Exploring the dark side of the immune microenvironment in Follicular Lymphoma
Published in Cancer
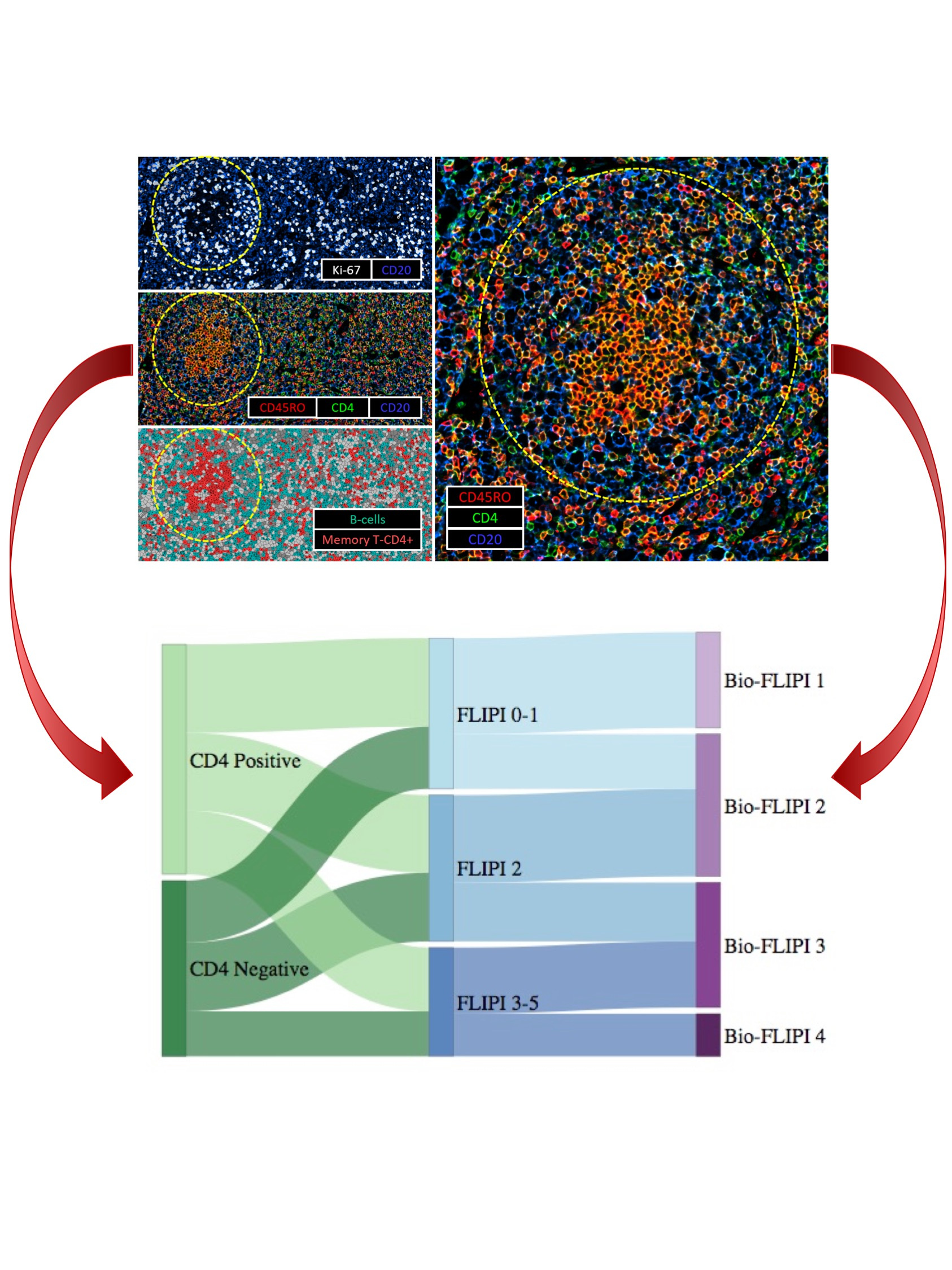
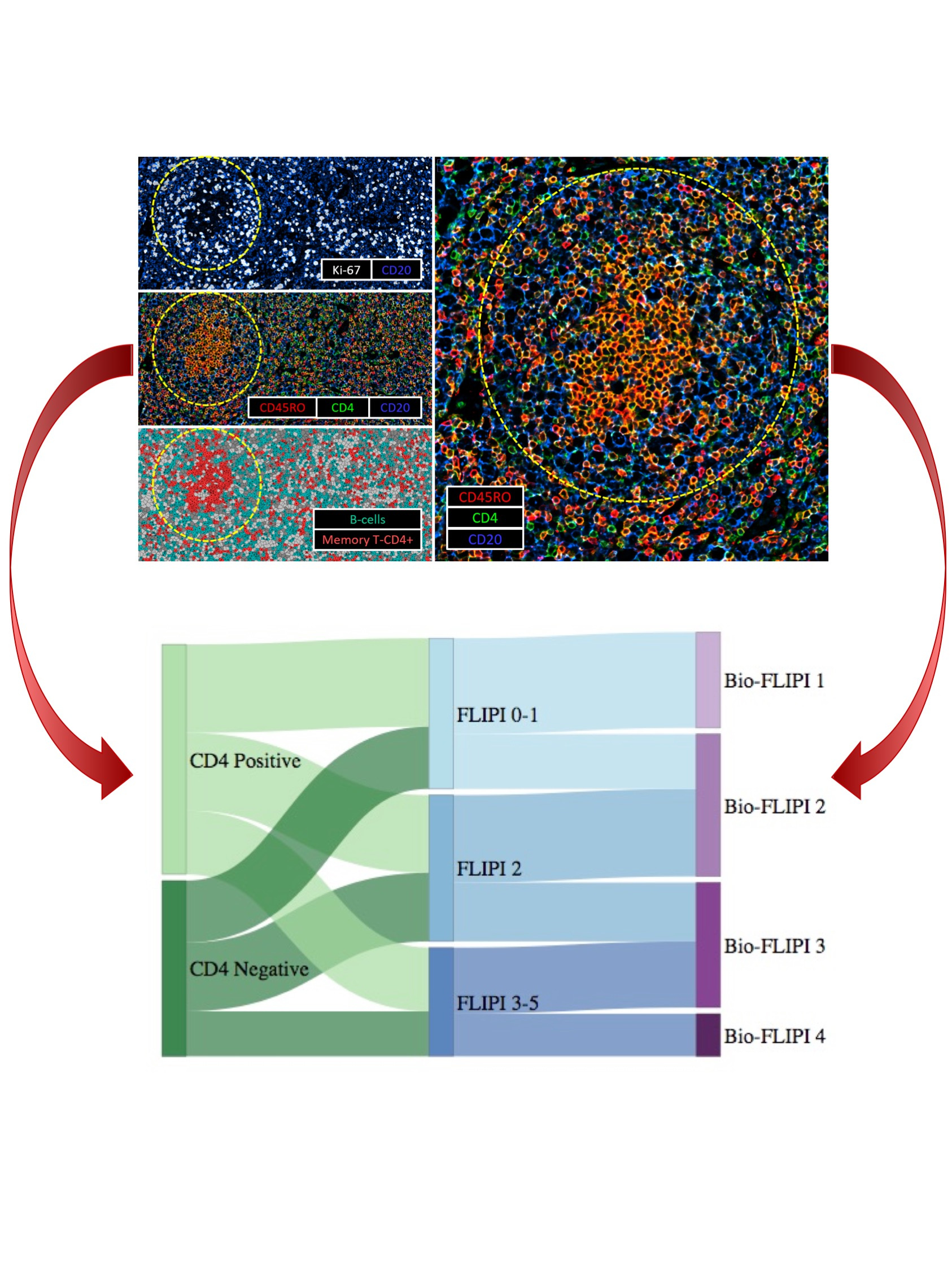
Despite a characteristic indolent course, a substantial subset of follicular lymphoma (FL) patients have an early relapse with a poor outcome. Several prognostic scoring systems have been developed to predict outcome in patients with FL. However, none of them can predict who will fail event-free-survival at 12 or 24 months (EFS12/EFS24), hence they are not routinely used to guide therapeutic decisions.1–3 Gene expression profiling of FL has demonstrated the importance of the tumor microenvironment in predicting disease progression and survival.4 Several immunohistochemical studies attempted to translate these findings to clinical practice by identifying T-cell subsets associated with outcome, however mixed and inconclusive results have been reported.5–9 One potential explanation for this discrepancy might be the evaluation of total, rather than cells in specific locations, such as intra- or peri-follicular biomarker expression. In particular, the immune architectural pattern of T cells within lymph nodes involved with FL has a strong clinical impact, since the topographic distribution of the immune cells reflects their dysregulated function. Based on this assumption, we hypothesized that phenotype and distribution of the immune cells in the tumor microenvironment would predict early failure.
To address this hypothesis, we used a discovery and validation study design to analyze the prevalence of T-cells subsets and macrophages in the pretreatment biopsy specimens of 496 newly diagnosed patients with FL grade 1-3A who were prospectively enrolled into the Molecular Epidemiology Resource (MER) cohort. To explore the relevance of immune cell distribution on the interaction with malignant B-cells, we specifically determined the expression of immune markers inside and outside the malignant follicles. Initially, we stained tissue microarrays of a discovery set of patients (N=166). Markers associated with failing to achieve EFS12/24 at P≤0.15 were then brought forward to validate the positive results in a second cohort of patients (N=330). Then, the immune population associated with outcome was immunophenotypically characterized and spatially located in the follicle using mass cytometry (CyTOF) and the Co-Detection by indEXing (CODEX) multiplex immunofluorescence system. Finally, the influence of the tumor genetic landscape on the microenvironment was assessed by digital multiplex gene expression profiling (NanoString technology) platform on the same matched patients.3
We demonstrated the prognostic value of CD4+ T-cells specifically located within the follicle in close proximity to other T-cells. The intrafollicular CD4+ T-cell population associated with favorable outcome was immunophenotypically characterized as activated, non-exhausted central memory T-cells (TCM), which play a critical role in the immune surveillance of peripheral tissues. Lack of activated TCM cells may disrupt immune surveillance, enabling immune escape, which in turn allows FL B-cells to persist, facilitating lymphoid proliferation and transformation. Tumors accordingly manifest more aggressive features, thus providing the rationale for considering CD4+ activated TCM cells as an independent prognostic biomarker.
We have leveraged the characterization of the immune microenvironment of follicular lymphoma to suggest a novel predictive model, termed BioFLIPI, which integrates biologic and clinical features. By adding the intrafollicular CD4+ expression to the well-established clinical risk model, FLIPI, we developed an improved prognostic algorithm that can help to guide therapeutic decisions for newly diagnosed FL patients. Favorable-risk groups showed an indolent disease course and these patients therefore might be observed or receive lower-intensity approaches, while high-risk groups had an early relapse which suggests a need for more intensive treatment, or for clinical trials with novel agents, and consideration for maintenance therapy. Although the BioFLIPI demonstrated superior prognostic value than the FLIPI, 44% of the two Bio-FLIPI high risk groups still achieve EFS24, suggesting that additional events influence the distinct biological behavior.
While our model underlines the importance of tumor microenvironment, we reason that it may be influenced by the genetic composition of tumor. For example, the highly recurrent CREBBP mutation has been associated with reduced expression of antigen presentation machinery and impaired immune surveillance.10 Along the same lines, it is possible that CREBBP mutations might lead to a lack of intrafollicular CD4+ expression as part of the same immune escape mechanism. Remarkably, we found that CD4+ expression, BioFLIPI and the genetic risk model 23-GEP scores were independently prognostic. This highlights that the tumor genetic features and the CD4+ T-cell infiltration are both factors of importance for the outcome of FL patients. However, future studies will be needed to standardize the assessment of intrafollicular CD4+ expression, validate the prognostic value of the BioFLIPI risk model in independent patient cohorts, and integrate tumor genetic features into this biological model.
Conclusion
The BioFLIPI represents a promising predictor of treatment outcome in newly diagnosed FL patients that incorporates biological and clinical features, but it will need independent validation and eventually combination with genomic factors. While additional investigation to determine the mechanisms behind the reduced or absent expression of CD4+ T-cells inside the lymphoma follicles is warranted, the prognostic power of the BioFLIPI will be useful in the design of clinical trials as it identifies patients at the highest risk of early failure who may benefit most from more intensive therapies or novel frontline regimens.
References
1 Solal-Céligny P, Roy P, Colombat P, White J, Armitage JO, Arranz-Saez R et al. Follicular lymphoma international prognostic index. Blood 2004; 104: 1258–65.
2 Pastore A, Jurinovic V, Kridel R, Hoster E, Staiger AM, Szczepanowski M et al. Integration of gene mutations in risk prognostication for patients receiving first-line immunochemotherapy for follicular lymphoma: a retrospective analysis of a prospective clinical trial and validation in a population-based registry. Lancet Oncol 2015; 16: 1111–1122.
3 Huet S, Tesson B, Jais J-P, Feldman AL, Magnano L, Thomas E et al. A gene-expression profiling score for prediction of outcome in patients with follicular lymphoma: a retrospective training and validation analysis in three international cohorts. Lancet Oncol 2018; 19: 549–561.
4 Dave SS, Wright G, Tan B, Rosenwald A, Gascoyne RD, Chan WC et al. Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N Engl J Med 2004; 351: 2159–69.
5 Carreras J, Lopez-Guillermo A, Roncador G, Villamor N, Colomo L, Martinez A et al. High numbers of tumor-infiltrating programmed cell death 1-positive regulatory lymphocytes are associated with improved overall survival in follicular lymphoma. J Clin Oncol 2009; 27: 1470–6.
6 Farinha P, Masoudi H, Skinnider BF, Shumansky K, Spinelli JJ, Gill K et al. Analysis of multiple biomarkers shows that lymphoma-associated macrophage (LAM) content is an independent predictor of survival in follicular lymphoma (FL). Blood 2005; 106: 2169–74.
7 Farinha P, Al-Tourah A, Gill K, Klasa R, Connors JM, Gascoyne RD. The architectural pattern of FOXP3-positive T cells in follicular lymphoma is an independent predictor of survival and histologic transformation. Blood 2010; 115: 289–95.
8 Glas AM, Knoops L, Delahaye L, Kersten MJ, Kibbelaar RE, Wessels LA et al. Gene-expression and immunohistochemical study of specific T-cell subsets and accessory cell types in the transformation and prognosis of follicular lymphoma. J Clin Oncol 2007; 25: 390–8.
9 Canioni D, Salles G, Mounier N, Brousse N, Keuppens M, Morchhauser F et al. High numbers of tumor-associated macrophages have an adverse prognostic value that can be circumvented by rituximab in patients with follicular lymphoma enrolled onto the GELA-GOELAMS FL-2000 trial. J Clin Oncol 2008; 26: 440–6.
10 Mondello P, Tadros S, Teater M, Fontan L, Chang AY, Jain N et al. Selective inhibition of HDAC3 targets synthetic vulnerabilities and activates immune surveillance in lymphoma. Cancer Discov 2020; : CD-19-0116.
Follow the Topic
-
Blood Cancer Journal
This journal seeks to publish articles of the highest quality related to hematologic malignancies and related disorders.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in