The effect of mutations on demographic noise in microbial colonies
Published in Microbiology
Explore the Research
Mutability of demographic noise in microbial range expansions - The ISME Journal
The ISME Journal - Mutability of demographic noise in microbial range expansions
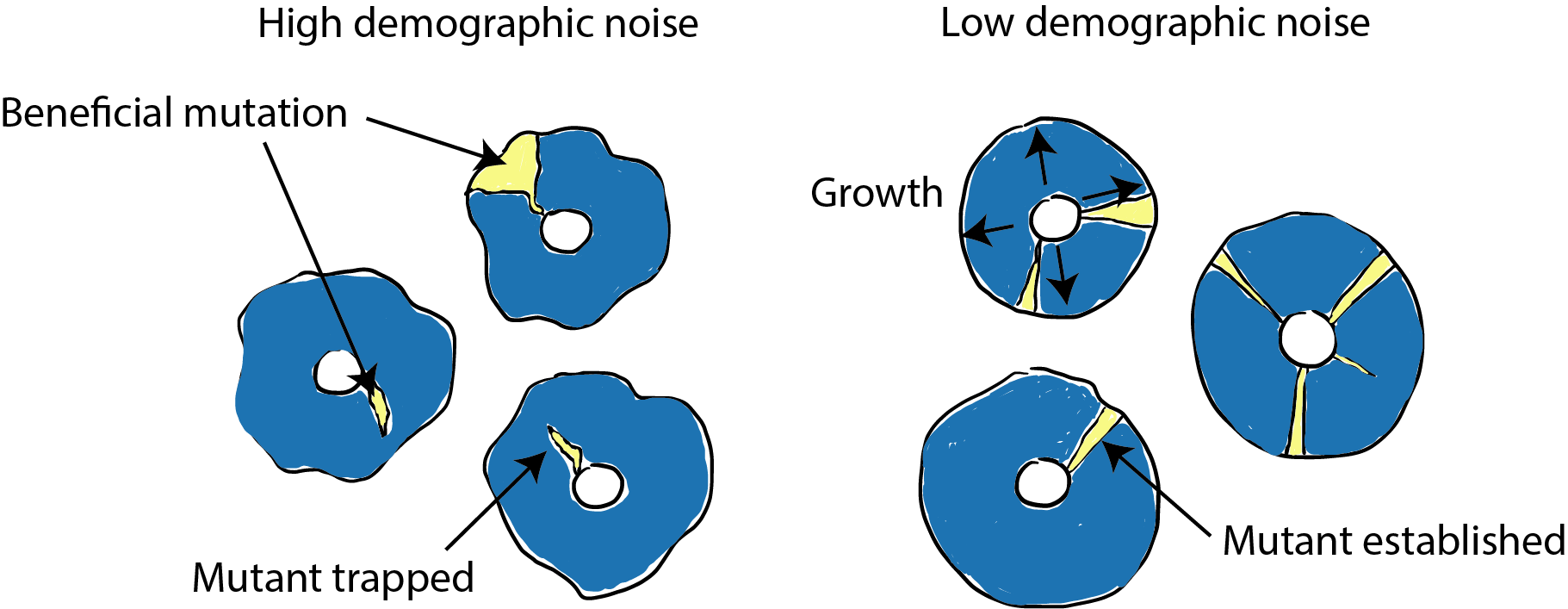
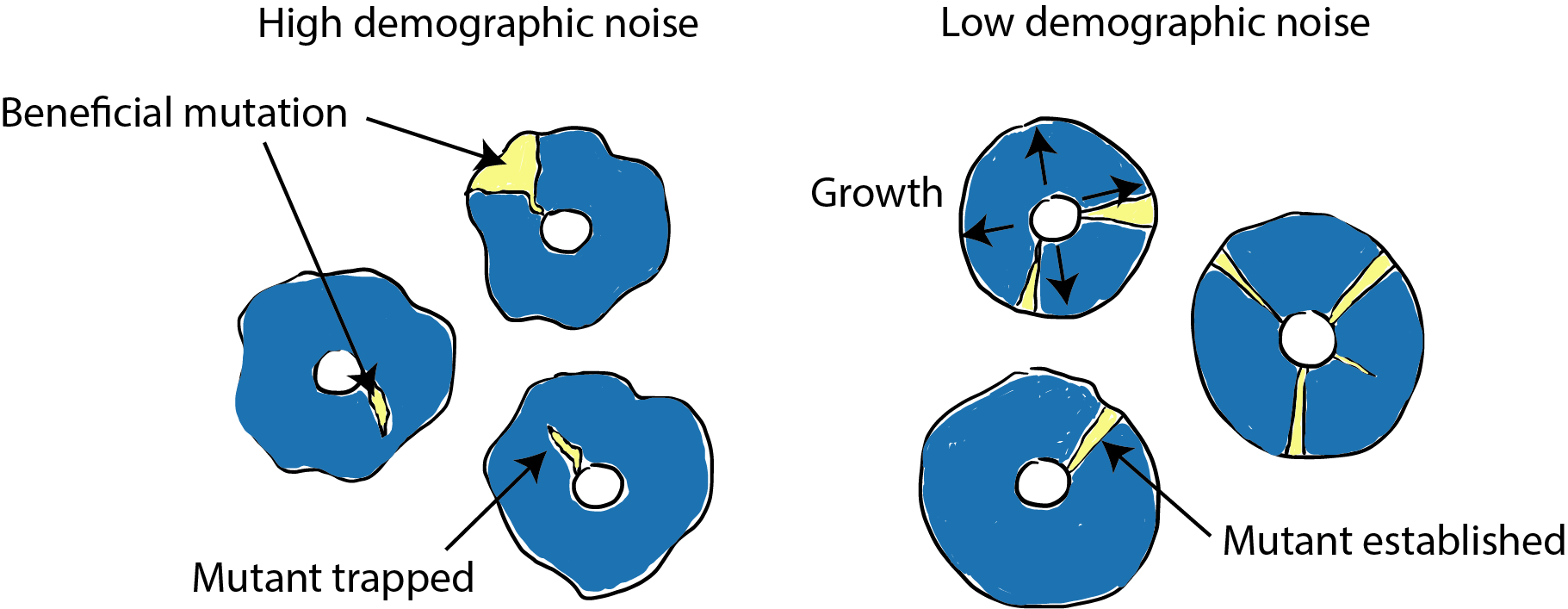
Demographic noise, or “genetic drift”, refers to the role of random chance in the course of molecular evolution.* When a new mutation arises, it is so rare that it can be easily lost from the population due to demographic noise. This means that beneficial mutations can disappear from the population before natural selection has an opportunity to act (Figure 1). Thus, quantifying the strength of demographic noise is important to predicting evolutionary outcomes for new mutations.
The strength of demographic noise depends on various factors; population size is the most commonly studied factor because in small populations, chance events have a larger role. Separately, we previously found that in microbial colonies, strains from the same species exhibited different strengths of demographic noise—in one instance, even in a strain that only differed by a single gene deletion – suggesting that genetics is another factor that affects demographic noise in colonies.
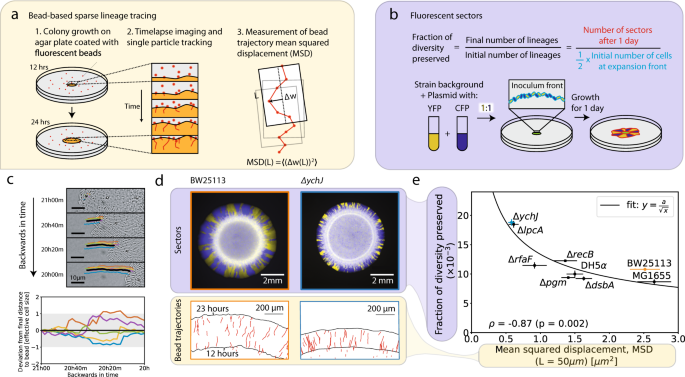
Given the role of genetics in setting the strength of demographic noise, we wondered how much single mutations can have an effect on demographic noise, analogously to studying how mutations affect fitness. We used an established E. coli gene deletion collection called the Keio collection as our model system, because loss of function mutations commonly occur in bacteria in a single mutation event. We wanted to observe the fate of lineages from cells in the initial population of the microbial colonies in order to measure demographic noise, which causes diversity to be lost from the initial population (Figure 2a).
In order to study demographic noise in a large number of genotypes, we developed a high-throughput method that uses fluorescent tracer beads to follow cell lineages. We add the beads onto the surface of a solid medium and measure their movement as a seeded colony grows (Figure 2b). We established that large displacements of the beads away from the average direction of motion correspond to higher demographic noise by showing that it is a proxy for the amount of diversity that is lost (Figure 2c). This is not necessarily intuitive, so let’s break down why that is the case: first, displacements of the fluorescent beads are a good proxy for the movement of the bacterial cells because the cells push the beads as they grow. Second, higher displacements away from the average direction of motion result from more random cell lineage trajectories, which suggest higher demographic noise (larger role for chance events).
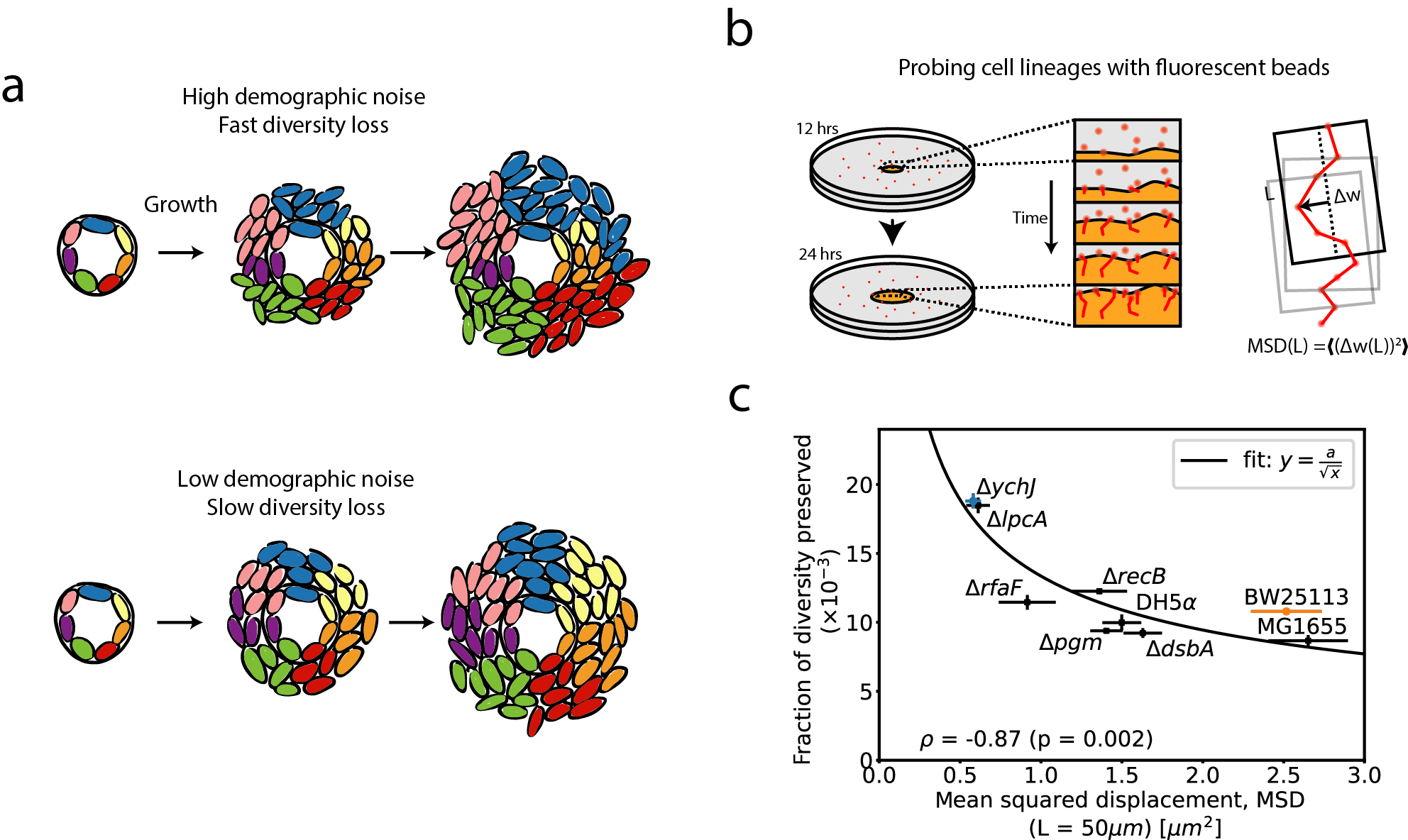
The scale of these experiments was unusual: in hopes of having enough statistical power, we traced moving beads from more than 1000 colonies and generated almost 3 TB of imaging data. Excitingly, we saw a signal of more variable demographic noise in single gene deletion strains than we would expect purely due to experimental noise. It was exciting and somewhat surprising to see a signal from randomly selected single gene deletion mutations, suggesting that the mutations which change the strength of demographic noise are not special but can commonly occur.
We wanted to understand what effect this phenomenon has on adaptive evolution, so we tested what happened to a new beneficial mutation that appears in an environment with altered demographic noise. Population genetics theory predicts that higher demographic noise should make it harder for a beneficial mutation to establish (grow to a large enough size for selection to take over) because it has to overcome relatively larger random effects (Figure 1).
Excitingly, we found that single gene deletion strains can enhance the probability that new beneficial mutations establish, specifically because of decreased demographic noise.+ Taken together, our result shows that mutations can change the strength of demographic noise and that these effects can change how evolution proceeds. We think this is a novel and interesting way to think about how single mutations affect evolution, effectively by adding an additional “knob” on the strength of demographic noise that evolution can tune.
If you want to learn more, check out our paper in ISME J, where we also address the question of how mutations affect demographic noise in microbial colonies.
* Note that demographic noise is different from mutations, which also occur randomly. Mutations generate new genotypes, but demographic noise affects how the amount of each existing genotype in the population changes over time.
+ We expect that the other direction also happens, that single gene deletion strains with increased demographic noise can make it harder for beneficial mutations to establish, but we happened to not have tested those strains in this particular experiment.
Acknowledgements: We are grateful to Melanie Abrams and Arbel Harpak for providing valuable feedback on drafts of this blog post to make it more accessible and interesting to read.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in