Uncover natural variation left behind in wild emmer for modern wheat improvement
Published in Plant Science
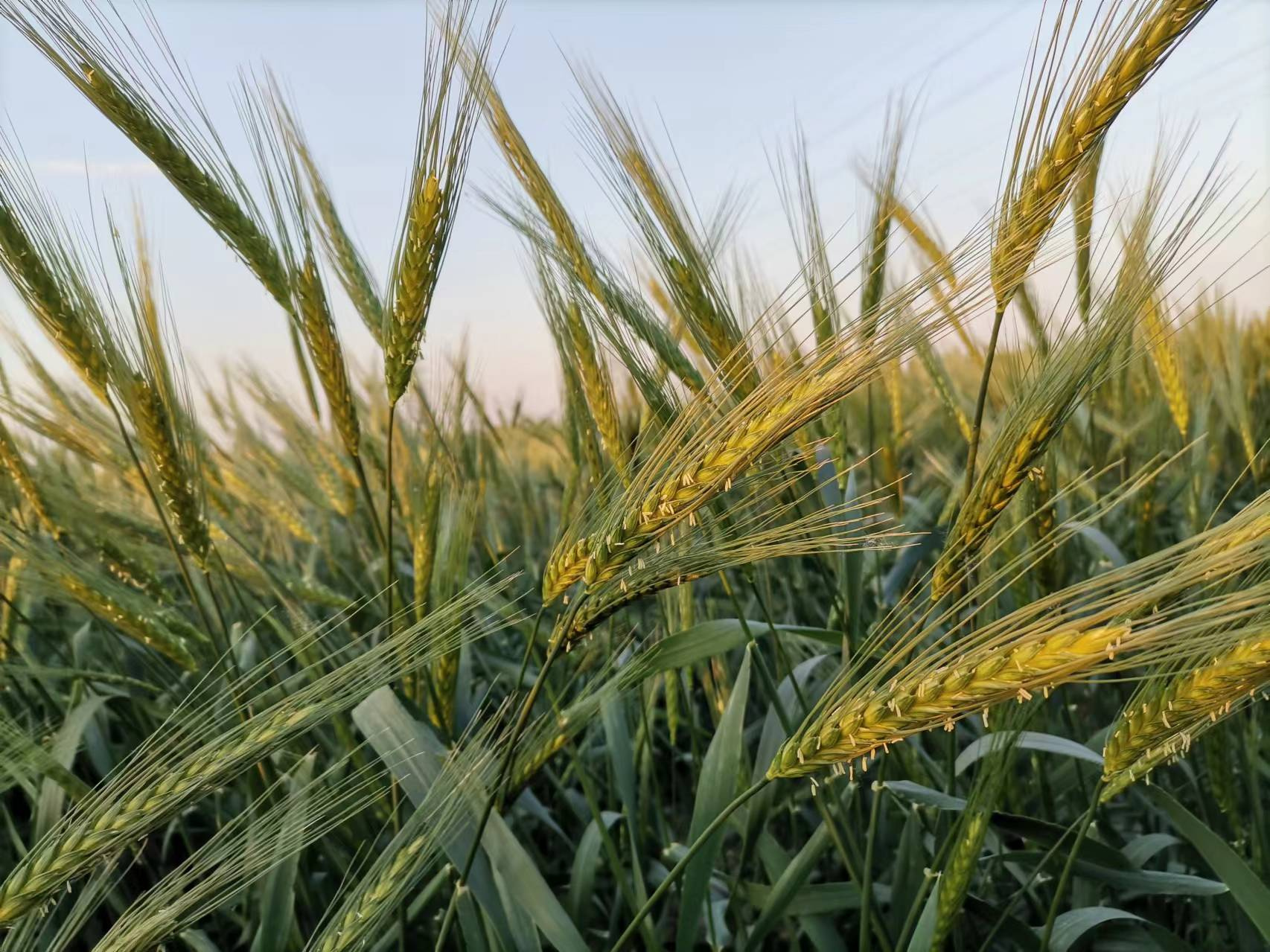

Wheat is one of the most important crop plants in the world that can be traced back to the universal cereal of Old World agriculture. Modern wheat, including tetraploid hard or durum-type wheat, Triticum turgidum durum (2n = 4x = 28, AABB), and hexaploid bread wheat, T. aestivum (2n = 6x = 42, AABBDD), ranks the first in the world’s grain production and accounts for more than 20% of the total human food calories. Due to the bottleneck limitation of domestication and hexaploidization processes, modern wheat suffers extremely narrow genetic diversity that led to genetic erosion and increasing susceptibility and vulnerability to environmental stresses, pests, and diseases. Wild emmer wheat, T. dicoccoides, (2n = 4x = 28, AABB) (Figure 1), is proved to be the direct progenitor for both durum and bread wheats and harbors rich genetic resources for wheat improvement. One of the best strategies to enrich the genetic diversity of modern wheat is to utilize the adaptive genetic resources of the wild emmer wheat.
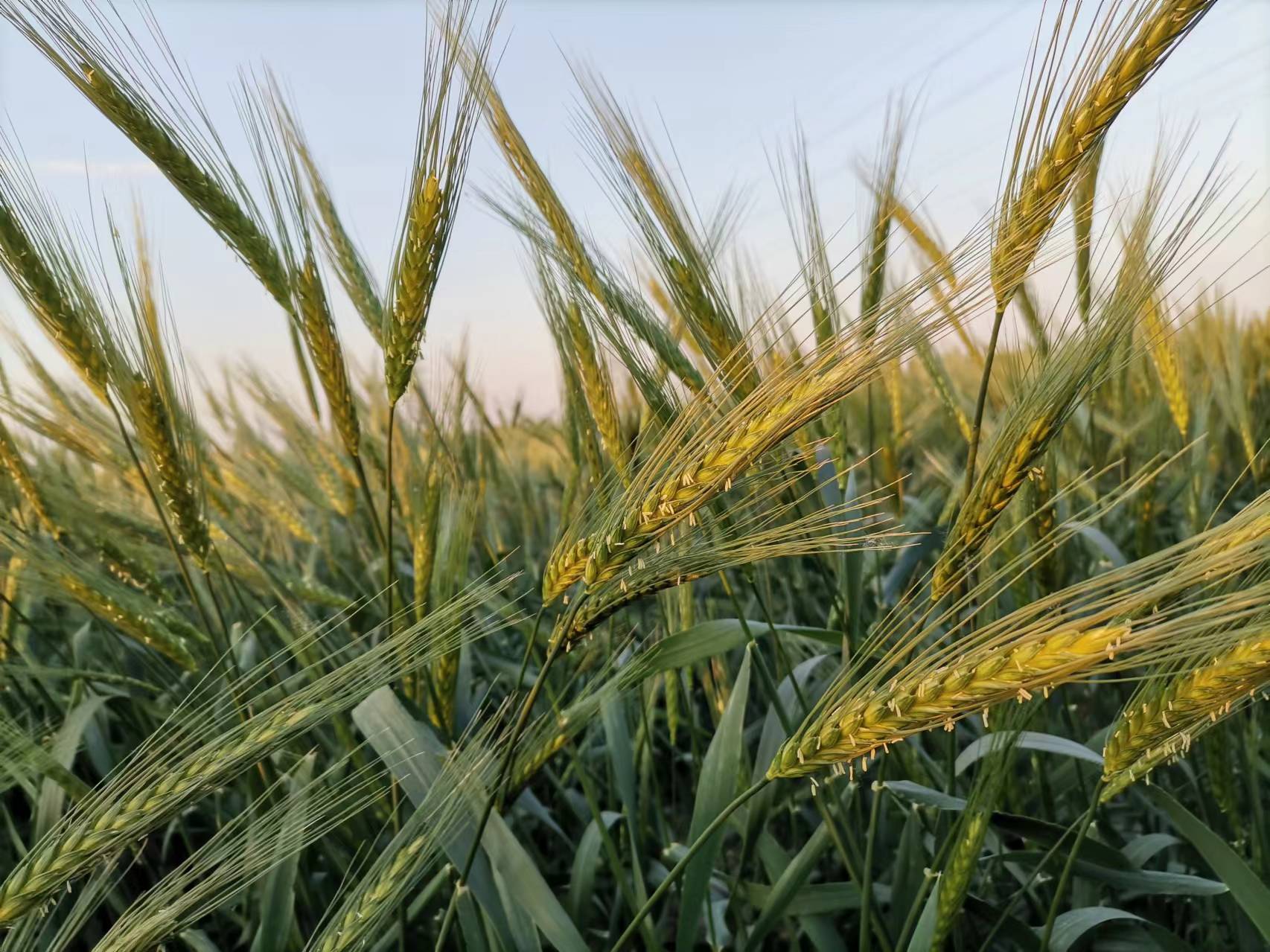

Wild emmer wheat possesses many important beneficial traits that can be used for modern wheat improvement. Those traits include but not limited to biotic stress resistance such as stripe (yellow) rust (Puccinia striiformis f. sp. tritici), stem rust (Puccinia graminis f. sp. tritici), powdery mildew (Blumeria graminis f. sp. tritici), and soil born wheat mosaic virus, abiotic stress tolerance such as drought tolerance, herbicide resistance, amylases and alpha amylase inhibitors, grain nutrition such as amino acid composition, grain protein content, storage protein genes (HMW glutenins), salt and micronutrients (Zn and Fe), as well as high biomass and yield potential related traits such as high photosynthetic yield, earliness, short stature, and high tillering capacity. However, due to the polyploid nature with complex genome, only a few genes were successfully isolated from wild emmer and functional characterized. Those genes include the high grain protein content GPC-B1, stripe resistance genes Yr36 and Yr15, powdery mildew resistance genes Pm41, Pm69, Pm60 alleles, and Pm36.
Pm36 was first identified in wild emmer - durum wheat backcross inbred lines (BC5F5) 5BIL-29 and 5BIL-42. Dr. Antonio Blanco and his colleagues from the University of Bari, Italy, used a semi-dwarf and high-yielding cultivar Latino of durum wheat as a recurrent parent to make cross with wild emmer accession MG29896 for its high grain protein content, acceptable seed size, and powdery mildew resistance. By using amplified fragment length polymorphism (AFLP), simple sequence repeat (SSR), and expressed sequence tag (EST)-derived makers, Pm36 was mapped in 5BL chromosome that can be used for powdery mildew resistance breeding in durum wheat.
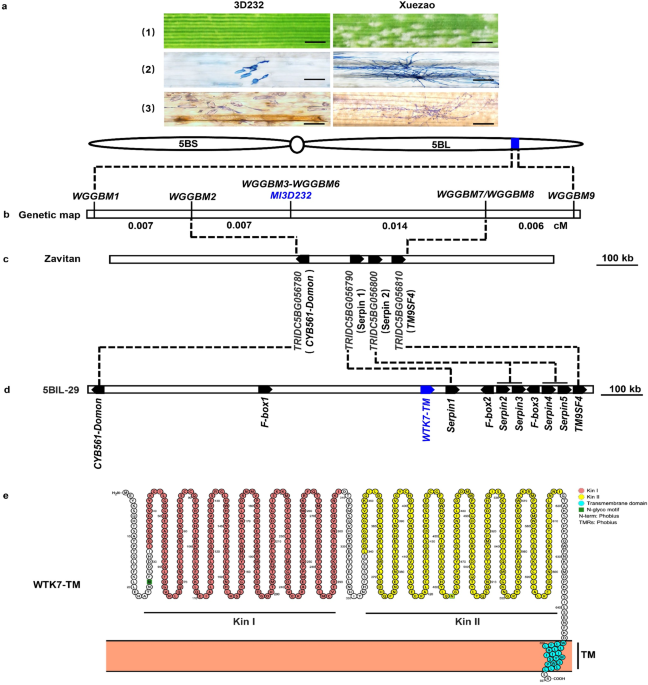
In the same period of time, Prof. Tsomin Yang from China Agricultural University developed a powdery mildew resistance wild emmer - common wheat backcross introgression line 3D232 (87-1*6/I222) by crossing and backcrossing susceptible elite common wheat line 87-1 with wild emmer accession I222, a powdery mildew resistance wild emmer wheat kindly provided by Dr. Z. K. Gerechter-Amitai, Agricultural Research Organization, The Volcani Center, Israel. 3D232 was found resistant to all 104 available Chinese Blumeria graminis f. sp. tritici isolates, indicating its broad-spectrum resistance and potential value in breeding disease resistance wheat cultivar. By applying molecular markers and bulked segregant analysis (BSA), the powdery mildew resistance locus in 3D232 was located on chromosome 5BL using polymorphic SSR, EST-derived sequence tagged site (STS) markers. Further comparative genomic analysis and linkage maps revealed that the powdery mildew resistance gene Pm36 in 5BIL-29 and 5BIL-42 and Ml3D232 in 3D232 were located in the same genetic interval and co-segregated with same molecular markers XBD37670 and XBD37680, indicating they are likely the same gene or alleles in the same locus.
In order to isolate the gene responsible for powdery mildew resistance in 3D232, a F2 segregation population consisting of 15,893 individual (31786 gametes) derived from the cross between a highly susceptible common wheat line Xuezao and 3D232 was used to fine map Ml3D232 in a genetic interval of 0.021-cM, which corresponds to 273.8 kb genomic region of wild emmer wheat Zavitan reference genome sequence. Only four high-confidence genes were predicted in this genomic region and two of them, CYB561-Domon and TM9SF4, were found constitutively expressed in the leaves of 3D232 and Xuezao seedlings. Since CYB561-Domon and TM9SF4 performed differential expression patterns and coding sequence polymorphisms between 3D232 and Xuezao, those two genes were separately delivered into highly susceptible wheat cultivar Fielder by Agrobacterium-mediated transformation for functional characterization. Unfortunately, all positive transgenic plants for CYB561-Domon and TM9SF4 genes in the independent T1 transgenic families were highly susceptible to powdery mildew, suggesting they are not the causal gene of Ml3D232.
We then wondering if these is genomic structure variation in the Ml3D232 gene region? Synteny analysis of the Ml3D232 genomic region in all available assembled wheat and wild relatives genome sequences indicated no significant sequence insertion/deletion and rearrangement in this region on 5BL chromosome. This result raised the possibility of genomic structure variation for the Ml3D232 genomic region in this particular wild emmer accession. Take the advantage of PacBio SMRT long-read sequencing, the tetraploid wild emmer-durum introgression line 5BIL-29 carrying Pm36 was sequenced to generate HiFi reads approximately 19.4-fold coverage of tetraploid wheat genome. To improve the genome sequence assemble, 117.3-fold Hi-C reads were also produced further. A tetraploid wheat genome assembly containing 2,207 contigs was obtained, and a 7.1 Mb contig ptg000422 spanning the entire Pm36 physical mapping interval (1.17 Mb) was captured. Unsurprisingly, a ~900 kb genomic sequence insertion in the Pm36 physical region harboring two segmental duplications containing 7 more predicted genes was found and a putative tandem kinase with predicted transmembrane domain (WTK7-TM) was further confirmed to be the Pm36/Ml3D232 gene.
Haplotypic variation analysis revealed that Pm36/Ml3D232 (WTK7-TM) and all the so far cloned genes (GPC-B1, Yr15, Yr36, Pm41, Pm69, Pm60 alleles, and Pm36) from wild emmer wheat are only presented in the wild emmer gene pool. Geographic distribution analysis also suggested that Pm36, Pm41, Yr15, and Yr36 were co-located in the southern wild emmer wheat populations, providing further evidence for host-parasite co-evolution in the wild emmer wheat natural habitats. The absence of Pm36, Pm41, together with Yr15 and Yr36, in wild emmer wheat natural populations from southeastern Turkey where wheat is believed to be domesticated, as well as in cultivated tetraploid and hexaploid wheat, revealed that these disease resistance genes were left behind, rather than lost during wheat domestication and polyploidization.
The majority of cloned wheat resistance genes, including Pm41, Pm60 alleles, and Pm69 isolated from wild emmer wheat, encode for intracellular nucleotide-binding leucine-rich repeat receptor (NLR) proteins with coil-coil (CC) N-terminal, conserved nucleotide-binding (NB) and C-terminal leucine-rich repeat (LRR) domains. Tandem kinase protein (TKP) was first characterized in barley as a new type of resistance protein for durable stem rust resistance gene Rpg1. Recently, new members of TPKs have been characterized in wheat and wild relatives for stripe rust (Yr15/WTK1), stem rust (Sr60/WTK2 and Sr62/WTK5), leaf rust (Lr9/WTK6-vWA), powdery mildew (Pm24/WTK3, WTK4, Pm36/WTK7-TM, Pm57/WTK6-vWApm), and wheat blast (Rwt4) resistance. Among them, Yr15/WTK1 and Pm36/WTK7-TM were uncovered from wild emmer wheat and proven to resistant to diversified P. striiformis f. sp. tritici and B. graminis f. sp. tritici isolates. Pm36/WTK7-TM has been used to develop advanced breeding lines compatible with commercial wheat cultivars in plant architecture and yield potential (Figure 2) to safeguard the food security. These results highlight the importance of conservation and exploitation of wild emmer wheat gene pool for modern wheat improvement.

Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in