Directing Min protein patterns with advective bulk flow
Published in Physics

Patterns are phenomena that amaze us, not only by their aesthetics but also by how omnipresent they are in biology (and elsewhere in the world, for that matter). A pattern emerges when one or several system constituents do not spread homogeneously over the available space as one might expect. Instead, they accumulate in specific locations to form static or dynamic concentration clusters, the total of which we refer to as a ‘pattern’. [1] While the stripes of a zebra or the spots of a leopard are beautiful outcomes of multicellular pattern formation, we have become aware of several intracellular pattern-forming systems involved in the coordination of vital cellular processes. Examples are the protein concentration patterns that play essential roles in directing basic phenomena of life, such as cell division, polarity, and motility. [2-4]

The most-studied example of intracellular pattern formation is the Min protein system of the bacterium Escherichia coli (E. coli). Here, two proteins – called MinD and MinE – cyclically bind and unbind at the bacterium’s inner membrane, as determined by their specific interactions with each other and the membrane. Combined with diffusive transport, these interactions result in pole-to-pole oscillations. This spatiotemporal pattern helps position E. coli’s cell division machinery in the middle of the rod-shaped cells. As it requires only two proteins, the Min system is considered a minimal and paradigmatic model system for biological pattern formation. [5]
To better understand how these proteins function, we can take them out of their natural habitat and study them in a reasonably well-controlled environment, the parameters of which we can set. In such in vitro studies, a glass slide is coated with an artificial lipid membrane to which the proteins can bind. Supplied with ATP (the adenosinetriphosphate molecule which provides energy required for biochemical reactions), MinD and MinE form magnificent membrane patterns, such as travelling waves or spirals. Under the well-controlled conditions of the in vitro assay, we can systematically study these patterns and, in combination with theoretical studies, gain insight into the underlying mechanisms. [5,6] As we gradually tweak and extend our models to achieve better agreement between what we see in microscope images and simulation results, we improve our understanding of how this system works – and thus generalize some of our findings to biological pattern formation. [2,4]
Protein patterns arise from the interplay of chemical reactions and the spatial transport of proteins; that is, they are a reaction-diffusion system – as famously pioneered by Alan Turing. [7] Depending on the system in various organisms, transport can occur either passively (through diffusion) or actively (through flows). Unlike diffusion, transport by a flow has a preferred spatial direction. So far, little research has been conducted on the influence of fluid flow on protein patterns. [8] The in vitro Min-protein system is an ideal model to study this flow effect on pattern formation and test predictions of the theoretical models. [9] Combining the theory by the Frey group (LMU Munich) and experiments performed by the Dekker group (TU Delft), we found that flow can reveal system properties that might otherwise not become apparent.
This project started in 2020 with theoretical investigations and numerical simulations of Min protein patterns under flow performed by Fridtjof Brauns and Jernej Finžgar, in the group of Erwin Frey. To find out if the simulation results would compare to experimental data, they contacted the lab of Cees Dekker, with whom the team had collaborated before. [10] In 2021, as microscopy labs were more easily accessible again, the simulations were complemented by experiments with Min proteins in flow cells. Along the way, a data analysis pipeline was designed that could be used to quantify the protein patterns’ (sometimes ambiguous) response to flow. [11] Notably, this collaboration stretched not only over a global pandemic but also over job changes for some of us. With three shared first authors spread over the Netherlands, Germany, and California, we had to carefully schedule our numerous intercontinental discussion sessions. In light of these challenges, we are even happier to be able to share our findings here.
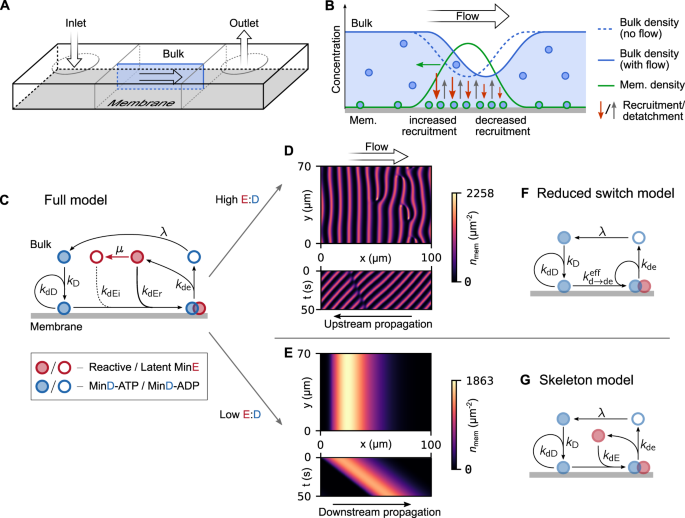
In simulations as well as experiments with Min patterns in flow, we could show that laminar flow causes alignment of the membrane-bound protein patterns and propagation predominantly occurring along the bulk flow’s direction. Strikingly, the patterns can move either with the flow (downstream) direction or against it (upstream). This latter scenario might appear counterintuitive at first, as intuition would tell us that patterns, like anything, could be expected to move along with the flow.
However, bulk flow does not directly affect the surface-bound proteins but only those in bulk. Bulk and surface are coupled by the exchange (attachment and detachment) of proteins, By systematically varying experimental parameters, we found that the propagation direction depends on the ratio of protein concentrations (i.e., the MinE versus MinD concentration ratio), with an overall good qualitative agreement between simulations and experiments. Theoretical modeling further revealed that the patterns’ direction of movement subtly depends on specific interactions between the Min proteins.
Despite many advances in recent years, the Min system still presents many unresolved puzzles. For instance, while theoretical models successfully reproduce many experimental observations qualitatively, we keep encountering a quantitative mismatch between simulation and experimental outcomes. This hints that our understanding of the molecular mechanisms driving the Min-protein patterns is still incomplete, with some hidden mechanisms. Extending our theoretical model to account for such details and exploring their effect on the patterns is an ongoing effort.
As we see it, fluid flows provide a versatile tool for controlling protein patterns and investigating molecular mechanisms of pattern formation – not just for the Min system. Combined for example with the Min protein system’s demonstrated capacity for cargo transport,[12] it could also open up engineering opportunities for aligning and directing surface-bound components. But perhaps even more importantly, from a scientific point of view, our work shows that even after decades of research, we are still far from knowing everything there is to know, even about such a “simple” pattern-forming system as the Min protein system.
References
- Schweisguth, F. & Corson, F. Self-Organization in Pattern Formation. Developmental Cell 49, 659–677 (2019).
- Halatek, J., Brauns, F. & Frey, E. Self-organization principles of intracellular pattern formation. Phil. Trans. R. Soc. B 373, 20170107 (2018).
- Merino-Salomón, A., Babl, L. & Schwille, P. Self-organized protein patterns: The MinCDE and ParABS systems. Current Opinion in Cell Biology 72, 106–115 (2021).
- Burkart, T., Wigbers, M. C., Würthner, L. & Frey, E. Control of protein-based pattern formation via guiding cues. Nat Rev Phys (2022) doi:10.1038/s42254-022-00461-3.
- Ramm, B., Heermann, T. & Schwille, P. The E. coli MinCDE system in the regulation of protein patterns and gradients. Cell. Mol. Life Sci. 76, 4245–4273 (2019).
- Mizuuchi, K. & Vecchiarelli, A. G. Mechanistic insights of the Min oscillator via cell-free reconstitution and imaging. Phys. Biol. 15, 031001 (2018).
- Turing, A. The chemical basis of morphogenesis. Phil. Trans. R. Soc. Lond. B 237, 37–72 (1952).
- Vecchiarelli, A. G., Li, M., Mizuuchi, M. & Mizuuchi, K. Differential affinities of MinD and MinE to anionic phospholipid influence Min patterning dynamics in vitro: Flow and lipid composition effects on Min patterning. Molecular Microbiology 93, 453–463 (2014).
- Denk, J. et al. MinE conformational switching confers robustness on self-organized Min protein patterns. Proc Natl Acad Sci USA 115, 4553–4558 (2018).
- Brauns, F. et al. Bulk-surface coupling identifies the mechanistic connection between Min-protein patterns in vivo and in vitro. Nat Commun 12, 3312 (2021).
- Meindlhumer, S., Kerssemakers, J. & Dekker, C. Quantitative analysis of surface wave patterns of Min proteins. Front. Phys. 10, 930811 (2022).
- Ramm, B. et al. A diffusiophoretic mechanism for ATP-driven transport without motor proteins. Nat. Phys. 17, 850–858 (2021).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in