Targeting fungal membrane homeostasis with imidazopyrazoindoles impairs azole resistance and biofilm formation
Published in Microbiology

Candida species represent a leading cause of these infections, with the opportunistic pathogen Candida albicans reigning as the fourth most common cause of hospital-borne infection (Brown, et al., 2012). This growing threat has been met with the prophylactic overuse of a limited arsenal of antifungals, leading to the rapid emergence of multidrug-resistant pathogenic fungi (Revie et al., 2018). The rate at which resistance continues to emerge is greatly outpacing the rate of antifungal discovery, highlighting the urgent need for novel therapeutic strategies (Revie et al., 2018). A promising strategy to combat emerging drug-resistant fungi is through combination therapy, as combining drugs has the potential to increase drug effectiveness and can slow the evolution of drug resistance (Spitzer, et al., 2017).
To take advantage of this strategy, we teamed up with a group of scientists at the RIKEN Center for Sustainable Resource Science to screen a large collection of over 20,000 natural products and natural product-like molecules for antifungal activity against the prominent fungal pathogen, C. albicans. We performed these screens in the absence and presence of the widely used azole antifungal, fluconazole, to identify compounds that act synergistically to inhibit the growth of C. albicans. To select compounds with favourable antifungal properties, we assessed each of hit compounds’ toxicity against mammalian cells, activity against drug-resistant isolates, and spectrum of activity across evolutionarily diverse human fungal pathogens. Using this pipeline, we narrowed in on one particularly potent compound, NPD827, that increased the activity of fluconazole by over 32-fold against azole-sensitive strains and over 8-fold against azole-resistant strains of C. albicans. Further, NPD827 was able to potentiate the effect of azole antifungals against a subset of strains of pathogenic species from across the fungal kingdom, including Aspergillus fumigatus, Cryptococcus neoformans, and the emerging pathogen, Candida auris.
Excited by this potent activity against diverse fungi, we investigated the mode-of-action of NPD827 by conducting genetic, biochemical, and biophysical experiments with a collaborative team of experts in these fields. United by our stubbornness to decode the story behind NPD827’s activity, we discovered that NPD827 has profound effects on lipid homeostasis, particularly in the presence of sterol biosynthesis-perturbing agents, such as the azole antifungals. This disruption of lipid homeostasis in turn induces membrane-associated stress responses, including the accumulation of lipid-storage organelles termed lipid droplets and the activation of calcineurin-dependent and unfolded protein stress responses. The impact of NPD827’s activity was visualized by the accumulation of endosome-like vesicles within internal compartments and build-up of the membrane-bound, drug-efflux pump, Cdr1.
Intrigued by the effect of NPD827 on membrane biology, we set out to determine if NPD827 perturbed the ability of membrane-bound proteins to properly function. In testing the effect on drug-efflux, we discovered that NPD827 activity led to the accumulation of fluconazole, as well as the fluorescent efflux substrate Rhodamine-6G, within the cell. Interestingly, these inhibitory effects on drug efflux occurred on rapid timescales, suggesting that NPD827 induced almost immediate changes to the membrane. This led us to the hypothesis that NPD827 may be directly impairing membrane dynamics by altering the biophysical state of the plasma membrane, rather than inhibiting a specific protein target. To explore this possibility, we performed fluorescence recovery after photobleaching (FRAP) lateral mobility analysis to monitor membrane protein dynamics, as well as isothermal titration calorimetry (ITC) to monitor compound interactions with membrane sterols. These biophysical experiments supported our model that NPD827 physically associates with membrane sterols, altering the properties of the fungal membrane and reducing membrane mobility.
Finally, we partnered with additional collaborators to explore the therapeutic potential of NPD827 in treating C. albicans infections. These final studies revealed NPD827 impairs virulence in a Caenorhabditis elegans model of candidiasis, blocks C. albicans filamentation in vitro, and prevents biofilm formation in a rat model of catheter infection by C. albicans.
Overall, this manuscript identifies and characterizes the activity of a molecule with a non-protein-targeted mode of action that re-sensitizes the leading human fungal pathogen, C. albicans, to azole antifungals and impairs critical virulence traits. Importantly, this work would not have been possible without the tremendous support from a team of collaborators that helped us take a hit compound from a high-throughput screen and characterize its unusual and unique mode-of-action, while also highlighting the therapeutic potential of targeting membrane dynamics to cripple drug-resistant and virulence attributes. It was a richly rewarding journey and a true testament to what can be accomplished when great minds come together.
Written by Dr. Nicole Revie and Dr. Nicole Robbins
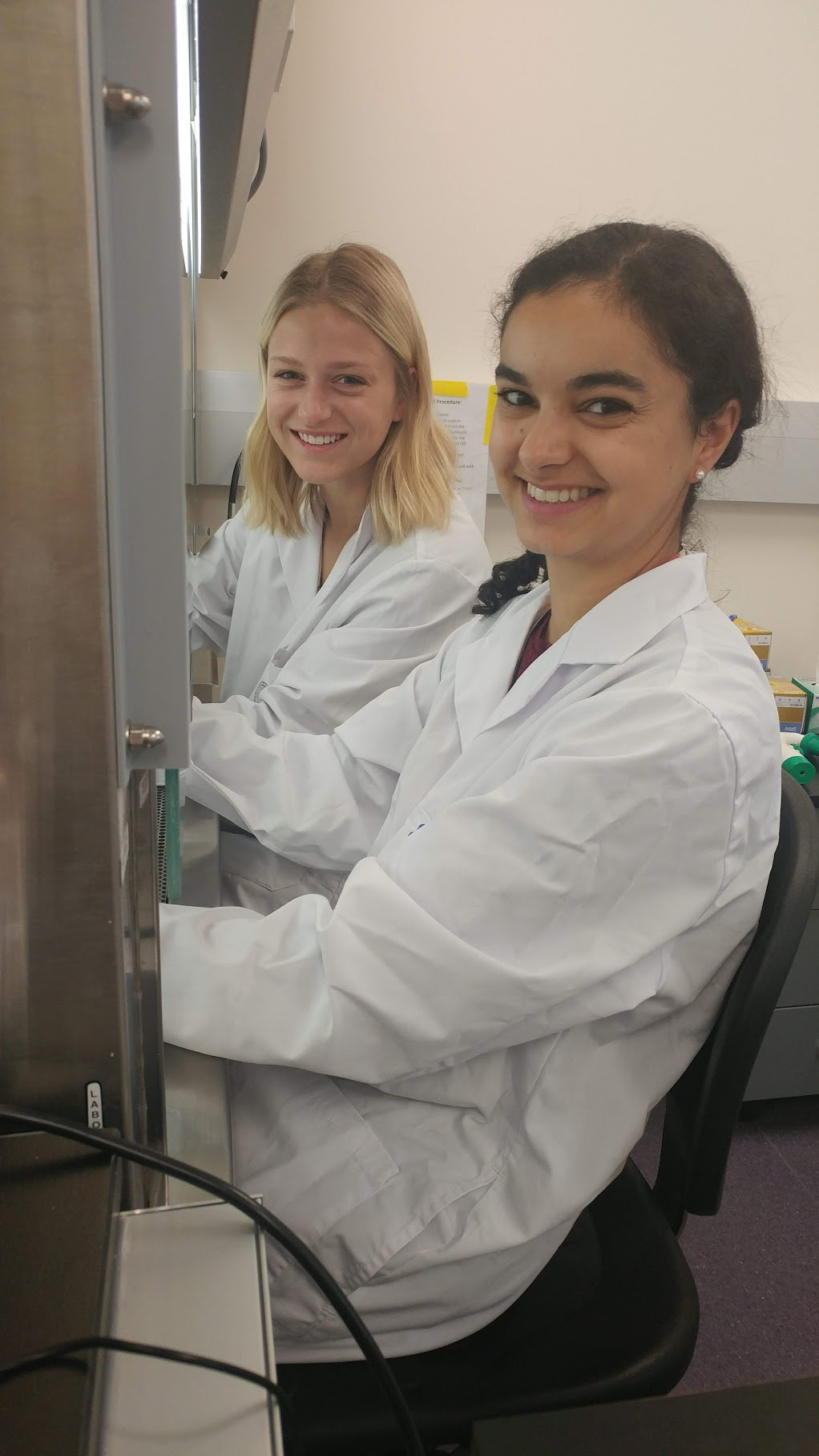
Photo: Drs Nicole Revie and Kali Iyer, the dynamic duo.

References:
Fisher, M. C., Hawkins, N. J., Sanglard, D., & Gurr, S. J. (2018). Worldwide emergence of resistance to antifungal drugs challenges human health and food security. Science, 360, 739– 742.
Brown, G. D., Denning, D. W., Gow, N. A. R., Levitz, S. M., Netea, M. G., & White, T. C. (2012).
Hidden killers: human fungal infections. Science Translational Medicine, 4(165), 165rv13.
https://doi.org/10.1126/scitranslmed.3004404
Revie, N. M., Iyer, K. R., Robbins, N., & Cowen, L. E. (2018). Antifungal drug resistance: Evolution, mechanisms, and impact. Current Opinion in Microbiology, 45, 70–76.
https://doi.org/10.1016/j.mib.2018.02.005.
Spitzer, M., Robbins, N., & Wright, G. D. (2017). Combinatorial strategies for combating invasive fungal infections. Virulence, 8(2), 169–185. https://doi.org/10.1080/21505594.2016.1196300.
Follow the Topic
-
Nature Communications

An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in