Who's a Bonehead? Novel Insights into Evolutionary History from Reptilian Skull Roof Structure
Published in Ecology & Evolution
Why are terrestrial vertebrates so highly diverse? In times of anthropogenic change and mass extinctions, this key question to evolutionary research is more relevant than ever. We often attribute this phenomenon to modifications in accordance with different environments and lifestyles. However, it is not always easy to reconstruct from the fossil record how historic key events determined the destinies of entire lineages millions of years ago. Therefore, 160 years after Darwin's theory of evolution, our understanding of certain evolutionary mechanisms remains incomplete.
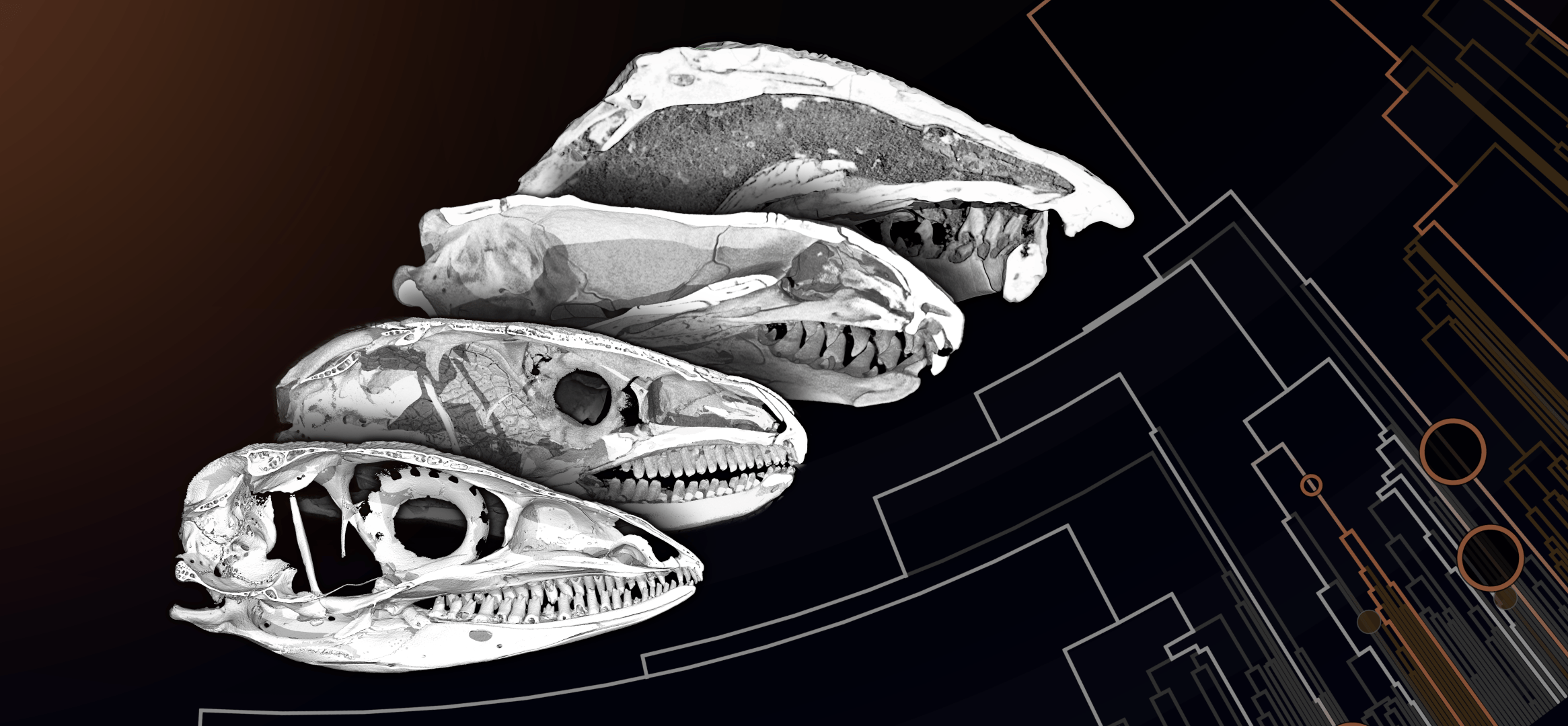
In order to better understand the complex interrelations between vertebrate lifestyle, form, and evolution, we looked into the skull roof structure of squamate reptiles – i.e., lizards and snakes. We wanted to understand to what extent their bone structure reflects certain lifestyles. Although many of these reptiles use their skulls as a digging tool, this question has never been systematically investigated. In our paper First Evidence of Convergent Lifestyle Signal in Reptiles Skull Roof Microanatomy, we therefore employed computer simulations to reconstruct the evolution of a specialised burrowing lifestyle over a period of 240 million years. Remarkably, we found that burrowing evolved independently in 54 squamate lineages. These reptiles are therefore particularly well suited as a model system for the study of convergent evolution.
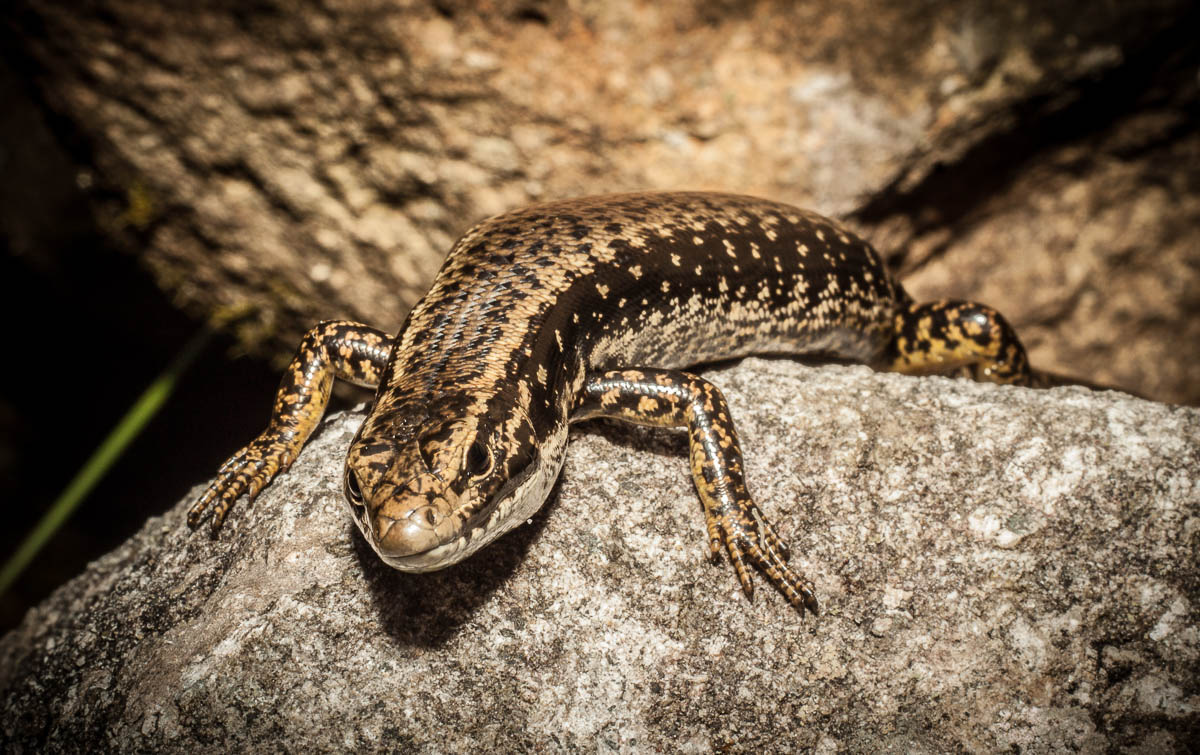

Scincoid lizards, such as this representative from the Australian east coast, are among the most diverse squamates – both ecologically and taxonomically. Nonetheless, we were surprised to identify 20 independent acquisitions of a specialized burrowing lifestyle in this clade alone. Photo by Roy Ebel.
In the second phase of our study, we compared the skull roof structure of lizards and snakes in accordance with their lifestyles. To this end, we made use of high-resolution computed tomography (micro-CT) for the 3D visualisation of bone tissue and employed a new, effective protocol for data analysis. Doing this, we achieved a large sample size, allowing us to draw conclusions about the entire squamate reptilian clade with over 11,000 species. We found that burrowing lizards and snakes have repeatedly evolved a particularly dense and compact skull roof in independent evolutionary processes. We further identified typical proportions of both the skull and between the skull roof bones as convergently evolved modifications associated with this lifestyle.
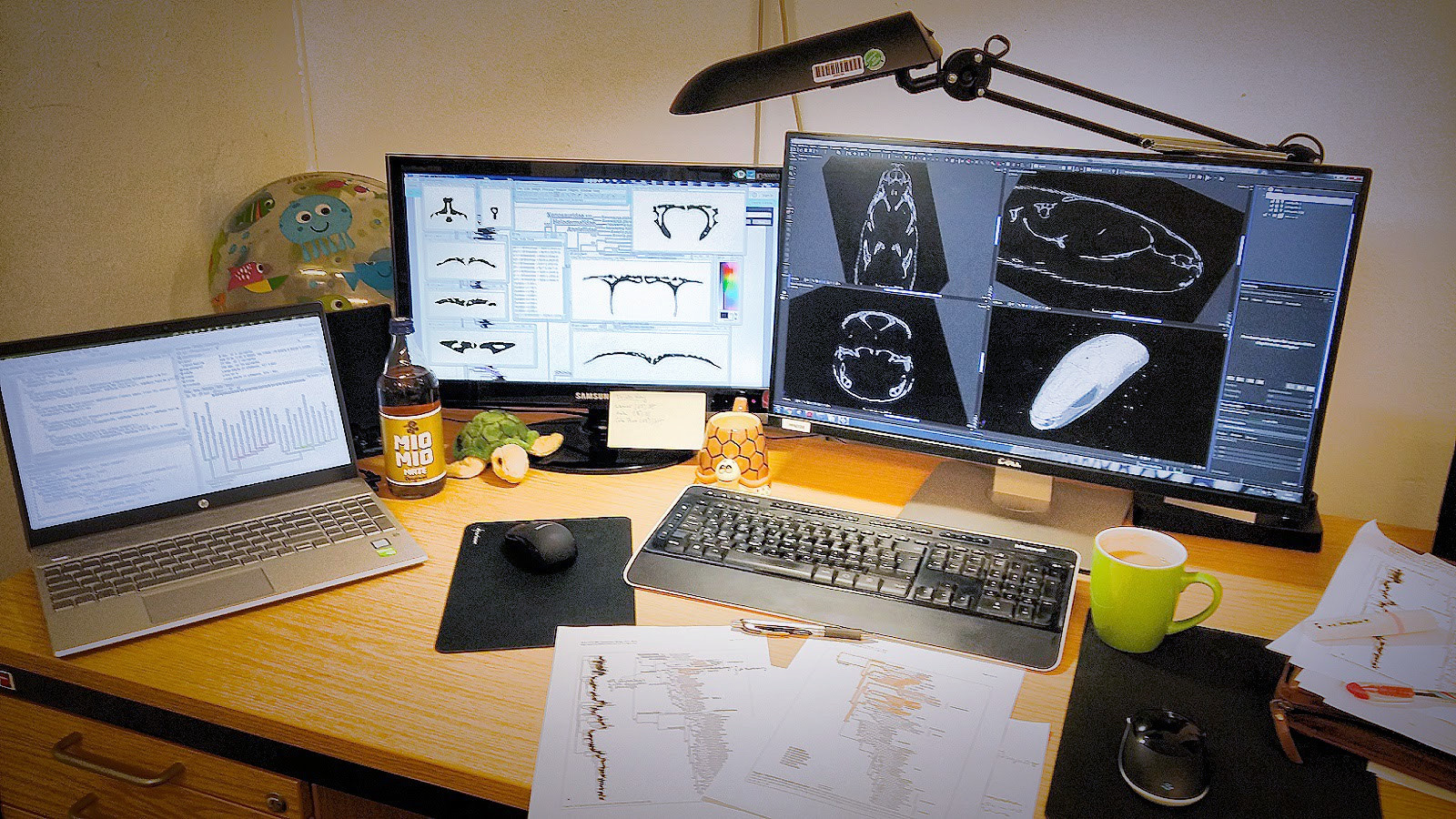

This desk at the CT-Vis-Lab of the Museum für Naturkunde Berlin says quite a bit about our working routine. From right to left: VG Studio MAX for processing 3D-volumes acquired with µCT, ImageJ for processing extracted slices, and R-Studio for the analysis of convergence and lifestyle signal. As a coincidence, the stages of data reduction correlate with decreasing screen size. ...and who would have thought that this is how a biologist's workspace may look like? Photo by Roy Ebel.
In our study, we present a novel case of convergent evolution: in different lineages, very similar structures have independently evolved in response to a common lifestyle. Such similarities reflect a certain function, in this case burrowing, and may therefore provide little information on the phylogenetics relations between the considered taxa. Nonetheless – or precisely therefore, the knowledge of these processes is of outstanding importance: by means of skull roof structure, we can now reconstruct the lifestyle of reptiles that became extinct millions of years ago. Our findings thus cast completely new light on the evolutionary history of certain lineages. This may particularly apply to snakes, for which both an aquatic and burrowing origin have been controversially discussed for decades. Our work now adds to a growing body of evidence for the latter. Having said this, our findings may also have important implications beyond squamate reptiles. A burrowing lifestyle has likely played a key role in the evolutionary history of turtles and certain amphibians. Recent studies even suggest that the mammalian stem lineage may not have survived the largest mass extinction in earth's history at the end of the Permian about 250 million years ago without adopting a burrowing lifestyle. It is therefore invaluable to understand such historical lifestyle transitions.
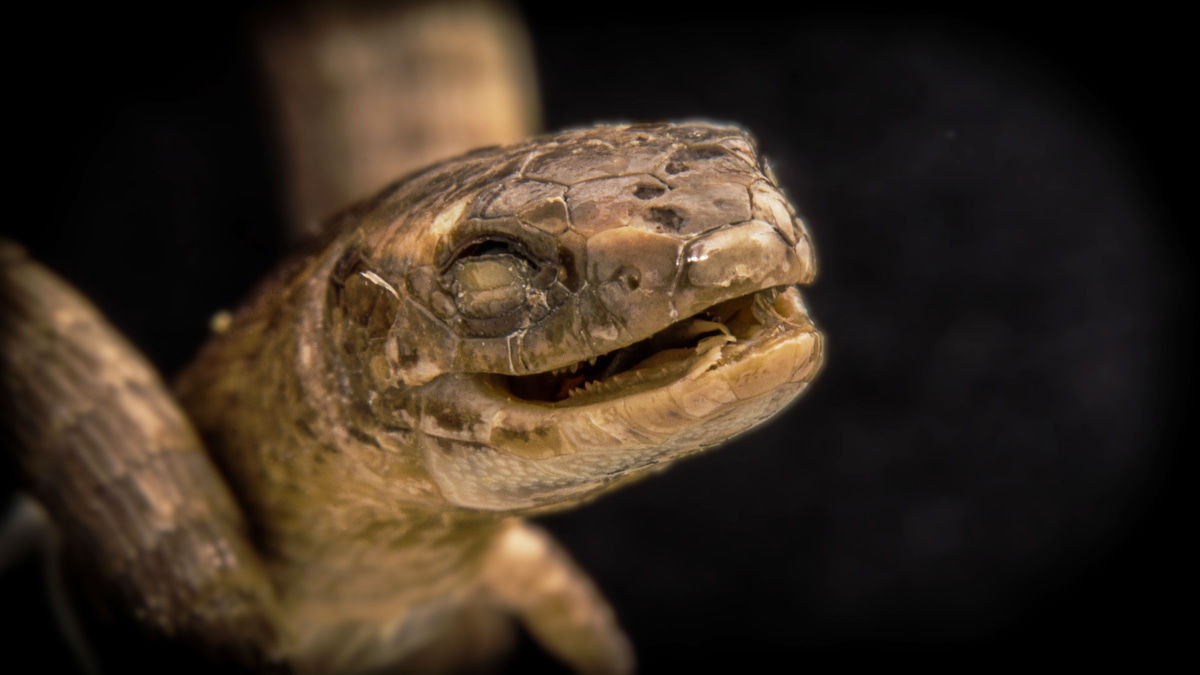

This little critter is a rare burrowing gymnophthalmid lizard from Brazil, which quickly became a lab favourite for its happy smile. We were able to include the specimen in our dataset due to a loan from the American Museum of Natural History (AMNH R64876). In total, our dataset comprises specimens from 13 herpetological collections worldwide. Photo by Roy Ebel.
Should our work have sparked your interest in the fascinating field of squamate skull morphology and its implications for their ecology and evolution, you may be pleased to hear that we uploaded the µCT-scans employed in our study into the Data Repository of the Museum für Naturkunde Berlin. They will thus be available for future research projects worldwide.
Follow the Topic
-
BMC Biology
This is an open access journal publishing outstanding research in all areas of biology, with a publication policy that combines selection for broad interest and importance with a commitment to serving authors well.
Related Collections
With Collections, you can get published faster and increase your visibility.
Cancer metabolism
BMC Biology is calling for submissions to our Collection on Cancer metabolism. Cancer metabolism is a developing field that explores the biochemical and physiological changes that occur in cancer cells, which often exhibit distinct metabolic pathways compared to normal cells. Alterations such as the Warburg effect, where cancer cells preferentially utilize glycolysis for energy production even in the presence of oxygen, play a pivotal role in tumor progression and survival. Understanding the intricacies of cancer metabolism provides insights into how tumors adapt to their microenvironments and highlights potential therapeutic targets for intervention.
Future research in cancer metabolism promises to yield transformative insights that could reshape therapeutic approaches and improve precision medicine. The continued focus on metabolic pathways may lead to the identification of new biomarkers for cancer diagnosis and prognosis, as well as novel strategies to enhance the sensitivity of cancer cells to treatment. As we deepen our understanding of the metabolic landscape of tumors, we may uncover innovative strategies that exploit these vulnerabilities, ultimately leading to novel and more effective cancer treatments, as well as improved patient outcomes.
Recent advancements in the field, including the identification of metabolic reprogramming strategies and the influence of diet on tumor growth, have opened new avenues for research. Investigations into the roles of lipids, fatty acids, and dietary interventions, such as ketogenic diets, are revealing potential methods for manipulating tumor metabolism and enhancing the efficacy of existing treatments. We invite researchers to submit their work to this Collection, which aims to showcase groundbreaking research and technologies addressing cancer metabolism and support the advancement of this field, encompassing a wide array of topics related to metabolic pathways and their implications for cancer biology and therapy.
Potential topics include but are not limited to:
- Metabolic reprogramming in cancer cells
- The Warburg effect and its implications
- Role of mitochondria in cancer metabolism
- Impact of dietary interventions on tumor metabolism
- Glycolysis and lipid metabolism in cancer
- Animal models
- Imaging and method developments
- Metabolic engineering
This Collection supports and amplifies research related to SDG 3 (Good Health and Well-being).
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Oct 30, 2026
Biology of neurodegenerative diseases
BMC Biology is calling for submissions to our Collection on the biology of neurodegenerative diseases. This Collection aims to bring together multidisciplinary knowledge to better understand the mechanisms driving progressive neuronal dysfunction and loss.
We welcome studies investigating key pathological mechanisms, including protein misfolding and aggregation, neuroinflammation, oxidative stress, mitochondrial disorders, dysfunction of cellular protein sorting and degradation, altered RNA metabolism, blood-brain barrier impairment, brain vascular dysfunction, contribution of extracellular vesicles and synaptic dysfunction. Research exploring intracellular transport disruptions, neuronal network alterations, and genetic or epigenetic contributions to neurodegeneration is also encouraged.
We are particularly seeking submissions that employ state-of-the-art approaches, such as multi-omics, electrophysiology, high-resolution imaging, (induced) pluripotent stem cell-derived model systems, (e.g. microfluidics 2D co-culture, organoids and assembloids), and animal models to gain deeper mechanistic insights into neurodegenerative processes and identify potential therapeutic targets.
This Collection supports and amplifies research related to SDG 3: Good Health and Well-Being.
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Sep 03, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in