How birders helped pinpoint hotspots for migratory bird conservation
Published in Ecology & Evolution
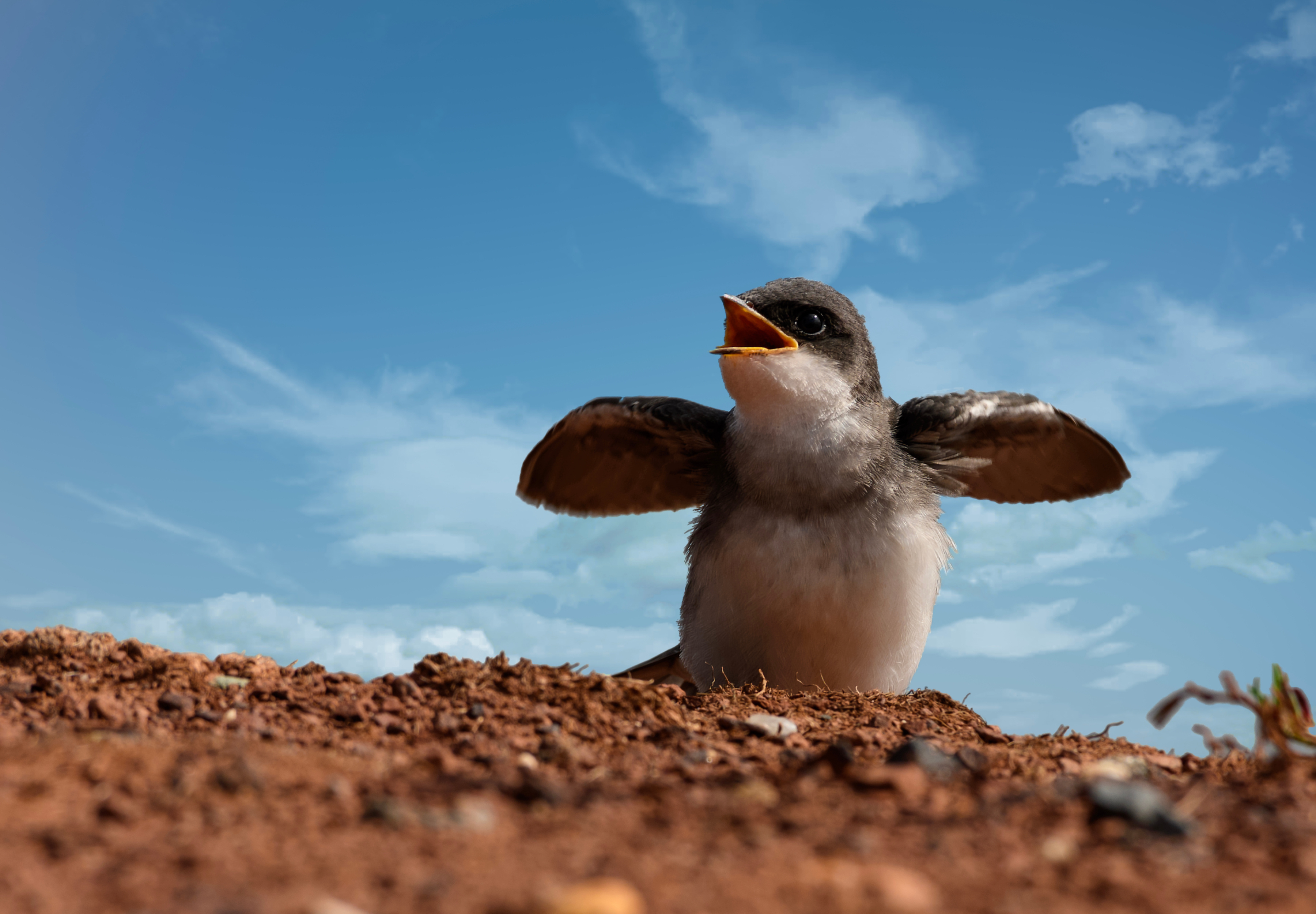

Our new paper published in Nature Communications provides a blueprint for conserving enough habitat to protect the populations of almost one-third of the warblers, orioles, tanagers, and other birds that migrate among the Americas throughout the year. This analysis represents first time a global citizen-science database (eBird) has been used in hemispheric-scale conservation planning. How did I get here?
I used to be a software developer. After vocational school in electrical engineering, I created games for casino machines using C++. The industry was crocked, but I loved coding. I got to a point that I could not recall what happened during my 30-minute commute to and from work. All I did was think about the code.
This preoccupation didn’t seem healthy to me, so I decided to change careers. My love of the outdoors drew me to Biology, and later that year I started university for the first time, studying animal behavior and ecology.
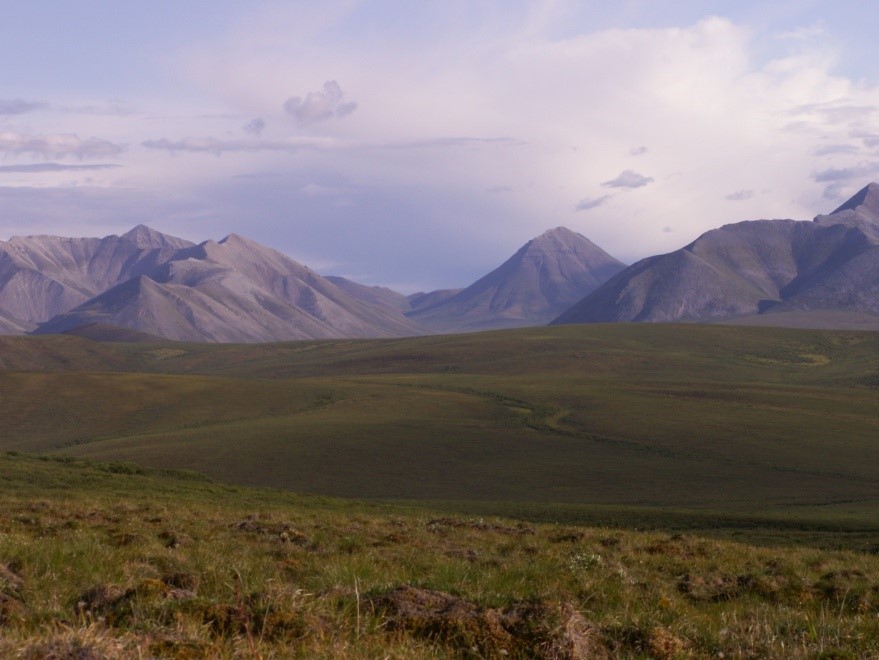


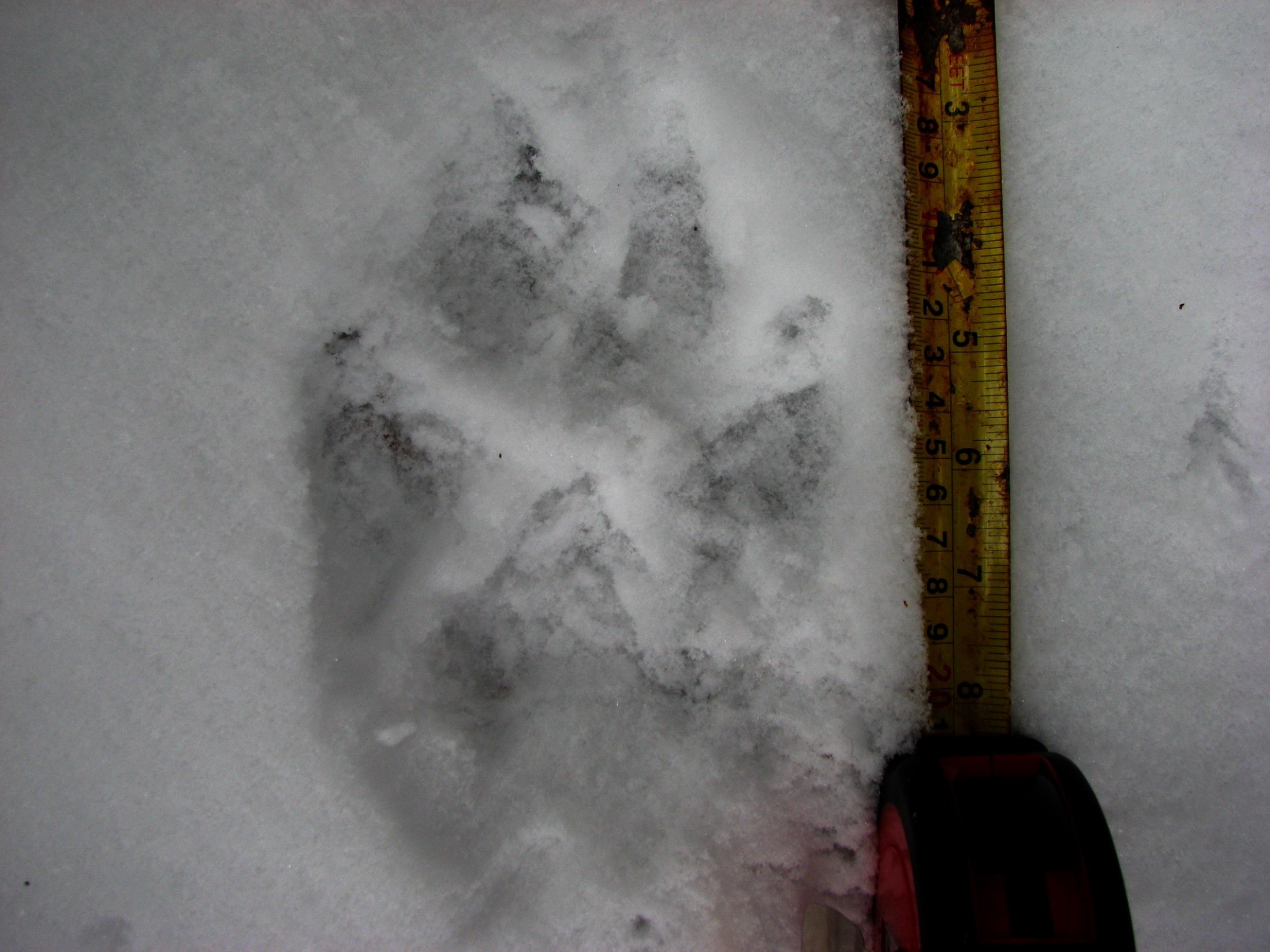
Ecology really seemed to be a good fit, as I was able to spend time outdoors, collect data and admittedly also have a lot of fun, just running through the snow chasing animal tracks (always against their direction of travel, as I didn’t want to run into cougars or wolves).


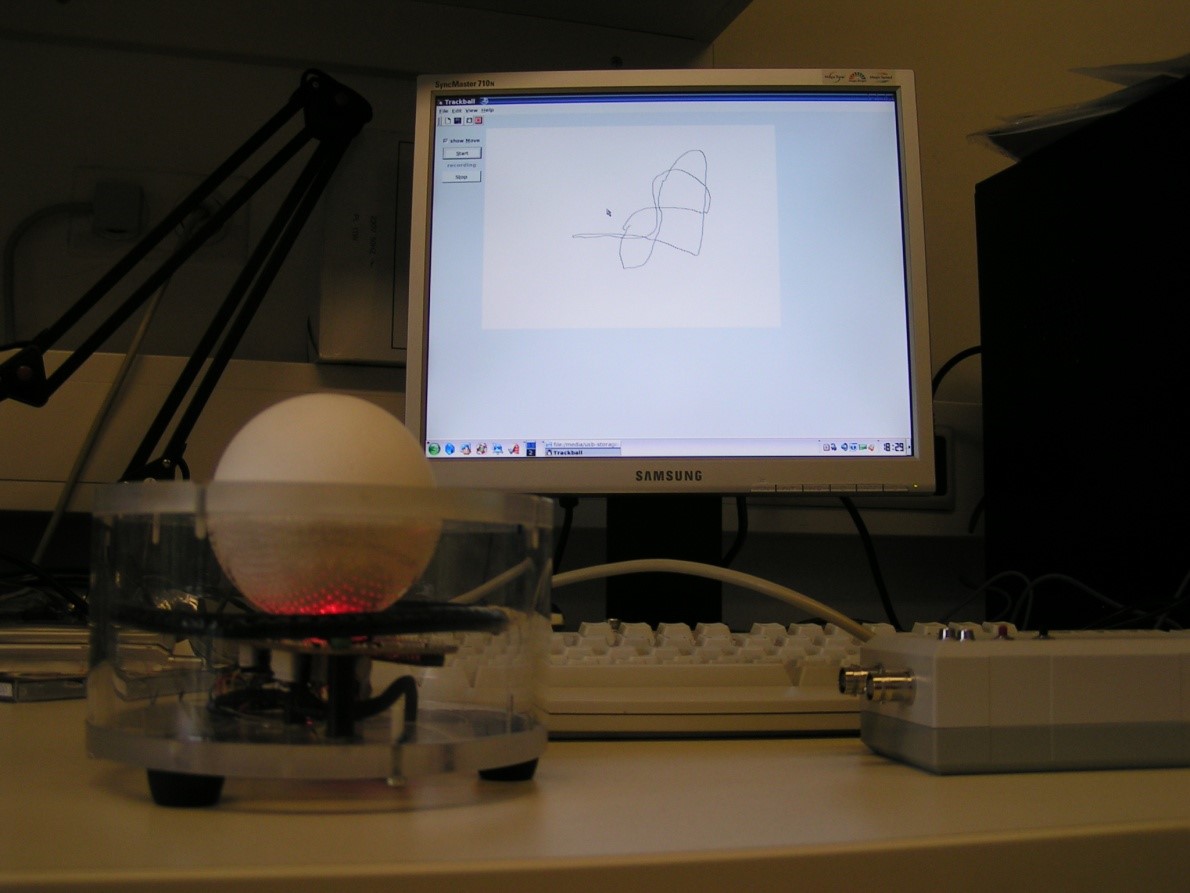
But of course, my past caught up with me. For one of my undergrad projects, I started to code again. I developed the graphical user interface for a cricket movement tracking device. I also really enjoyed digging up the Lego blocks I used in my childhood to create a low-cost calibration gadget for the device.
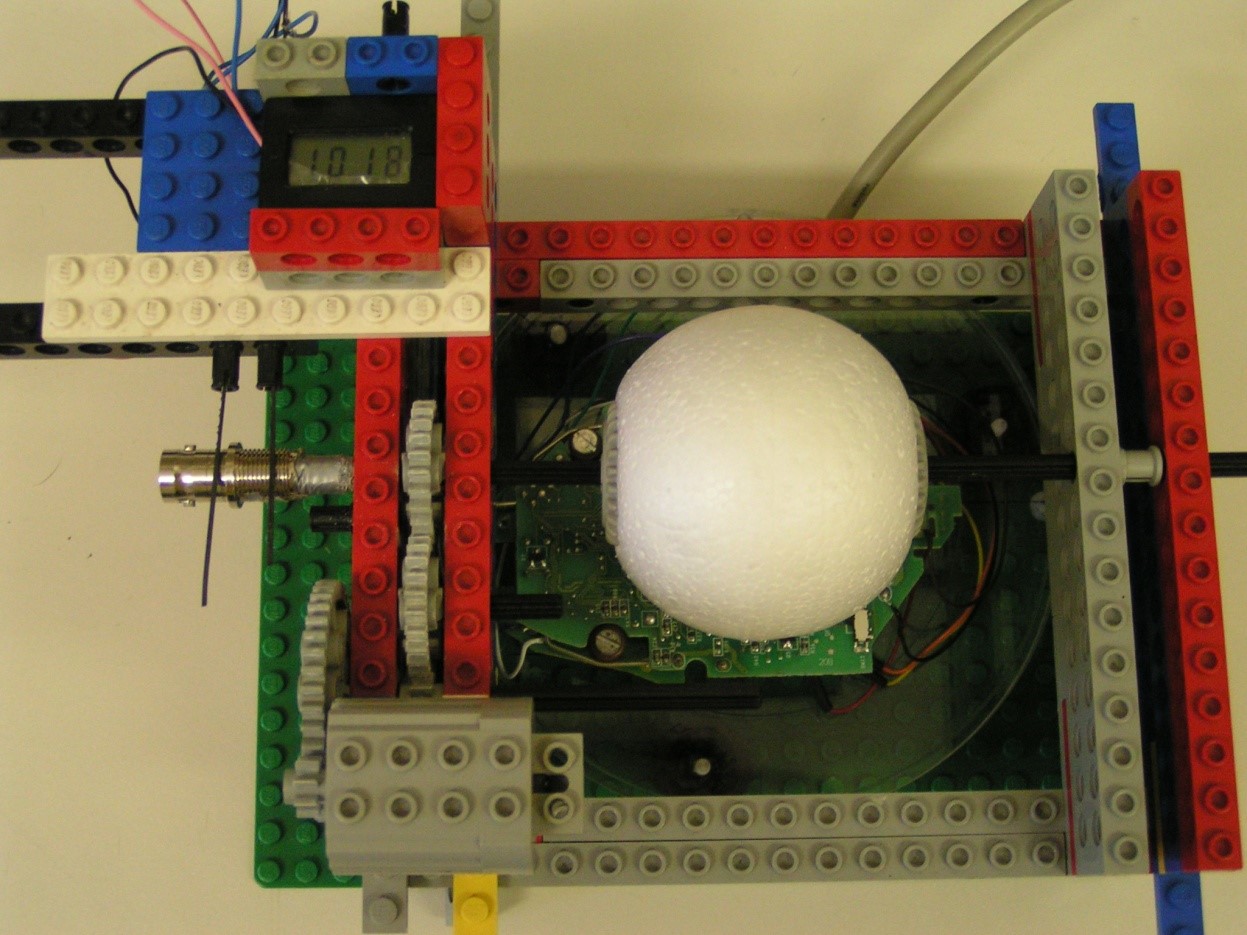
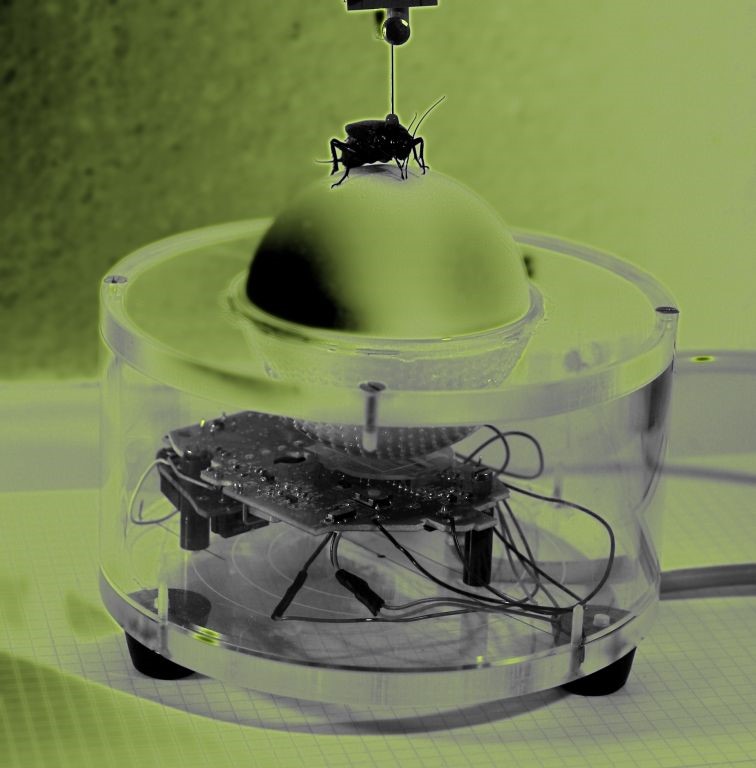



I really got back into coding when I started my PhD and learned to use R. I eventually started developing analytical tools for researchers and practitioners to use in conservation planning and outreach, ranging from data visualization to complex decision support tools.
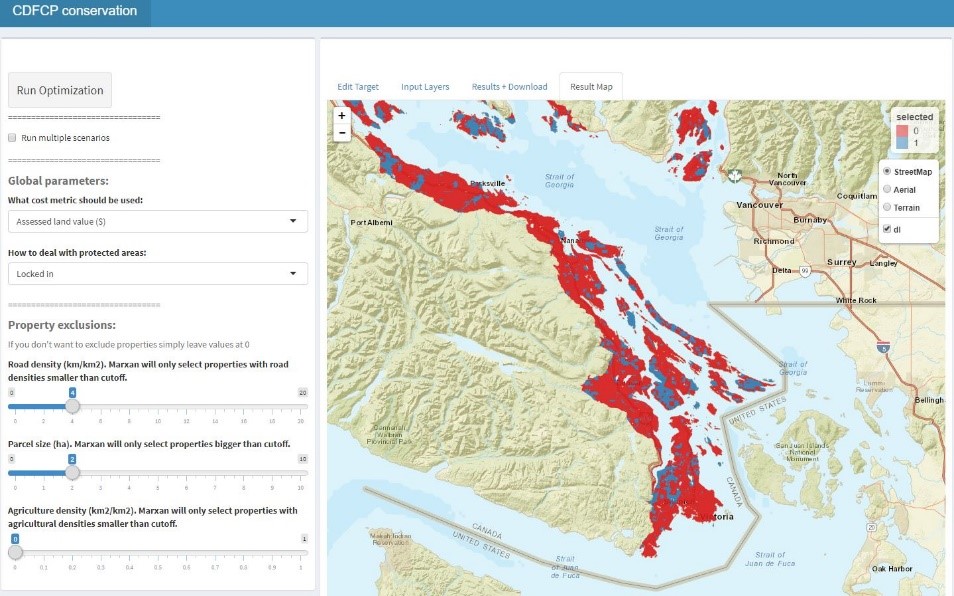
Enter the Liber Ero fellowship program, and the outstanding group of mentors and collaborators on the project that lead to this paper.

I have been very fortunate to have mentors and collaborators like Peter Arcese, Joe Bennett, Amanda Rodewald and Scott Wilson. Amanda was my ticket into the outstanding Cornell Lab of Ornithology's (CLO) global citizen science database eBird. Birders can download a free mobile app onto their smartphone or tablet and enter information about the birds they observe from anywhere around the world. Over 420,000 eBirder’s have entered more than half a billion sightings in the 15 years since eBird was created.


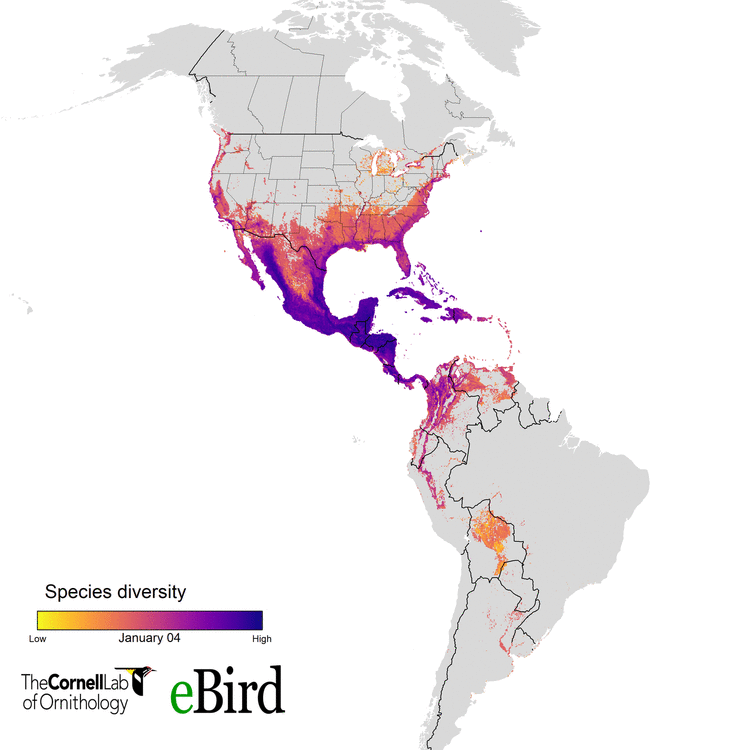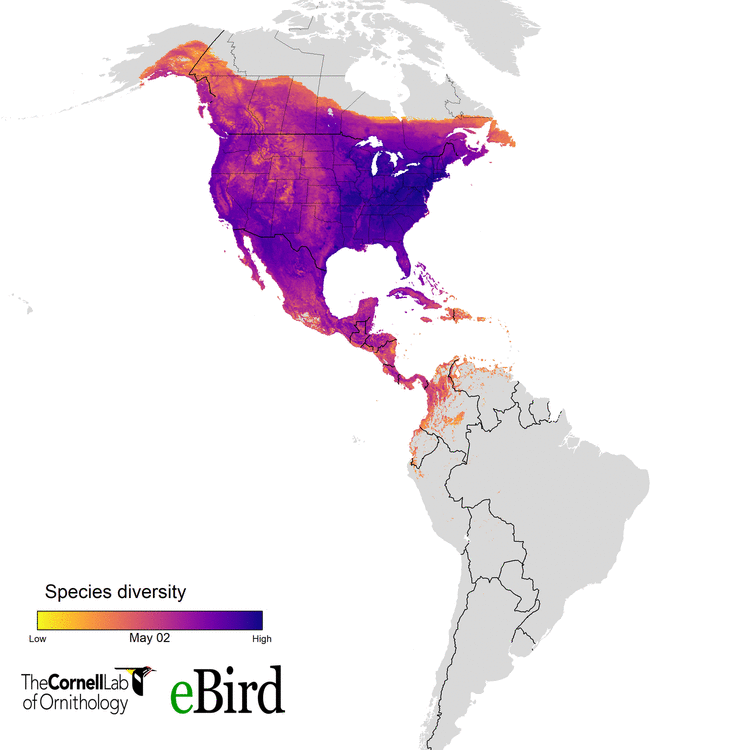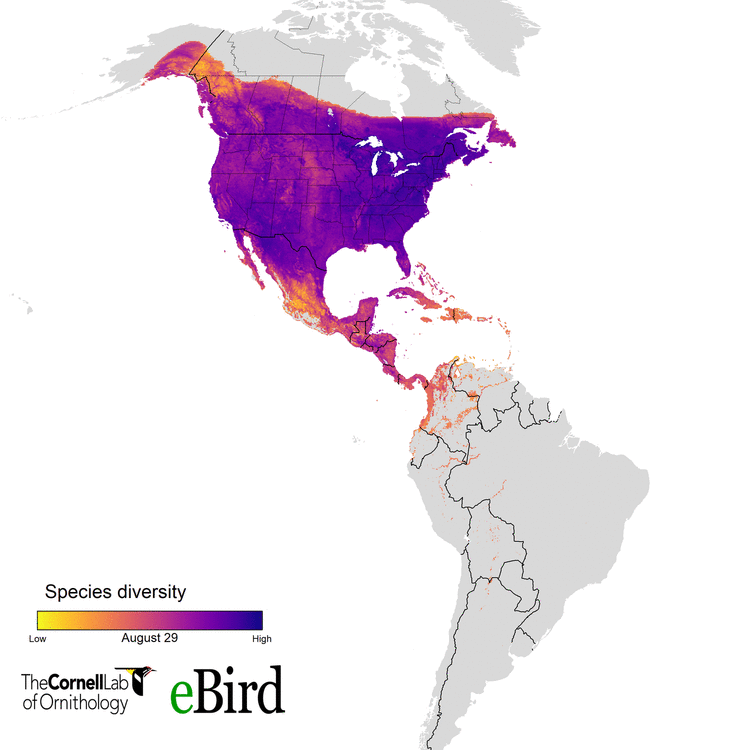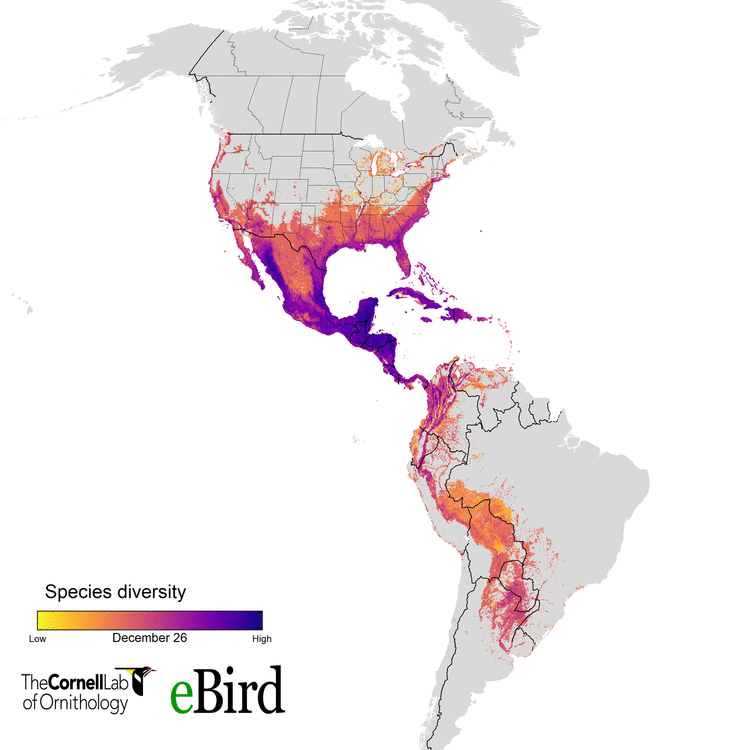
For the current study, we (specifically data wizards Daniel Fink and Tom Auer, from the CLO) used 14 million datapoints collected by hundreds of thousands of citizen scientists to create weekly abundance maps of 117 migratory species. With these massive datasets, we went into a systematic conservation prioritization R package called ‘prioritizr’. The lead developer of that package is the amazing Jeff Hanson. We calculated how to sufficiently conserve habitat across the Western Hemisphere for all the habitats 117 migratory species use, throughout their annual cycle of breeding, migration, and overwintering. This work provides guidance on the location and amounts of lands that must be protected for 30 percent of the populations of these birds.
eBird and more than 420,000 eBirders around the world make it possible for researchers like us to consider the full annual journey of migratory birds - information that can translate to better conservation. With the help of current and future eBirders, we are hopeful that we will be able to make informed and cost-effective decisions to protect migratory species.
Although our planning scenarios focused on migratory birds, our approach could be easily adjusted and replicated in other migratory species and systems with sufficient data. In the case of birds, citizen science data and advanced prioritization tools allowed us to reveal marked efficiencies in area-based plans spanning the full annual cycle and multiple jurisdictions to conserve 117 individual species simultaneously.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in