GRA45, the “blacksmith” assembling the armor of effectors defending Toxoplasma gondii against IFNγ attacks
Published in Microbiology
The eukaryotic pathogen Toxoplasma gondii can establish an infection in all nucleated cells from almost all warm-blooded animals. To defend themselves against infection, these animals produce the cytokine interferon-gamma (IFNγ), which rallies an army of interferon-stimulated genes including Immunity-Related GTPases (IRGs) and Guanylate-Binding Proteins (GBPs), which function as spears to pierce Toxoplasma’s replication niche - the parasitophorous vacuole (PV)-. Toxoplasma counteracts these IFNγ attacks by building a protective armor consisting of parasite effectors secreted from its unique organelles, rhoptries and dense granules (secreting ROPs and GRAs, respectively), on the PV membrane (PVM). Although some of these Toxoplasma secretory effectors were previously identified1, 2, most of the effectors counteracting the attacks from IFNγ were largely unknown. Identifying such Toxoplasma effectors and understanding their role in building the protective armor will help to forge “armor breaker” therapies against Toxoplasma.
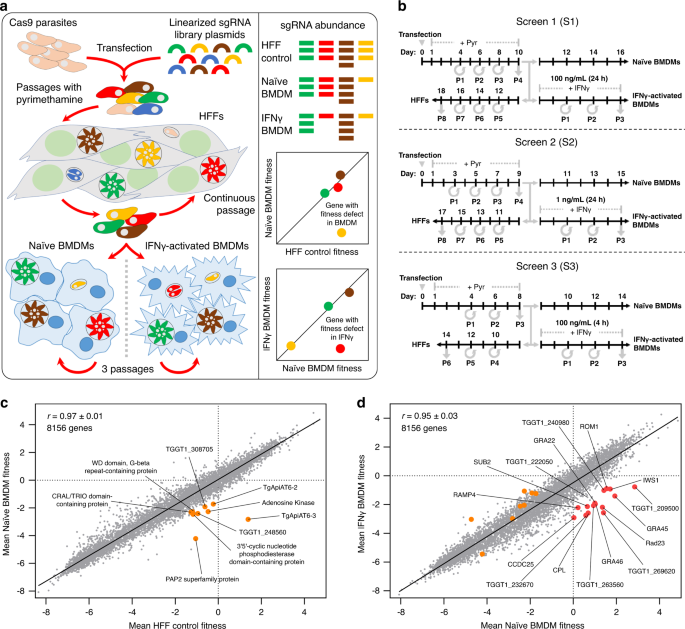
Our group has always been interested in discovering novel Toxoplasma effectors involved in various steps of host cell modulation. In this study, collaborating with the Lourido laboratory at the Whitehead Institute, we applied a genome-wide CRISPR/Cas9 loss-of-function screen3 to determine which of Toxoplasma’s 8,000 protein-coding genes could function as a potential “armor plate”. With the contributions from four postdoctoral researchers in the group, we identified several Toxoplasma effector-encoding genes determining IFNγ resistance in primary murine macrophages. Among seven confirmed Toxoplasma genes, we further investigated one gene encoding dense granule protein GRA45, which plays a conserved role in defending against IFNγ attacks in macrophages from different host species, such as mice, rats, and humans. However, we found that GRA45 is unlikely to be the piece of “armor plate” covering the PVM because it only localizes inside the PV when discharged from dense granules.
Intriguingly, we noticed that GRA45 has structural homology to bacterial small heat shock protein (sHSP), suggesting it might act as a chaperone to maintain the correct folding and trafficking of other proteins. Searching the literature for Toxoplasma secretory protein trafficking led us to realize that most of the hydrophobic domain-containing GRAs have a wired trafficking route and after release from dense granules they specifically integrate into the PVM instead of into the parasite plasma membrane (PPM). Based on this, we discovered that GRA45 is critical for the correct localization of several GRAs into the PVM, revealing that GRA45 is the “blacksmith” assembling the armor on the PVM. When GRA45 loses its “hammer”, the conserved sHSP motifs, the armor-built GRAs on the PVM are disorganized and Toxoplasma fails to defend itself against IFNγ attacks.
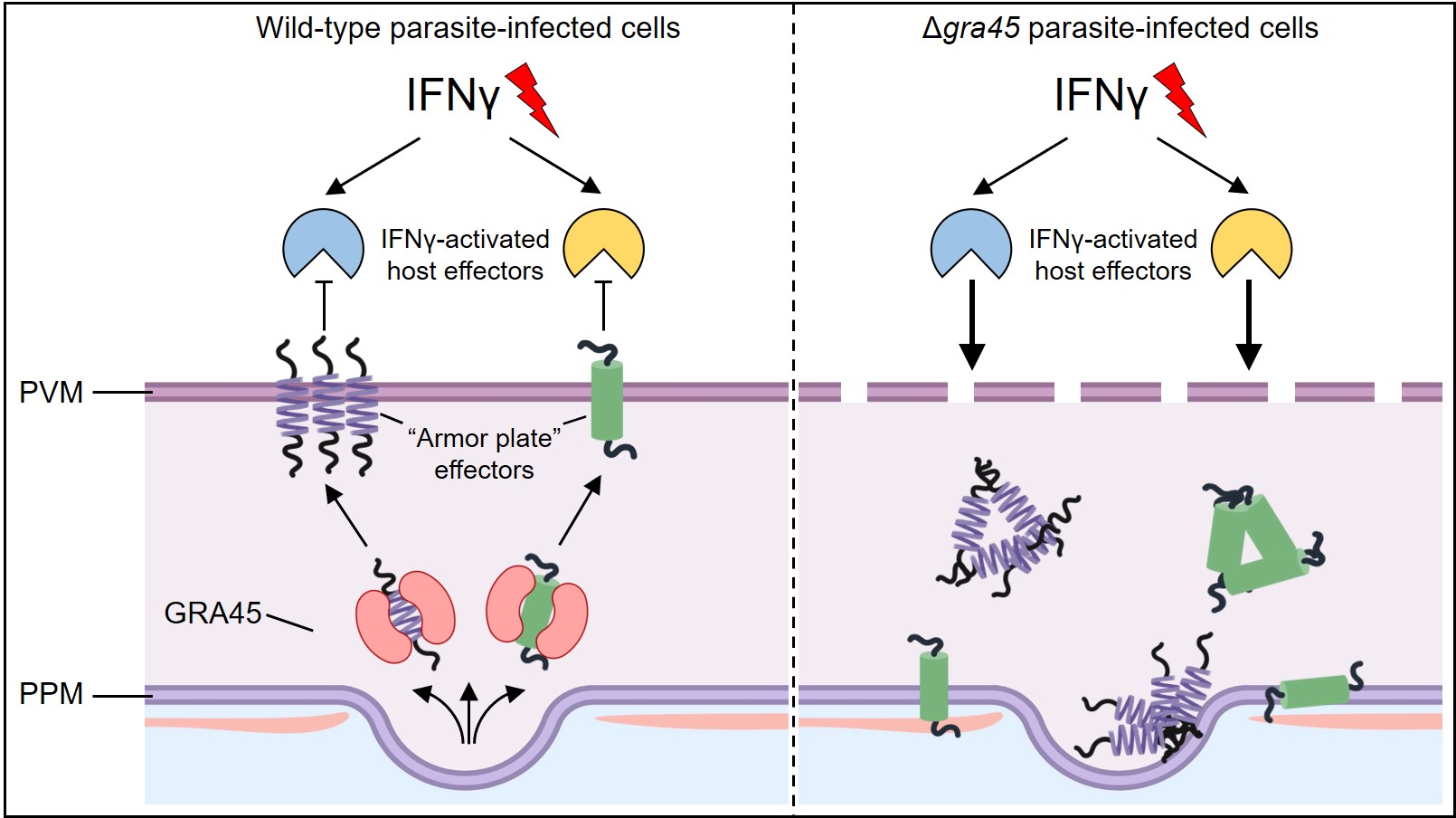
A model of wild-type or Δgra45 parasites counteracting IFNγ attacks.
In sum, the study identifies that GRA45 is an important “blacksmith” GRA forging the IFNγ-resistant armor for Toxoplasma but also providing a foundation for many future analyses of Toxoplasma pathogenicity. These findings also raise new questions such as: 1) what are the armor-built GRAs forged by GRA45? 2) How does GRA45 assemble the armor on the PVM? 3) what are the exact IFNγ-recruited troops attacking the PVM?
- Hunter CA, Sibley LD. Modulation of innate immunity by Toxoplasma gondii virulence effectors. Nat Rev Microbiol 10, 766-778 (2012).
- Hakimi MA, Olias P, Sibley LD. Toxoplasma Effectors Targeting Host Signaling and Transcription. Clin Microbiol Rev 30, 615-645 (2017).
- Sidik SM, et al. A Genome-wide CRISPR Screen in Toxoplasma Identifies Essential Apicomplexan Genes. Cell 166, 1423-1435 e1412 (2016).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in