Merging Enzymes with Chemocatalysis for Amide Bond Synthesis
Published in Chemistry
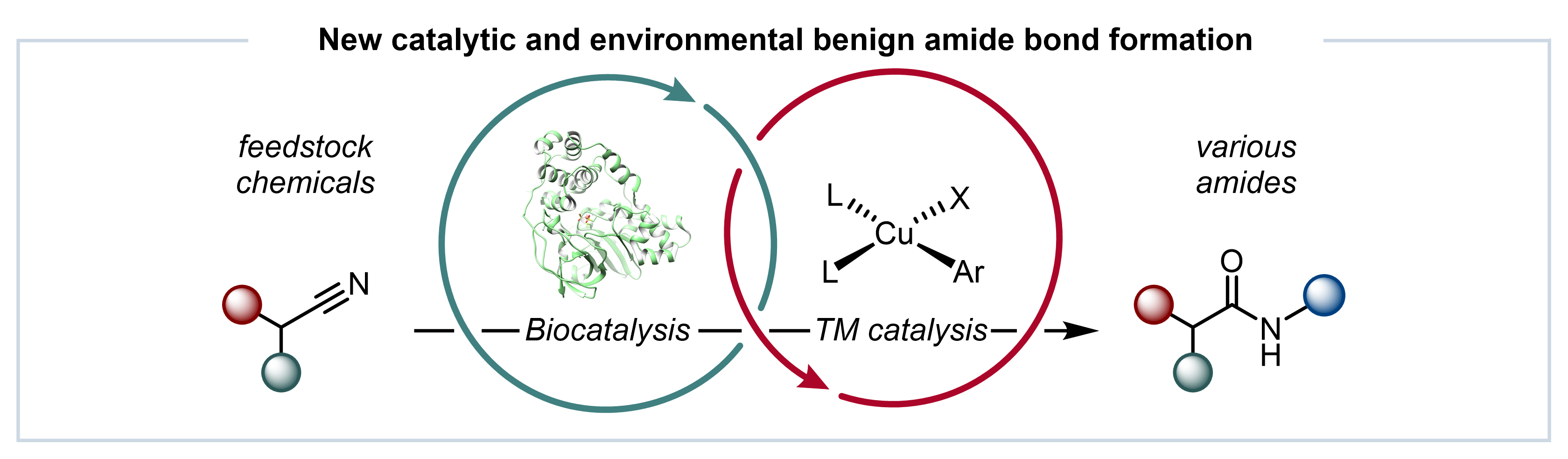
Amide bonds are ubiquitous structural features in nature and are of fundamental importance, as amides are the bonds that link amino acids to form proteins. In addition, many essential chemicals and functional materials are also produced using amide bond synthesis at some stage during their manufacture. Traditionally, amide bonds have been constructed by the coupling of amines and carboxylic acids. Amide bond coupling is very robust and is the most frequently used reaction in the pharmaceutical industry. However, amide coupling methodologies consist of multiple steps, each requiring large amounts of expensive and harmful chemicals, creating significant problems in purification and waste processing. Thus, benign catalytic methods with improved efficiency are needed to produce amides more sustainably.
The combination of chemocatalysis and biocatalysis in the same reaction (integrated catalysis) is a relatively young but rapidly evolving research field. A key aspect is, that integrated catalysis can improve the overall synthetic efficiency through decreasing the number of reaction steps and thereby reducing solvent consumption and waste production. As water is the favoured solvent of enzymes, integrated processes in aqueous media are also particularly desirable from a green chemistry perspective. However, combining enzymes and chemocatalysts in the same vessel is often not straightforward. The different operating conditions and mutual deactivation of the catalysts present the most challenging obstacles in the development of integrated processes.
Our research group has a major interest in developing novel and more sustainable approaches using biocatalysts to produce amide bond containing molecules. We have also demonstrated how enzymes and chemocatalysts can be combined to develop new telescoping routes to important molecules, including pharmaceuticals. Based on these results, we aimed to address the challenges associated with amide bond synthesis by developing an integrated chemo- and biocatalytic approach. In this study, we report an integrated system, merging nitrile hydratase (NHase) enzymes with a Cu-catalysed N-arylation reaction in a single reaction vessel for the construction of amide bonds (Fig. 1A).
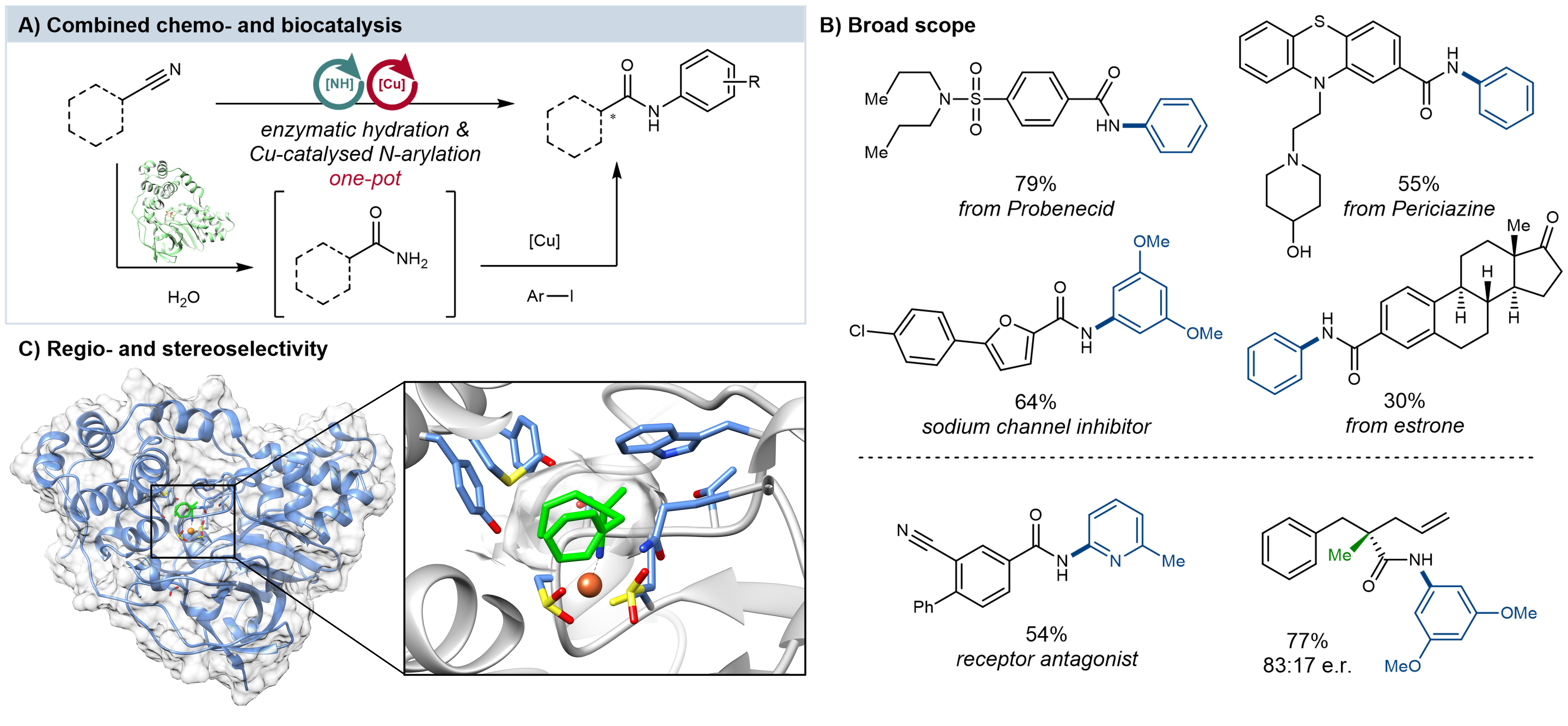
Figure 1: Integrated amide bond synthesis by combining enzymes and chemocatalysts. A) General concept for the integrated amide synthesis. B) Selected examples of the scope. C) NHase structure accommodating nitrile substrate. Regio- and stereoselectivity was applied for the selective synthesis of bioactive compounds and chiral amides.
We aimed to develop a general process that allows the synthesis of various amides at a preparative scale, while avoiding both the need for protecting groups and the use of harmful organic solvents, while utilising an earth-abundant, inexpensive and non-toxic transition metal-catalyst. We found that E. coli whole cells, overexpressing NHase enzymes from different origins, could be efficiently combined with Cu-catalysed Ullmann-type coupling in the same reaction vessel without any mutual deactivation (Fig. 1A). This one-pot cascade displays excellent functional group tolerance and broad substrate scope with >50 examples, including one-pot synthesis of bioactive natural products and drug molecules (Fig. 1B). Usage of NHase enzymes enabled us to synthesise chiral amides and the selective hydration of polynitriles by taking advantage of the biocatalyst’s inherent stereo- and regioselectivity (Fig. 1C). We also demonstrate the scalability of the integrated reactions, using micellar organo-compartments to drastically increase the efficiency of the reaction at very high substrate concentrations, beyond the scope of biocatalysis. Our findings provide an alternative approach, through the integration of chemo- and biocatalysis, delivering functionalised amide products, under mild and benign conditions, for a range of important future applications.
This research has now been published in Nature Communications.
https://doi.org/10.1038/s41467-022-28005-4
Luis Bering, Elliott J. Craven, Stanley A. Sowerby Thomas, Sarah A. Shepherd & Jason Micklefield
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in