New research advances the understanding of microbial photosynthesis
Published in Microbiology

The capture of the sun’s energy through photosynthesis in distinct photosynthetic organisms is the fundamentals of almost all life on Earth. Under the current circumstances of global climate change and food/energy crisis, there has been considerable interest in studying how photosynthetic microbes and plants trap and convert solar energy in nature.
Photosynthesis usually starts with photon absorption by light-harvesting antenna. The absorbed energy is then rapidly and efficiently transferred to a reaction centre (RC) for charge separation. Purple bacteria are the oldest photosynthetic species and carry out anaerobic photosynthesis in diverse ecological niches. In purple bacteria, light-harvesting 1 (LH1) complexes often associate with the RC to form a RC-LH1 core supercomplex, containing a high content of non-covalently attached pigments such as bacterial chlorophylls and carotenoids, to increase the effective cross-section for light absorption of each RC (Mullineaux and Liu, 2020). This then supports photosynthesis to operate effectively over a wide range of environmental light intensities and qualities. For example, purple bacteria are highly efficient in utilising green and far-red light of wavelengths above 750 nm that plants and algae do not utilize.
Due to the simplicity of the photosynthetic machinery and ease of growth, purple photosynthetic bacteria have been used to study the fundamental mechanisms of photosynthesis. For decades, X-ray crystallography has been used by structural biologists to solve the structural composition of protein complexes. The Nobel Prize in Chemistry 1988 was awarded to Hartmut Michel, Johann Deisenhofer, and Robert Huber for their contributions to determining the photosynthetic reaction centre from a purple bacterium. Over the past few years, the cutting-edge cryo-electron microscopy (cryo-EM) technology (2017 Nobel Prize in Chemistry) has revolutionised the structural understanding of photosynthetic RC-LH1 complexes in a range of purple bacteria and has highlighted the variety of RC-LH1 complex structures among different species.
 The research in the Liu lab at the University of Liverpool has recently focused on special “open” RC-LH1 structures from the Rhodobacter species, which contain the gate(s) within the LH1 rings. For example, cryo-EM studies showed that the RC–LH1 structure from Rhodobacter veldkampii is a monomer and has a “C-shaped” LH1 ring with a large opening, unlike the closed LH1 ring found in many purple bacteria. The in-depth structural analysis further discovered that this opening is formed by an extra protein peptide, known as PufX, with a special orientation (Bracun et al., 2021). The study indicated that PufX works as a molecular “cross brace” to bridge the RC and LH1 and stabilise the RC–LH1 core structure. It also provides a big channel for the electron carrier molecules called quinones to diffuse across the LH1 barrier for efficient electron transport.
The research in the Liu lab at the University of Liverpool has recently focused on special “open” RC-LH1 structures from the Rhodobacter species, which contain the gate(s) within the LH1 rings. For example, cryo-EM studies showed that the RC–LH1 structure from Rhodobacter veldkampii is a monomer and has a “C-shaped” LH1 ring with a large opening, unlike the closed LH1 ring found in many purple bacteria. The in-depth structural analysis further discovered that this opening is formed by an extra protein peptide, known as PufX, with a special orientation (Bracun et al., 2021). The study indicated that PufX works as a molecular “cross brace” to bridge the RC and LH1 and stabilise the RC–LH1 core structure. It also provides a big channel for the electron carrier molecules called quinones to diffuse across the LH1 barrier for efficient electron transport.
Recently, the group has further studied the RC–LH1 core complexes from Rhodobacter sphaeroides using cryo-EM (Cao et al., 2022). In native Rhodobacter sphaeroides cells, RC–LH1 complexes exist as monomers and dimers. The dimeric core has a “S-shaped” LH1 ring with two large openings that encircle two RCs, representing the largest protein complexes in bacterial photosynthesis. High-resolution cryo-EM imaging deciphered that this opening is formed by two extra protein peptides, termed PufX and PufY. By genetically “deleting” these peptides, the researchers obtained several structural variants of the RC–LH1 complexes. Analysis of these structures validated that PufX is responsible for the interactions between two monomers to form the dimer, and the PufY peptide is important to keep the “gate” in the LH1 ring.
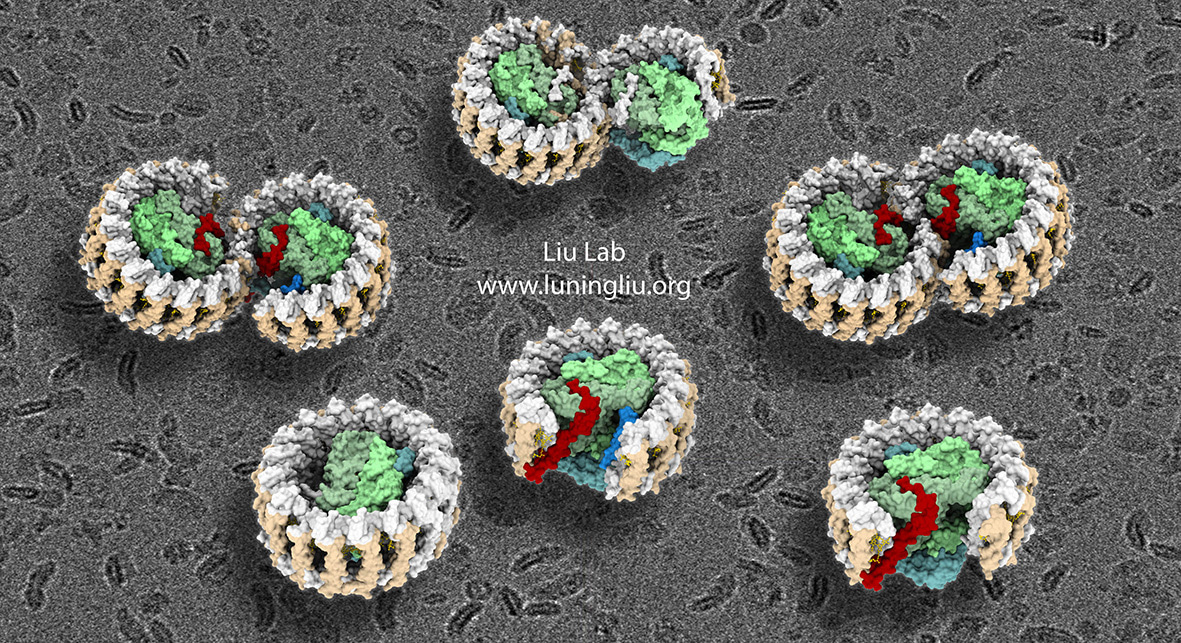
Through systematic studies of the dimeric structure and its variants, the researchers proposed how the dimeric photosynthetic core complex is generated and self-organised in nature. The researchers further used computational approaches to prove that the “gates” in the “S-shaped” LH1 ring provide the channels for the electron carrier molecules to diffuse across the LH1 barrier to perform efficient electron transport.
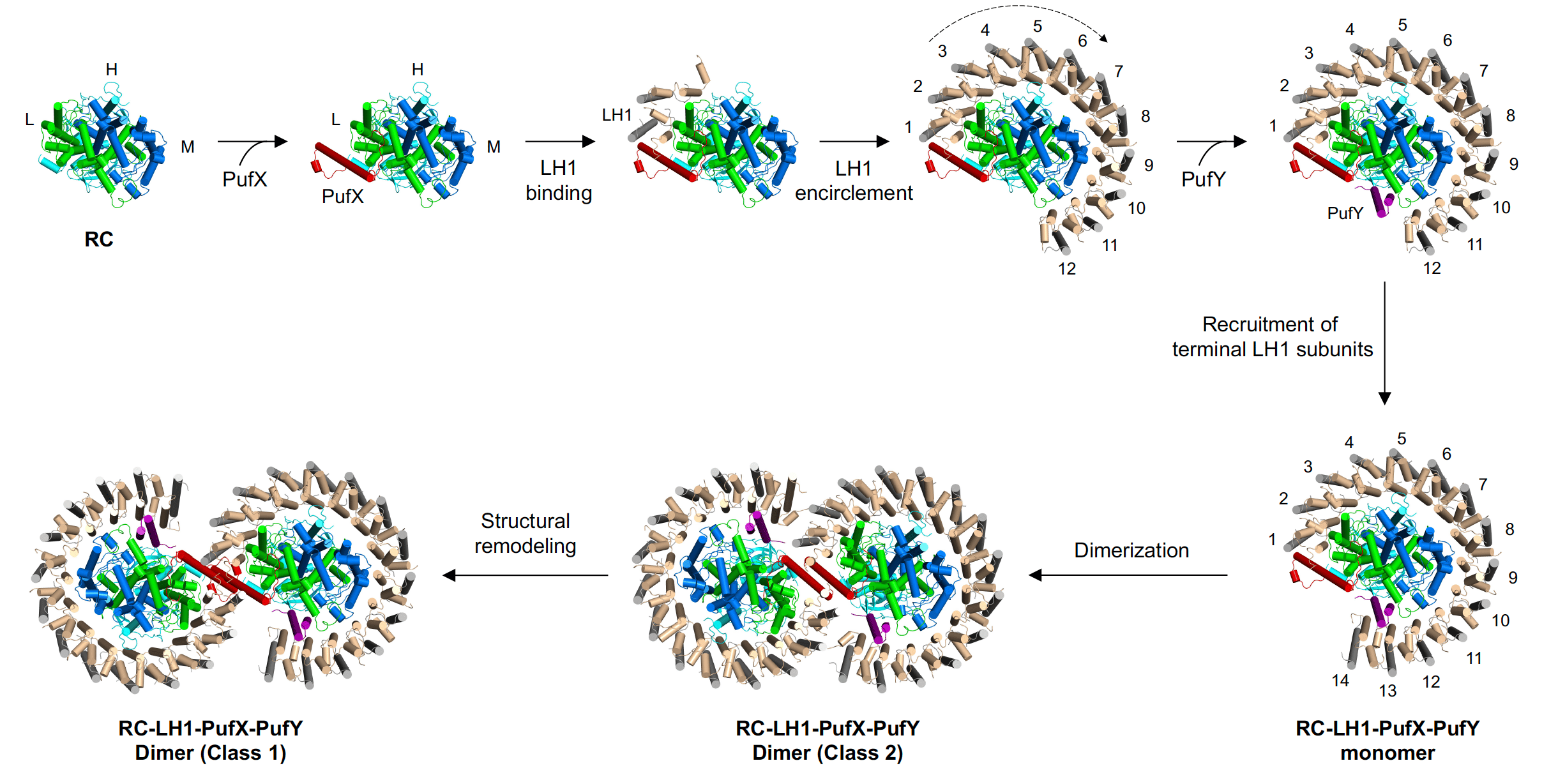
Possible assembly pathway of the dimeric RC–LH1 core complex
“This is the first time to study these important photosynthetic protein structures in a systematic manner,” said Professor Luning Liu at the University of Liverpool. “It is truly remarkable to unravel how native photosynthetic systems are elegantly generated and fulfil functions in cells, as we need to understand, at the molecular level, how photosynthesis is performed and regulated to allow photosynthesizing organisms to grow in their specific niche and how to make our planet colourful and sustainable. The fundamental finding is valuable for artificial photosynthesis engineering and bioenergy production.”
References:
Cao, P., Bracun, L., Yamagata, A., Christianson, B.M., Negami, T., Zou, B., Terada, T., Canniffe, D.P., Shirouzu, M., Li, M., et al. (2022). Structural basis for the assembly and quinone transport mechanisms of the dimeric photosynthetic RC–LH1 supercomplex. Nature Communications 13, 1977.
Bracun, L., Yamagata, A., Christianson, B.M., Terada, T., Canniffe, D.P., Schirouzu, M., and Liu, L.-N. (2021). Cryo-EM structure of the photosynthetic RC-LH1-PufX supercomplex at 2.8-Å resolution. Science Advances 7, eabf8864.
Mullineaux, C.W., and Liu, L.N. (2020). Membrane dynamics in phototrophic bacteria. Annual Review of Microbiology 74, 633-654.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in