Benchmarking algorithms for single-cell multi-omics prediction and integration
Published in Protocols & Methods, Computational Sciences, and Genetics & Genomics
Behind the paper
Our journey began when we attempted to use multi-omics prediction algorithms to analyze our scRNA-seq data. We found that the predictions for chromatin accessibility were unsatisfactory, and the accuracy of protein predictions varied significantly. Despite extensive efforts, we were unable to improve the overall prediction accuracy. We also tested many existing tools but encountered similar performance issues. As new single-cell multi-omics prediction algorithms continued to emerge, it became increasingly difficult to choose the best tool from the numerous available options. We realized that this challenge was likely shared by other research groups, prompting us to conduct a benchmark study to evaluate the existing prediction tools. Our goal was to help users select the best-performing tools while highlighting the current challenges in the field.
During this process, we identified another critical challenge in single-cell multi-omics data analysis: the need for robust and efficient computational strategies for data integration. Many algorithms had been developed for this purpose, which also required systematic evaluation. After discussing with the handling editor of our manuscript, we decided to incorporate evaluations of both single-cell multi-omics prediction and integration algorithms into our study. This comprehensive approach aimed to address the key challenges in single-cell multi-omics analysis.
The problem
Recent advances in single-cell multi-omics technologies have enabled the simultaneous detection of multiple types of genomic information within individual cells. This capability allows us to explore complex gene regulation mechanisms at the single-cell level and predict cellular fates with greater precision. However, due to high experimental costs and technical challenges, the adoption of single-cell multi-omics technologies remains limited compared to single-omics methods. This has created a critical need for predictive algorithms that can infer comprehensive multi-omics information from single-cell data. To address this gap, researchers have developed various multi-omics prediction algorithms, such as totalVI, scArches, and LS_Lab, which can predict protein abundance and chromatin accessibility from scRNA-seq data.
Another major challenge in single-cell multi-omics analysis is the need for robust and efficient computational strategies for data integration. This includes developing algorithms for specific tasks, such as associating different omics modalities (vertical integration) and correcting batch effects across multi-omics datasets (horizontal integration). Additionally, some algorithms focus on integrating single-cell datasets that share at least one type of omics information, known as mosaic integration.
While these prediction and integration algorithms have substantially advanced our understanding of biological and pathological processes, few studies have systematically compared their performance. This highlights the need for comprehensive evaluations to determine their effectiveness and applicability across diverse tasks.
The solutions
We developed a framework to systematically benchmark fourteen algorithms for predicting chromatin accessibility or protein abundance from scRNA-seq data using 36 multi-omics datasets and six evaluation metrics (Fig. 1). Additionally, we benchmarked eighteen algorithms designed for vertical, horizontal, or mosaic integration using 35 single-cell multi-omics datasets and ten evaluation metrics (Fig. 1). Our study also evaluated the performance of these algorithms using different data batches for training and prediction, as well as the computational resources consumed by each algorithm.
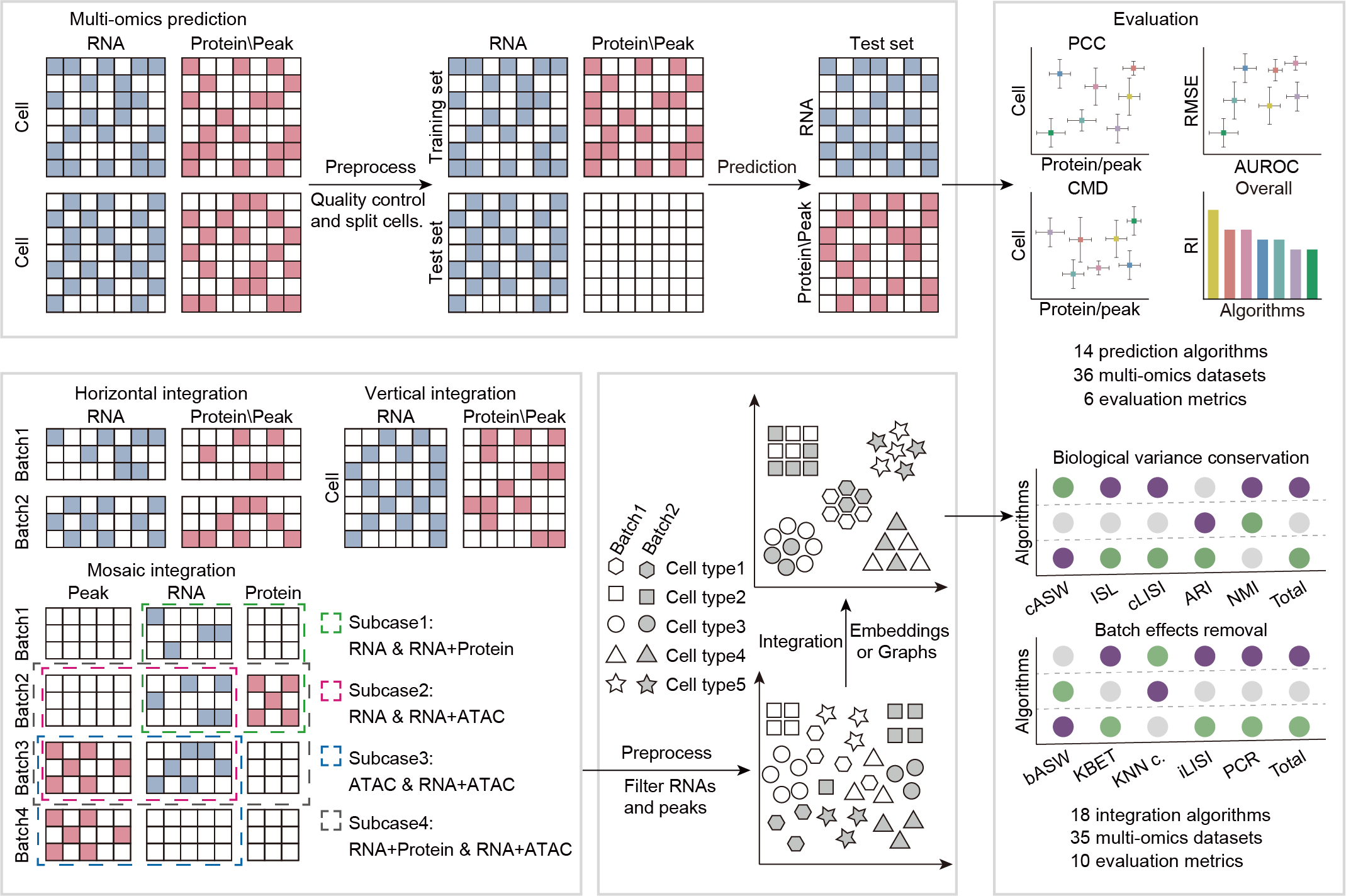
Fig. 1: Workflow for benchmarking. Benchmarking workflow for fourteen multi-omics prediction algorithms and eighteen multi-omics integration algorithms, including eleven algorithms that can predict protein abundance, nine algorithms that can predict chromatin accessibility, fifteen algorithms for vertical integration, nine algorithms for horizontal integration, and eight algorithms for mosaic integration. We adopted six and ten metrics for the multi-omics prediction and integration algorithms, respectively, to evaluate the performance of these algorithms, and we also assessed their robustness and consumed computational resources.
Our benchmark results demonstrated that totalVI and scArches outperformed other algorithms in predicting protein abundance, while LS_Lab was the most effective for predicting chromatin accessibility (Fig. 2a). For multi-omics data integration, Seurat, MOJITOO, and scAI were the top performers in vertical integration (Fig. 2b), while totalVI and UINMF excelled in both horizontal and mosaic integration scenarios (Fig. 2c & d). To support researchers, we provide a pipeline for selecting the optimal multi-omics prediction and integration algorithm.
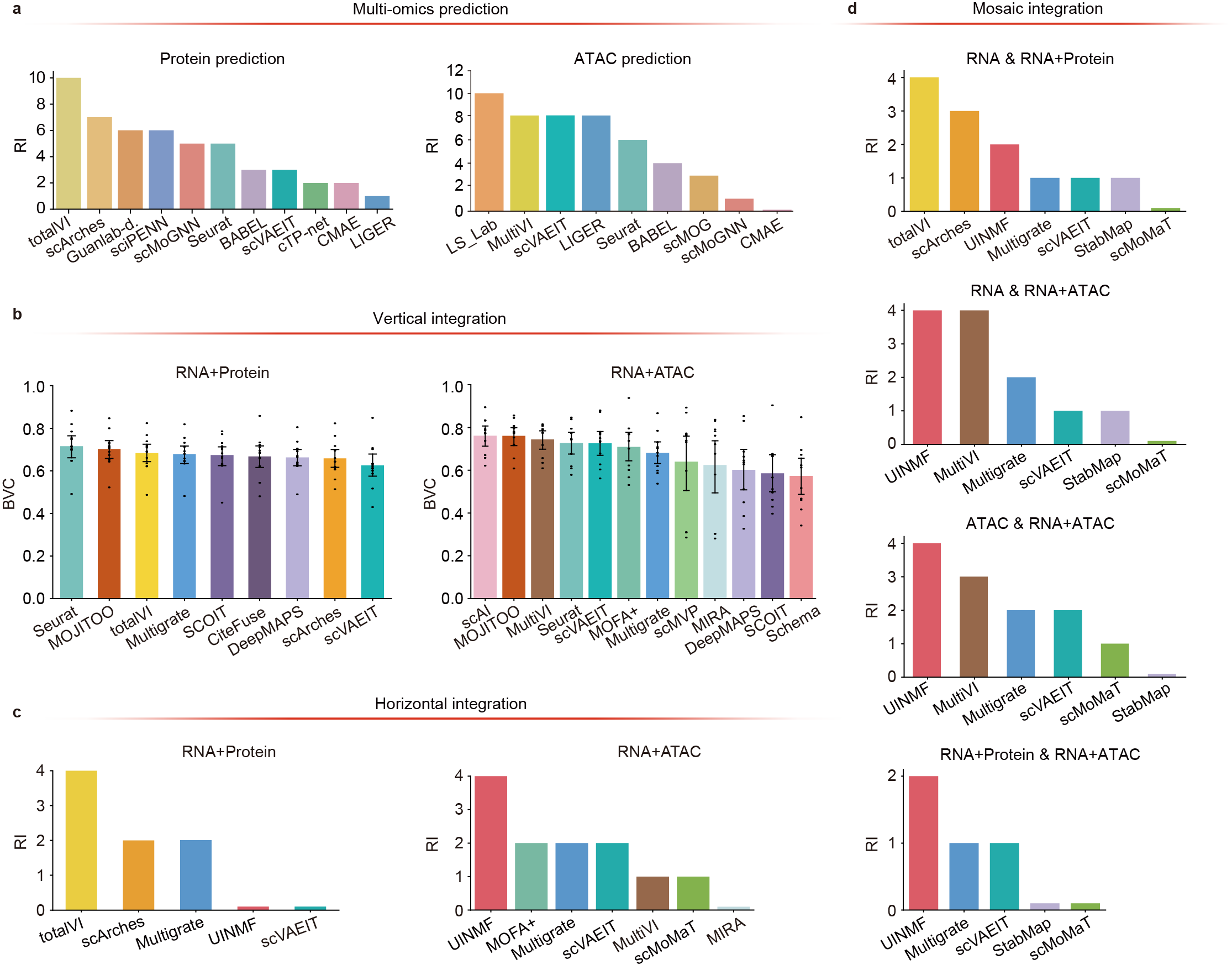
Fig. 2: Performance of prediction and integration algorithms. a,The overall performance of eleven protein abundance prediction algorithms and seven chromatin accessibility prediction algorithms in both intra-dataset and inter-dataset scenarios. b, The overall performance of vertical integration algorithms, including nine algorithms integrating RNA expression with protein abundance and twelve algorithms integrating RNA expression with chromatin information. c, The overall performance of horizontal integration algorithms, including five algorithms focused on integrating multiple RNA+Protein batches and seven algorithms focused on integrating multiple RNA+ATAC batches. d, The overall performance of eight mosaic integration algorithms across four subcases.
Follow the Topic
-
Nature Methods
This journal is a forum for the publication of novel methods and significant improvements to tried-and-tested basic research techniques in the life sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Methods development in Cryo-ET and in situ structural determination
Publishing Model: Hybrid
Deadline: Jul 28, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in