After the Paper: A retroviral mystery tour
Published in Microbiology
Explore the Research

Genetic organization of a chimpanzee lentivirus related to HIV-1 - Nature
Nature - Genetic organization of a chimpanzee lentivirus related to HIV-1
If a new microbe turns up today in a handful of humans, associated with an unusual pathology, you can be reasonably sure the nucleic sequence will be established before the syndrome has been adequately described. In the click of a BLAST the sequence will be linked to something from a mammal, bird or insect. Search posies will then round up some of the unusual suspects, isolate the microbe, sequence 10 or more isolates and finger the adaptive mutations. A bream of papers will then follow. The outbreak may die down and not be seen for a decade; alternatively, the microbe may continue to cross over to humans yet remain at low levels. All will be dubbed emerging infections in grant proposals no matter how minor. Iterate the above narrative n times.
Just occasionally, an unusual microbe will go big time. The AIDS pandemic exploded in Central Africa in the post-WW2 period although the virus crossed over to humans earlier, sometime in the first half of the 20C.
HIV/AIDS needs no introduction. It brought out the worst and the best in people. It transformed clinical infectious disease research; it discombobulated immunology which had been far too happy working with inbred mice; it took virologists totally off guard. Rapidly HIV was in the vanguard of everything as the research $$$ poured in. Fortunately, all this happened at the time when molecular biology was ramping up into exponential phase. The PCR tsunami broke in 1989. HIV/AIDS even morphed language. When I was young it was good to be seropositive for a microbe. Today, if you’re seropositive, the sentence stops.
It showed us just how little we knew.
Fast backwards to pre-HIV retrovirology, a field dominated by the oncoviruses of mice and chickens, understandably because you could do experiments with them. There was bovine leukemia virus, but who was interested in cows? A sick animal would be slaughtered. Having said that, remember; work with a chicken tumor retrovirus gave us the oncogene, a phenomenally important discovery (1). It saw the morphing of endless retrovirologists into cancer specialists. A few lentiviruses were isolated from sheep, goats and horses but they caused neurological disease, arthritis or infectious anemia, decidedly un-royal, aka they didn’t induce cancers. The spumaviruses were the retrovirologists black hole. Yet in the absence of a pathogenic human retrovirus animal oncoviruses dominated. In 1980 a serious tremor occurred with the isolation of human T cell leukemia virus (HTLV-1) from humans (2,3). Another tremor quickly followed when the much rarer HTLV-2 was isolated in 1982 (4). Where did these retroviruses come from? Quickly it emerged that there were simian lookalikes in some species of macaques.
Enter HIV in 1983 (5). Despite the HTLV/STLV connection, the immediate reaction was where the hell did it come from? My group was the first to publish the sequence of the HIV genome in 1985 (6). It was larger than any of retroviral genome, while the synteny of its orfs was unique. Uninformed stories in the press fingered the KGB, others the CIA. The answer was neither; no retrovirologist then, or since, had the knowledge to design anything so sophisticated.
A paper pointed out a one way cross with a horse lentivirus (7), while another showed similar electron micrographs for HIV and visna virus, a sheep lentivirus (8). Interestingly the visna genome was as large as that of HIV, while its base composition was almost identical (9). Being a young cloner, nothing made more sense than genome data. We powered ahead and published the sequence of visna virus 7 months later (10). The two genomes had similar orf synteny and were closely related phylogenically. That they were cousins was shocking: lentiviruses were known to cause a slow remorseless disease - 100% of experimentally infected sheep ultimately developed disease.
One of the biggest questions of the early days in HIV/AIDS was what fraction of seropositive individuals would develop the disease? I remember the figure of 10% being bandied around. Why 10%? Simply because the proportion of seropositive individuals presenting with AIDS was around 10%. The logic was one of an acute infection even though we have known for ages that retroviral infections are forever. With visna as a cousin the unavoidable conclusion was that vast majority seropositive individuals would progress to AIDS. This was the beginning of the dark tunnel from which we emerged a decade later with tritherapy.
As the reader may well know a lentivirus was identified in rhesus macaques presenting with AIDS in 1985 (11). How was this simian immunodeficiency virus (SIV) overlooked? The premium had been on oncogenic and experimentally tractable viruses of course. Circa 1985, immunosuppression was typified by children living in bubbles or terminal stage cancer patients. Acquired immunosuppression was unheard of. HIV-2 was quickly discovered in 1986 and its genome proved isogenic to the macaque lentivirus (12,13). As things would have it, this virus was not natural to the rhesus monkey but had come from humans experimenting with cancerous material. It transpired that an endogenous asymptomatic lentivirus of sooty macaques had jumped over to humans spawning HIV-2, again sometime in the early 20C. By contrast HIV-1 and HIV-2 differed by two mutually exclusive orfs so while we were further on, the mystery continued. By now researchers were starting to smell a rat and started looking at African apes and monkeys.
Martine Peeters and Eric Delaporte, two Paris based researchers, were working at the primate research center in Libreville Gabon. Their main task was screening patients by ELISA and Western blotting (WB). Given the simian connection they decided to screen sera from the animals in the primate center. They identified a four year old female chimpanzee with chronic lymphoadenopathy. She presented a troubling WB. Fortunately, they were working in the lab of Françoise Brun-Vezinet who had mastered culturing HIVs and succeeded in making the second isolate of HIV-2. Not surprisingly a chimpanzee virus was quickly isolated and called SIVcpz (14). My group was fortunate to be already working with Françoise, who with Christine Rouzioux a few years earlier, made the first really good HIV-1 ELISA.
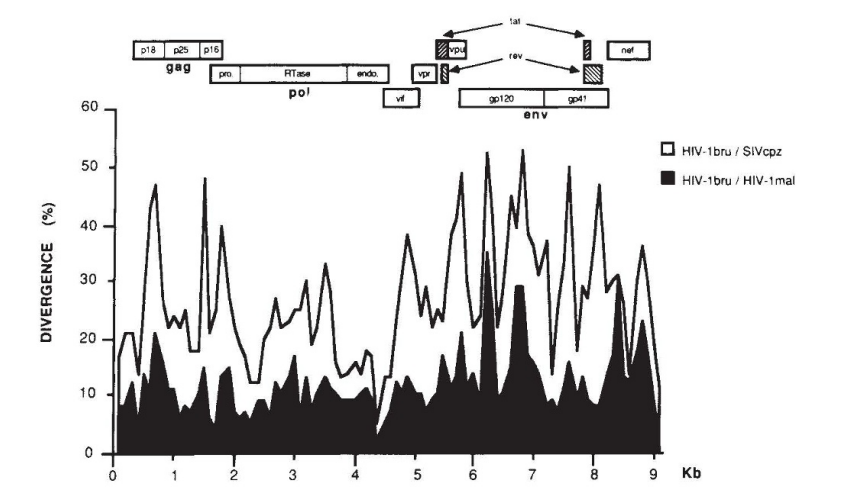
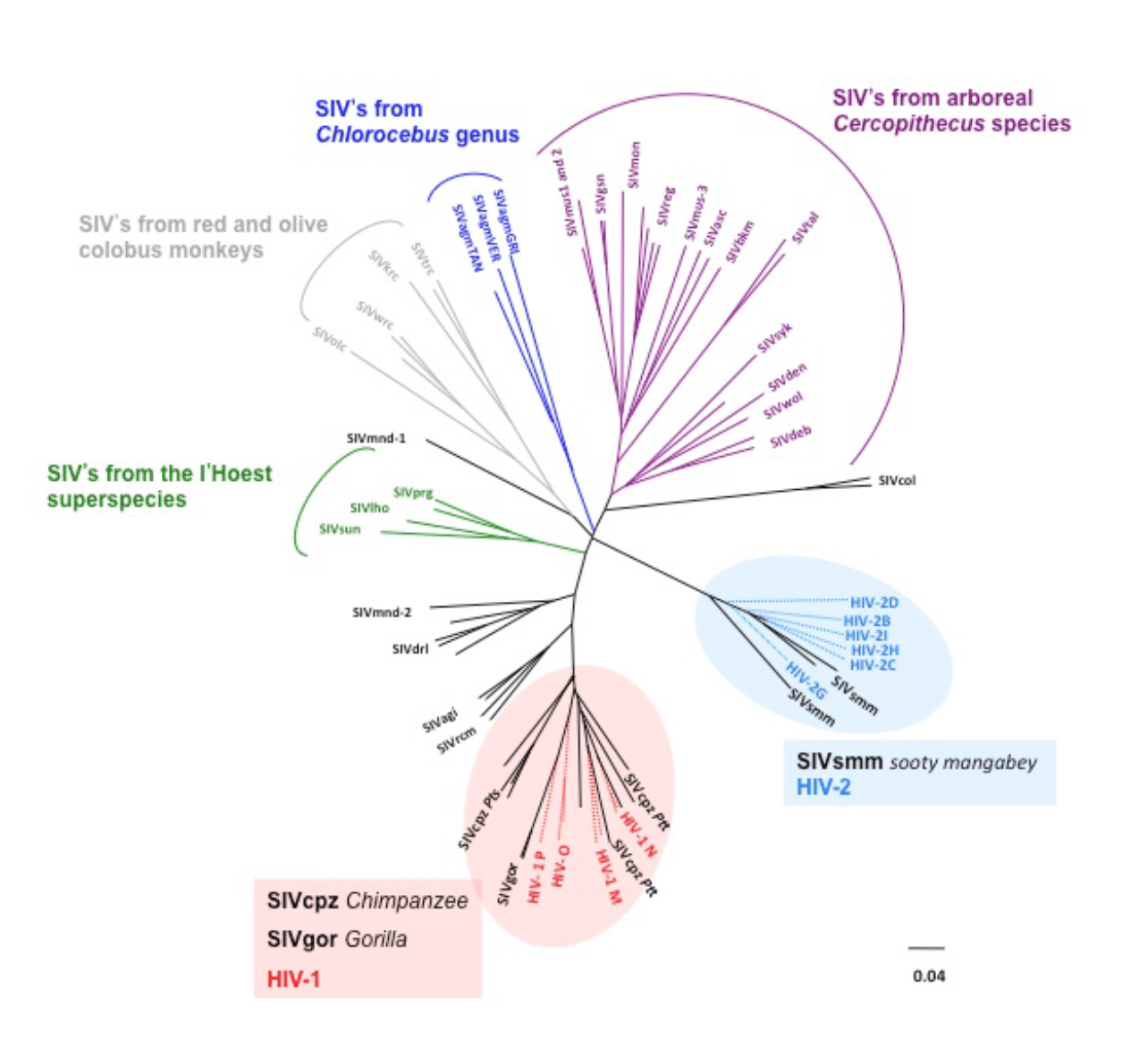
Quickly we set about cloning and sequencing SIVcpz. We used the methodology of the day which was DNase fragmentation, cloning in M13 phage dsDNA and dideoxy sequencing of the ssDNA matrices. The only difference was the use of fluorescent dideoxynucleotides as opposed to using 32P-labelled dATP and later 35S-labelled dATP. These technical transitions were fun to have lived through. While SIVcpz was indeed divergent it was completely isogenic with HIV-1 (15). And there it was; HIV’s closest ancestor was in chimps, just as our closest ancestor is the chimp. Adaptation wasn’t difficult. Obviously, isolation of the virus was the key step. But HIV virology was powering on and genetic methods were becoming de rigueur. Phylogenic trees sprung out of everywhere while PCR could be used to study viruses in uncultured samples and provided a means to identify lookalike viruses. Today there are more than 45 natural SIVs in non-human primates.
HIV/AIDS was unusual for a myriad of reasons; everything was new while there were no reference points and with them the cluster of experts who dominated the field. There was room for all and anybody and the younger generation with their command of molecular techniques ran riot.
With hindsight it does seem odd that zoonoses were not mentioned earlier on, but in fairness, the shock of a new and deadly virus in the population, blood and blood products was enough to concentrate minds. Diagnostics above all, followed by drugs and vaccines were top of the agenda. Origins and animals would wait.
For me, the visna lentivirus sequence provided the biggest shock of all (10) revealing the deadly nature of HIV. Dark days followed. The biomedical community came up with tritherapy by 1995 and from there on things have been getting better – a triumph of modern medicine. The one black spot left is the absence of any efficient vaccine after 35 years of trying.
We still don’t have good vaccines for TB and malaria after many more years you will retort. We have the firepower, more is not needed. We need to encourage and nurture thoughtful and imaginative young researchers who do not think like their peers. They may be quiet types who don’t fare well in the harsh and competitive peer review process. But please, were talking of human disease not a career system!
References
1. Stehelin et al., Nature 260:170 (1976)
2. Poeisz et al., PNAS 77:7415 (1980)
3. Yoshida et al., PNAS 79:2031 (1982)
4. Kalyanaraman et al., Science 218:571 (1982)
5. Barré-Sinoussi et al., Science 220:868 (1983)
6. Wain-Hobson et al., Cell 40:9 (1985)
7. Montagnier et al., Ann Inst Pasteur 135:119 (1984)
8. Gonda et al., Science 227:173 (1985)
9. Wain-Hobson et al., Nature 313:743 (1985)
10. Sonigo et al., Cell 42:369 (1985)
11. Daniel et al., Science 228:1201 (1985)
12. Clavel et al., Nature 324:691 (1986)
13. Guyader et al., Nature 326:662 (1986)
14. Peeters et al., AIDS 3:625 (1989)
15. Huet et al., Nature 345:356 (1990) - poster image is Figure 1 from this paper
Follow the Topic
-
Nature
A weekly international journal publishing the finest peer-reviewed research in all fields of science and technology on the basis of its originality, importance, interdisciplinary interest, timeliness, accessibility, elegance and surprising conclusions.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
very interesting and informative perspective but let us not call chimpanzees our nearest “ancestors “ just our nearest relatives.