See-N-Seq: RNA sequencing of target single cells identified by microscopy via micropatterning of hydrogel porosity
Published in Bioengineering & Biotechnology
Microscopes have long delighted us by giving us an opportunity to peer into the private lives of living cells. We have watched their intricate interactions and classified them based on their diverse and intriguing characteristics. These observations inspired methods such as single cell sequencing that revealed how variations in gene expression give rise to this diversity of features. However, despite the tremendous advances made in both microscopy and sequencing methods, there is still no way to simply select a cell using the microscope for subsequent gene expression analysis.
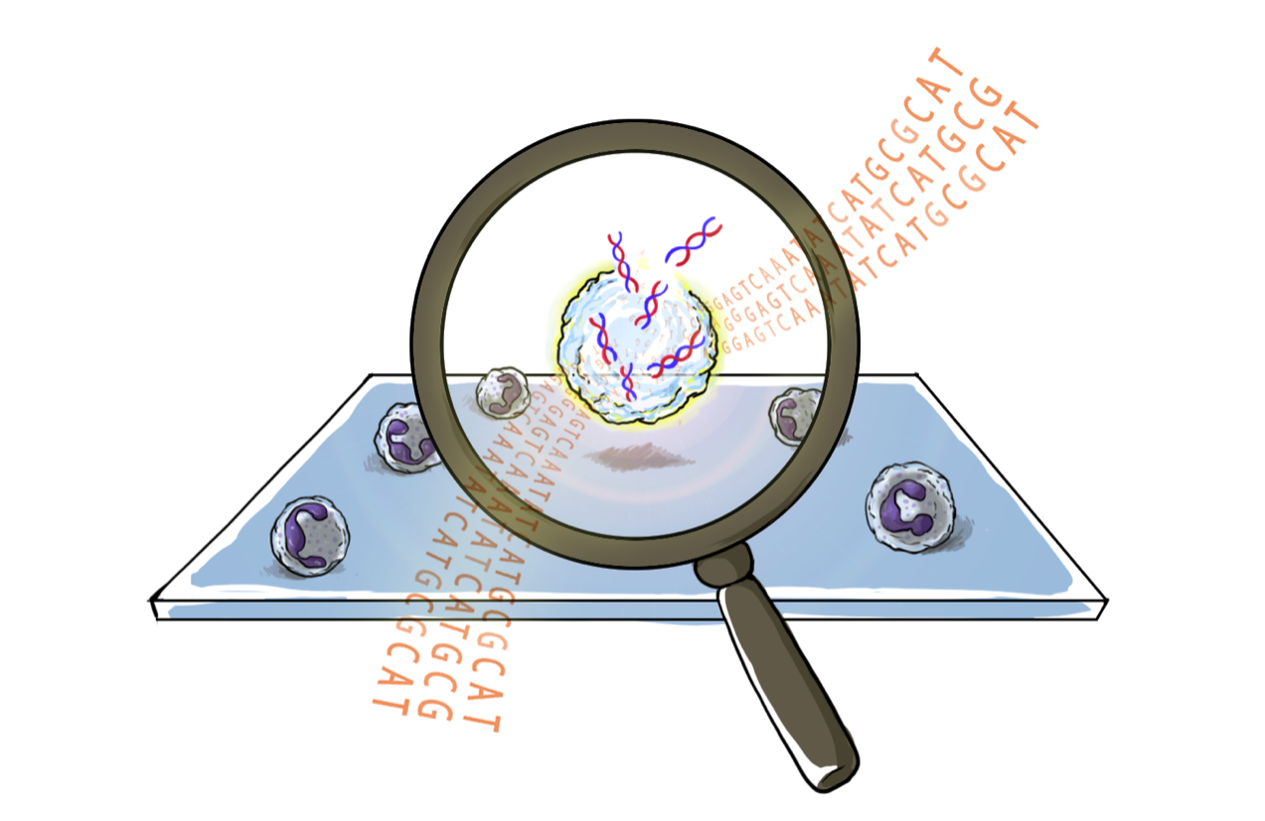
Our research team has been driven by the challenge of separating cells for gene expression profiling. We have previously addressed the needle-in-a-haystack challenge of finding relevant cells using physical and antigenic biomarkers for cell separation. However, these constitute only an indirect means of accessing the target cells. We wondered, what if we could directly sequence the cells that we can see using a microscope? If we could categorize, or if we could understand the “software” shaping cellular decision-making, then we could figure out how to manipulate these systems in health and disease.
Current methods that were available for this challenge of image-based cell selection involve methods for physically transferring cells to a new receptacle, such as micropipette aspiration, laser capture microdissection or magnetic micro-raft. Physically transferring single cells is technically challenging, time-consuming and prone to cell loss. To overcome these limitations, we envisioned the See-N-Seq technology, in which the microscope becomes a cell-sorting device where mRNA can be isolated directly from cells without requiring the transportation of single cells into different containers.
The underlying principle of See-N-Seq involves the use of two different photopolymerizable hydrogels. The first hydrogel is porous and permeable to biomolecules, such as staining reagents and nucleic acids. This first hydrogel simply holds cells in place so that they do not move during imaging and cell selection. A second hydrogel is then infused, which is impermeable to biomolecules. When a laser polymerizes this second hydrogel, it is used to encapsulate non-target cells in an impermeable matrix. Once the laser has encapsulated all non-target cells, the addition of cell lysis reagent induces mRNA release from only the exposed target cells, selected by microscopy. Consequently, a researcher can observe a cell population under the microscope, select a cell with an interesting phenotype and perform mRNA capture directly within the imaging microwell.
This method is valuable for sequencing cells that are “caught in the act” of infrequent and transient cellular events, which provides a key window into the cellular decision-making associated with these specific events. Examples of transient cellular events include receptor-ligand interactions, cell-to-cell contact, and micro-environmental modifications. Also, this approach is particularly useful when the cells can only be identified by imaging, such as investigation of protein localization.
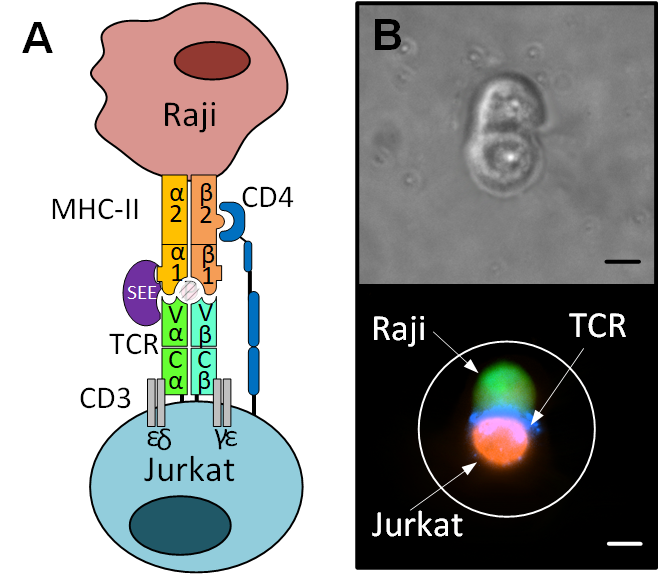
The particular cellular event that we investigated with See-N-Seq is the immunological synapse. In an immune response, phagocytic cells consume pathogens and other foreign material. Some of these phagocytes also present the foreign molecules on their surface, in a process called “antigen presentation”. When these antigen presenting cells come into physical contact with Helper T-cells, they form a joining of two cells, called the immunological synapse. Helper T-cells play a pivotal role in deciding the immunological response to infection and the immunological synapse is thought to play an important part in this decision process. Specifically, the helper T-cells may differentiate into type-1 cells (Th1) that promote a cellular immune response or they may become type-2 cells (Th2) that promote an antibody response.
We used cell lines that represent antigen presenting cells (Raji cells) and helper T-cells (Jurkat cells) in order to investigate relatively homogenous cell populations. We used See-N-Seq microscopy based selection to only sequence the cells with visually confirmed immunological synapse, and then we used the gene expression profile of these cells to determine whether the outcome of this synapse was to produce Th1-like or Th2-like cells. Surprisingly, even though all cells were provided the same stimulus, some cells polarized toward Th1 cells while others resembled Th2 cells. These findings are remarkable because they suggest that cells forming an immunological synapse in response to the same activating conditions can undergo distinct decision-making processes that result in divergent transcriptional programs.
See-N-Seq provides a simple and effective method to isolate RNA from specific single cells identified by imaging without needing to physically transport single cells. The ability to associate context with transcriptomic snapshots of single cells could be employed to characterize T-cells in other situations, such as thymic education, exhaustion due to chronic cytokine stimulation, and tumor-mediated reprogramming. In each of these contexts, researchers will have the opportunity to transition from profiling the transcriptional landscape to understanding how gene expression drives cellular decision-making. We anticipate that See-N-Seq has the potential to uncover decision-making processes in critical cells that ultimately dictate health and disease outcomes.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
From RNA Detection to Molecular Mechanisms
Publishing Model: Open Access
Deadline: May 05, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in