks through eDNA:eRNA hydrids, and either DNA binding, or non-canonical interactions contribute to eRNA stability in the matrix.
We reported previously the importance of non-canonical base pairs to matrix viscoelasticity. The increased tendency of RNA to form non-canonical base pairs could enable a functional role in matrix networks. Also, RNA is widely reported to mediate the phase separation of biomolecules inside cells into intracellular compartments. Our study suggests that this phenomenon could occur outside the cells. Indeed, RNA is more than just a discarded intermediary molecule in Pseudomonas aeruginosa biofilm formation, that contribute directly to forming the viscoelastic structural matrix scaffold responsible for so many important biofilm properties.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in