Malaria genomic surveillance using nanopore sequencing: Shifting the focus into endemic countries
Published in Genetics & Genomics
Introduction
Sophia Girgis & William Hamilton - Wellcome Sanger Institute, UK
Malaria remains a global health priority with an estimated death toll of over 600,000 people per year, mainly affecting young children in tropical low- and middle- income countries. Malaria is a mosquito-borne infectious disease caused by Plasmodium parasites, with P. falciparum responsible for the majority of deaths. The repeated evolution and spread of resistance to key antimalarial drugs has thwarted efforts to eliminate malaria over the last 70 years. Partial resistance to artemisinins - a cornerstone of current treatments for P. falciparum malaria - was first described in southeast Asia and has now been reported in multiple countries in eastern Africa. The expansion of molecular surveillance capacity in Africa is critical, to monitor emerging drug and diagnostic test resistance and the deployment of new malaria vaccines.
The aim of this study was to design and implement an end-to-end molecular surveillance workflow using a handheld MinION sequencing device from Oxford Nanopore Technologies (ONT) and a portable laptop computer, suitable for deployment in malaria endemic settings outside of large genomic centres. A key motivation was to drive the decentralisation and ‘democratization’ of genomics.
The study was based at two sites in Ghana, west Africa, with contrasting epidemiology: the capital, Accra, and Navrongo, a rural town located in northern Ghana. The assay used a multiplex PCR targeting five key drug resistance markers and the leading vaccine and monoclonal antibody target, circumsporozoite protein (csp). The protocol could be implemented on readily collected samples from people infected with malaria parasites, such as dried blood spots derived from fingerpricks. The data analysis pipeline could run directly on a high-specification laptop in around 20 minutes, and is available on GitHub: https://github.com/sanger-pathogens/nano-rave.
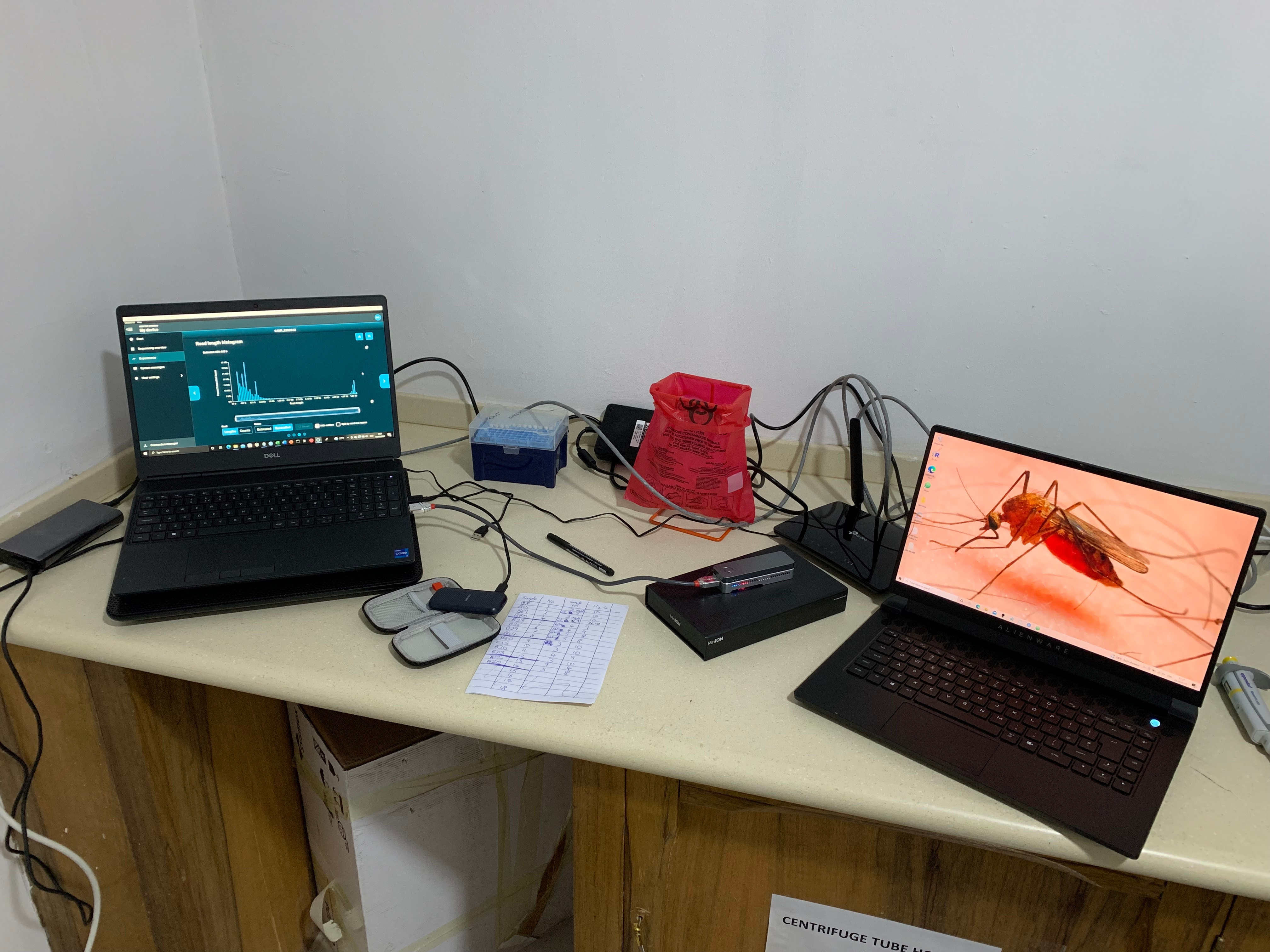

The study was conceptualised and undertaken in partnership between the Wellcome Sanger Institute (United Kingdom), University of Ghana, and the Navrongo Health Research Centre (NHRC). Below are some of the personal perspectives from the scientists based in Accra and Navrongo.
Sequencing in Accra
Edem Adika - University of Ghana, Ghana
The malaria transmission landscape in Ghana is divided into three zones: the northern and middle belts that experience relatively high malaria transmission and the southern belt that has lower malaria transmission. Although Accra, on the coast, has lower transmission, malaria is perennial here, fuelled by a longer two-part rainy season that spans between April and October.
For this study, we collected blood samples from patients at the Ledzokuku-Krowor Municipal Assembly (LEKMA) hospital in eastern Accra, which offers all the services of a standard hospital and also has a dedicated malaria laboratory, making it well-suited for malaria surveys.
It was fantastic to use the MinION device to rapidly generate useful data and to demonstrate the feasibility of this approach to support malaria surveillance across Ghana. For example, the assay could rapidly identify resistance markers to sulfadoxine and pyremethamine (SP), which are used for preventive treatment in children and pregnancy. We did not find the ‘high-level’ resistance markers associated with failure of these public health interventions.
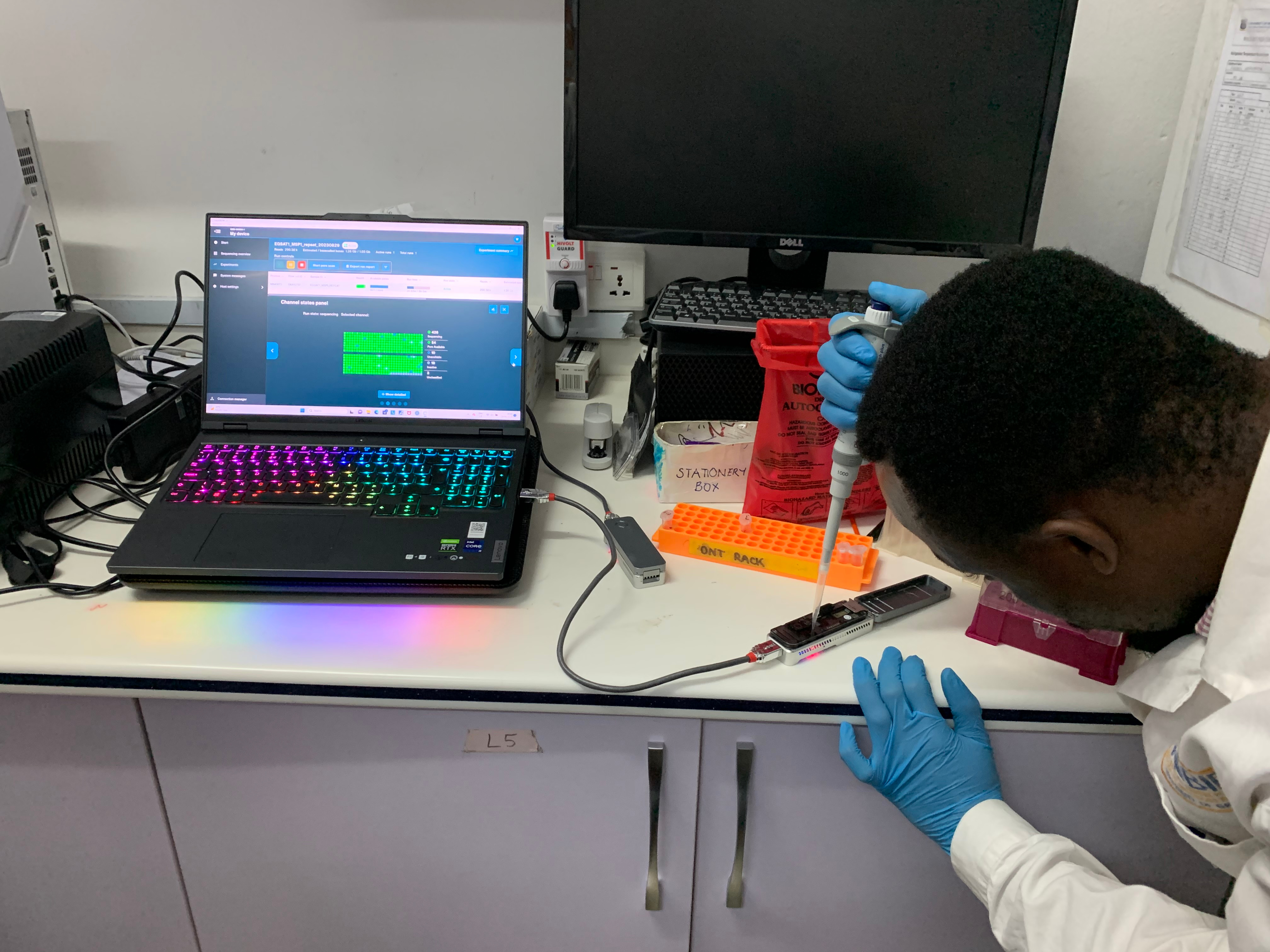

Edem Adika (co-lead author for the study), loading a MinION flow cell in the West African Centre for Cell Biology of Infectious Pathogens (WACCBIP), University of Ghana, Accra, Ghana. Image credit: William Hamilton.
The learning experience made the endeavor even more exciting for me personally - despite the 18 hour car journey from Accra to Navrongo! I hope that using ONT for malaria surveillance will soon be achieved at scale in resource-limited settings, generating data within endemic countries to support malaria control efforts. In addition, MinION sequencing could be expanded to incorporate other infectious diseases such as TB, HIV and hepatitis B virus, which are also extremely important threats to health in Ghana.
Sequencing in Navrongo
Felix Nenyewodey & Dodzi Senoo Jnr - Navrongo Health Research Centre, Ghana
Despite several high impact interventions - including indoor residual spraying, distribution of dual insecticide treated bednets, and the use of artemisinin-based combination therapies (ACTs) for treatment - malaria mortality remains high among the people living in the Navrongo Health and Demographic Surveillance (NDHSS) area in Navrongo, a rural setting in the Upper East Region of Ghana. The risk is especially high for young children and pregnant mothers.
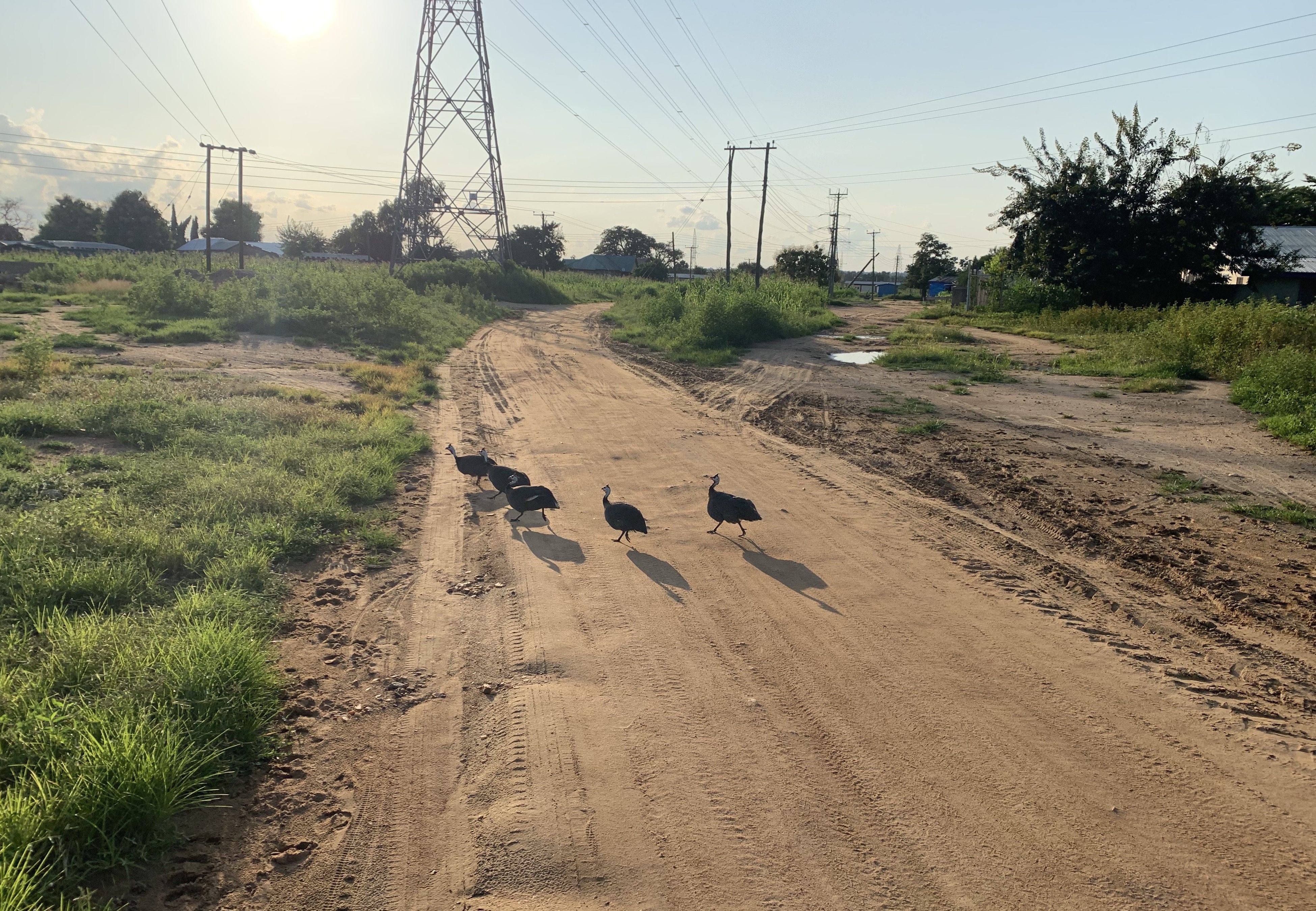

Malaria transmission here is divided into two major seasons: the rainy season (lasting 4 months, approximately July - October) and a dry season (November - June). The rainy season is a breeding time for mosquitoes as there are abundant water sources. This leads to a high incidence of malaria in communities compared to the dry season, which has much lower incidence. Socioeconomic deprivation and a lack of availability of diagnostic equipment in some areas makes controlling malaria even more challenging.
In this context, nanopore sequencing P. falciparum in Navrongo from patients with malaria and genotyping variants that cause drug resistance - within two days of collecting the samples - felt like being in a sci-fi movie. This portable device (the MinION) opened our minds to new scientific possibilities; dreams of studying genomics for malaria and other important pathogens now seem more achievable and closer to becoming reality. We also believe that ONT could make cancer genomics more attainable to those of us based in developing countries.
Overall, it was really a great experience doing MinION sequencing in Navrongo; knowledge from the study will continue to inspire us in our future careers.
Conclusions
Lucas Amenga-Etego - University of Ghana, Ghana
Nanopore sequencing undoubtedly has a niche in the pathogen genomic surveillance ecosystem. Its portability and comparatively lower costs put it forward as a platform for addressing the sequencing gap in sub-Saharan Africa. Although, several challenges remain to ensure this potential is fully realised, such as ensuring a reliable and affordable supply of reagents on the continent, and developing scalable data analysis pipelines. We would also like to push for rapid pathogen whole genome sequencing, in addition to the targeted amplicon-based approach used in this study.
Notwithstanding the progress made in establishing sequencing capacity in sub-Saharan Africa and other malaria endemic regions during the COVID-19 pandemic, there remains an important gap in the breadth of sequencing across malaria endemic regions. In order to address this gap, there is a need for upscaling sequencing capacity development in endemic countries to bridge the Global North and South sequencing divide - including bioinformatics and data analytics infrastructure. This should be a key goal for the global pathogen genomics community going forward.
Edited by William Hamilton and Lucas Amenga-Etego.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in