Going beyond the microscope to study the spatial orchestration of proteins on single cells
Published in Physics, Cell & Molecular Biology, and Immunology

A few years back, we assembled an interdisciplinary team of scientists and engineers to tackle the problem of scalability and throughput of spatial analyses of biomolecules. As microscopes are limited by light and the taking of pictures in 2D, we felt it was a field with room for disruptive innovation. The microscope was invented 350 years ago and discovered the existence of cells and microbial life. Subsequently it gave the scientific community all its insights of how life operates at the cellular level. From then onward, massive improvements have been made in resolution and signal strength of microscopes. However, these methods are still based on light and the use of various wavelengths to see the objects of study, just like the naked eye.
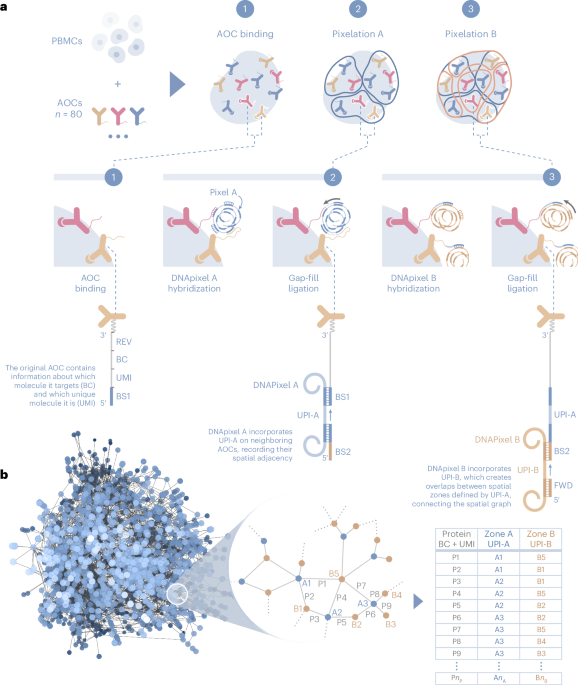
Having previously developed DNA-enabled protein detection for plasma with the Proximity Ligation/Extension Assays (PLA and PEA), the team understood the theoretical potential in employing DNA sequences as reporter systems to achieve scalability. But could we use the same philosophy for locating the relative positions of cellular biomolecules starting with the surface of cells? What would a GPS navigation-like system for localizing proteins and other biomolecules enable? The project we envisioned would be a challenge in not only immuno-detection by DNA-tagged antibodies, but also in molecular biology techniques of manipulating synthetic DNA and algorithms to analyze data generated by high throughput sequencing. This highly interdisciplinary project could deliver a next generation tool for spatial analyses. The thinking was based on the fact that there are a limited number of colors to choose from when doing microscopy but with DNA, the number of unique identifiers for targets and positions is endless. We attempted to locate the DNA-tagged antibodies using our DNA-pixels which are uniquely sequence encoded DNA molecules of nanometer size. We used a serial reaction to effectively perform colocalization twice with pixel overlap to enable a complete spatial pattern per cell surface, targeting 80 immune cell surface proteins.
In the early phase of the project an application for a grant from Wellcome Leap was approved and we gratefully received their support in building this technology. Even though there was very little supporting data at the time of the application.
As the first results were generated from the method which we had begun to call Molecular Pixelation (MPX), we immediately realized the data was a three dimensional spatial graph, ready to be mined by spatial statistics. Each cell became represented as a distinct graph even when we had all the cells free in the same suspension. It was a thrill explaining it to others as a method that takes a picture of thousands of cells in 3D, from all angles at once, always in focus, with high resolution, by just mixing reagents in a standard PCR-type test tube. No more countless hours spent looking into a microscope in a dark room with samples stuck to a glass slide. It was truly a WOW moment to see the first data ever plotted by a computer from a chemistry generated “image” of the cell surface based on protein locations. The team gathered around the monitor for a traditional photo opportunity to commemorate. MPX could bring omics-type research capability to a 3D spatial context and we’d taken the first steps.
We applied MPX to various research questions in immunology and found that it correlated well with standard microscopy and the spatial graph even reflected the cell shape and size. Objective spatial metrics for each protein of non-randomness of distribution (polarity score) and colocalization scores were developed. We wondered how the research community would receive a “chemical” image of a cell not based on photons and discussed the philosophical question whether MPX-data is an “image” at all, or just a graph representation of spatial proteomics data…?
As human beings studying the world around us, seeing is believing, and we mostly observe by seeing shapes and different wavelengths of photons. We are all drawn to pictures. Just think of the beautiful images generated of the universe by the space telescopes. They look really striking, but they don't directly teach us about the laws of physics and how the universe works. The real science lies in the data analyzed by computers and not by the human naked eye. The space images are pseudo colored by graphics artists to inspire us.
As our MPX project progressed, the data analysis team was challenged to visualize the results by presenting each protein as a color in the spatial graph. This became a visual mess, as our eyes can not differentiate between that many colors (80) in the same object at once, really highlighting the need for computational analysis instead of human visual observation. But we still need cool images to showcase our work. At the end of the day, we are still human beings :-)
We are very happy to now introduce Molecular Pixelation to the research community with our Nature Methods publication and look forward to its implementation and further refinement by following its exciting potential applications and development trajectory, beyond the limits of light.
Many thanks to the incredible development team, early users of MPX, and supporters of the project.
Follow the Topic
-
Nature Methods
This journal is a forum for the publication of novel methods and significant improvements to tried-and-tested basic research techniques in the life sciences.
Ask the Editor - Immunology, Pathogenesis, Inflammation and Innate Immunity
Got a question for the editor about the complement system in health and disease? Ask it here!
Continue reading announcementRelated Collections
With Collections, you can get published faster and increase your visibility.
Methods development in Cryo-ET and in situ structural determination
Publishing Model: Hybrid
Deadline: Jul 28, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in