Exploring the enigma of whole-genome duplications during early vertebrate evolution: unraveling the hagfish tale
Published in Ecology & Evolution and Genetics & Genomics
Beyond the botanic world, WGD is also a hot subject in the realm of vertebrate evolution. Numerous studies have posited a connection between WGDs and the morphological evolution of early vertebrates. This is classically exampled by the shift from one single Hox gene cluster in invertebrates to four in most vertebrates—bar teleost fishes due to an additional WGD.
Within the Vertebrata, the hagfish stands out not only as a fascinating creature but also as a pivotal one, given its place in the phylogenetic tree. Hagfish and lamprey are the only surviving representatives of a lineage of vertebrates without a jaw (agnathans), and together form the group called cyclostomes (round mouth), which, importantly, share with the rest of vertebrates (known as jawed vertebrates), the last common ancestor of all modern vertebrates (including us, humans). However, a minority of persists in adhering to the conventional perspective, which excludes the hagfish from a group fromed by lampreys and jawed vertebrates. Interestingly, both hagfish and lamprey possess six Hox clusters. However, the role of WGDs in this process was still shrouded in mystery. Furthermore, the evolutionary history of WGD events along the early evolution of vertebrates was a very interesting yet unresolved question.
To unravel the mysteries of vertebrate WGDs and shed light on their evolutionary impact, we must turn our attention to the hagfish. This was the envision that two authors of this paper, Drs Juan Pascual-Anaya and Wen Wang, settled in the late 2016. Juan collected invaluable testis and embryo samples from the wild, while Wen’s Ph.D. student, Yandong Ren, tackled the challenging task –due to the characteristics of the hagfish genome (Kohno et al., 1998)— of assembling the hagfish genome. Collaborating with the Vertebrate Annotation Team of Ensembl, which was spearheaded by Dr. Fergal J. Martin, our hagfish genome was meticulously annotated. Subsequently, Dr. Yong E. Zhang and I, a Ph.D. student in his lab, conducted the analyses of WGD evolution.
Fast forward to a hot summer day in 2018, Juan flew from Japan to Xi’an, China, to visit Wen's lab, where we engaged in an extensive discussion about the project. By then, Yandong had achieved a scaffold-level genome assembly for the hagfish, and I had spent a year wrestling with the enigma of ancient WGDs without a decisive breakthrough. The absence of chromosome-level assemblies for crucial species such as lamprey or amphioxus at the moment, the closely related outgroup species of vertebrates, presented a formidable hurdle in deciphering these early WGD events. Finally, that meeting led to a key decision for the project: to obtain Hi-C data to elevate the hagfish assembly to a chromosome-level resolution. During this period, the potential utility of echinoderm genomes was brought to my attention. Subsequently, it proved instrumental, as the karyotype of the sea cucumber closely mirrors that of the amphioxus, making it a valuable vertebrate outgroup.
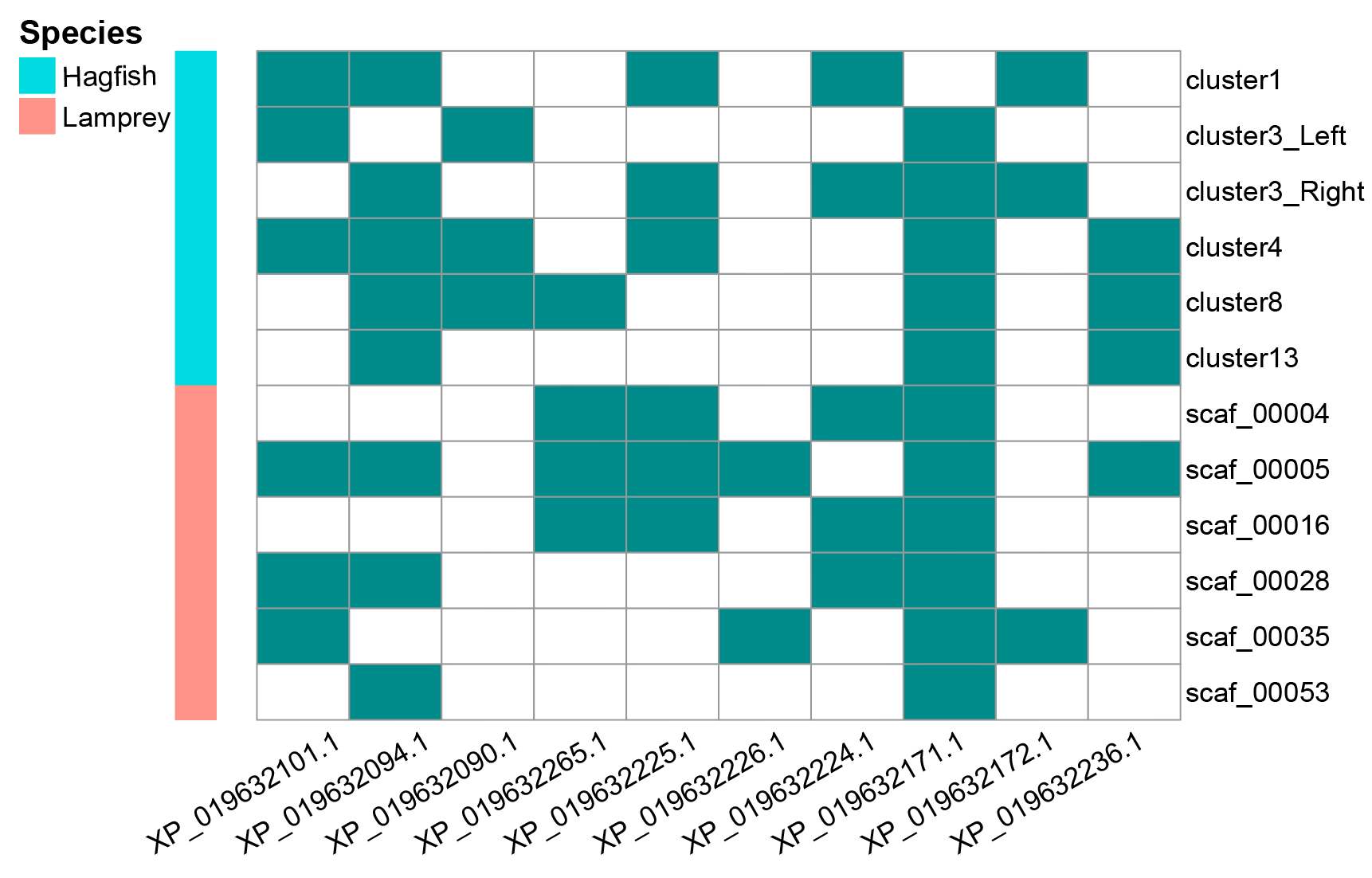
Figure 1. A very preliminary exploration of the existence of Hox genes on cyclostome chromosomes led to the latter development of overlapping ratio metrics. Gene IDs with “XP_” prefix are lancelet Hox genes identified based on known chicken Hox genes.
Subsequently, steady progress was made. Exploiting conserved synteny (gene orders in chromosomes), I aligned gnathostome chromosomes (represented by chicken and spotted gar) with sea cucumber chromosomes, allowing the inference of 17 vertebrate ancestral chromosomes (ACs). The genes of Belcher’s lancelet were then anchored onto these ACs, allowing us to reconstruct 17 chromosome-level phylogenetic trees and resolve the details of gnathostome 2R. With the gnathostome ohnologues (duplicate genes generated by WGD, and named as such in honour to the late Susumu Ohno) in these trees as a control, we deduced that cyclostomes only shared the first round of gnathostome WGD. This deduction is reinforced by the absence of gnathostome pre-2R chromosomal fusions in cyclostomes.
Then, how about the WGD history of cyclostomes? The extensive genome shuffling in hagfish disrupted synteny, necessitating the development of an alternative strategy. However, before I could address this challenge, the onset of the COVID-19 pandemic occurred. While situated in my hometown, I continued my work remotely. In May 2020, prompted partly by curiosity and partly by reasons elusive to me, I investigated the presence of Hox genes on cyclostome chromosomes, using Belcher’s lancelet Hox gene as the reference. Leveraging my bioinformatics background, I opted to visualize gene arrangements in R language, representing presence with color and absence with blanks. Upon examining the preliminary plot, I began to recognize that homologous chromosomes should exhibit a higher retention of shared genes than initially anticipated. Transforming the presence/absence information of outgroup genes into a binary code (1/0), I devised an inter-chromosomal similarity index, or overlapping ratio metric. Applying this new metric, I confirmed two rounds of WGDs in gnathostomes. Subsequently, I extended this analysis to the hagfish and lamprey, inferring a genome triplication in the cyclostome ancestor, and supporting thus the previous findings done by Aoife McLysaght and her team together with that of Byrappa Venkatesh (Nakatani et al., 2021). Consequently, a single round of WGD transpires in the common ancestor of vertebrates, succeeded by lineage-specific WGD events in both gnathostomes and cyclostomes.
.jpg)
Dr. Masahiro Uesaka at RIKEN, Japan, prepared ATAC-seq data from hagfish embros, a very precious material that only the laboratory of Prof. Shigeru Kuratani has been able to obtain regularly (see Juan's one blog for more details). Uesaka-san, together with a graduate student from Yong’s lab, Jieyu Shen, used these data to investigate the cis-regulatory evolution of this species, with a specific focus on ohnologues. At the genomic scale, we found that the hagfish possesses intricate and sophisticated regulatory networks during development, akin to those found in gnathostomes (Marlétaz et al., 2018). When examining ohnologues, hagfish ohnologues mirrored those of gnathostomes not only in regulatory evolution patterns but also in their association with developmental functions. These findings suggested that, at the micro level, the different WGD histories have led to a comparable degree of regulatory innovation in both gnathostomes and cyclostomes. Nevertheless, at the macro level, gnathostomes are characterized by a more diversified morphological spectrum, likely attributed to gnathostome-specific WGD, as demonstrated by analyses conducted by the team of Prof. Phil Donoghue, at the University of Bristol. Hence, WGDs do not invariably result in radical morphological evolution in vertebrates.
Collectively, our collaborative endeavors delineated the panorama of ancient vertebrate WGDs and their functional implications.
Follow the Topic
-
Nature Ecology & Evolution
This journal is interested in the full spectrum of ecological and evolutionary biology, encompassing approaches at the molecular, organismal, population, community and ecosystem levels, as well as relevant parts of the social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Biodiversity and ecosystem functioning of global peatlands
Publishing Model: Hybrid
Deadline: Jul 27, 2026
Understanding species redistributions under global climate change
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in