Nanopores for single amino acid electrical recognition towards protein sequencing
Published in Bioengineering & Biotechnology
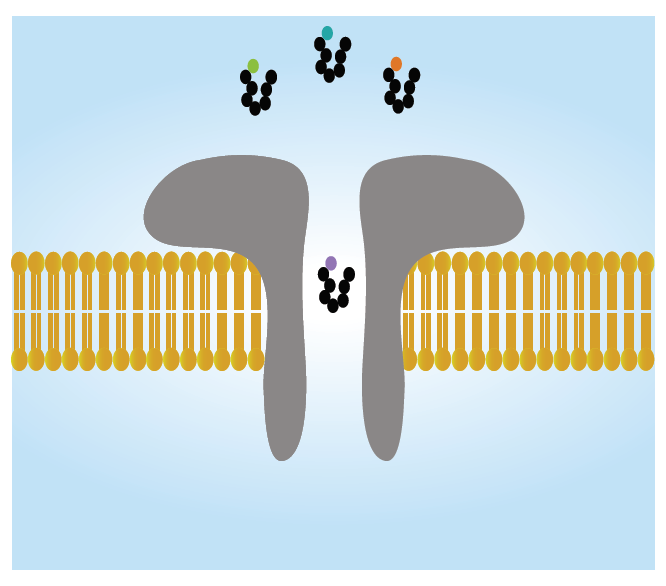
In 2017, I got the opportunity to join the group of Dr. Abdelghani Oukhaled as a PhD student. I met Dr. Oukhaled two years earlier at the University of Cergy-Pontoise as an undergraduate student learning concepts of polymer physics. During his class, Dr. Oukhaled described a single-molecule technology commonly known as nanopore-based electrical detection, that he had been already working with for several years. At the heart of this method is a nanometer-sized hole in a membrane separating two compartments both filled with an ionic solution. Under the action of an electrical field, an ionic current flowing through the nanopore is measured. The entry of a single molecule inside the nanopore causes a current impulsion whose form differs depending on the size, the conformation, and/or the composition of the molecule. I was amazed by the technology and its promise, in particular after seeing that it had already led to the development of a fast and low-cost nanopore-based DNA sequencing device. I applied to join Dr. Oukhaled’s team right away.
At that time, Dr. Oukhaled was particularly interested in using nanopores to develop a new method for protein sequencing, which is the next frontier of the nanopore technology. Development of nanopore-based protein sequencing faces additional challenges related to the structural properties of the proteins. Reading the sequence of a protein require a nanopore capable of distinguishing the twenty electrical signals specific to each amino acid. Several strategies have been employed to meet this challenge. However, up till now, neither biological nor artificial nanopores could provide a sufficient resolution to read proteins with a single amino acid precision.
When I joint his lab, Dr. Oukhaled proposed that I try sequencing proteins using a protein channel---the wild-type aerolysin nanopore. The aerolysin nanopore was a good candidate for protein sequencing because, according to our previous study, it has a superior resolution and sensitivity for the detection of short polycationic homopeptides. However, at this stage of investigations, it was not possible to detect an individual amino acid inside the aerolysin nanopore yet because that would not enter the nanopore of aerolysin by themselves. To address this problem, we came up with the idea of using a polycationic homopeptide, that we knew would be captured by the aerolysin nanopore, as a carrier that would ensure the entry of each individual amino acid inside the nanopore. We quickly ordered the peptide constructs, carried out nanopore translocation experiments and the results were fascinating.
From the get-go, our experiments demonstrate the detection of all twenty proteinogenic amino acids and the identification of at least 13 of the 20 natural amino acids, which was already a great achievement. Among those, to our great surprise, we found that the aerolysin nanopore could discriminate between amino acids of the same molecular mass, such as Leucine and Isoleucine, and between amino acids of very similar chemical structure, such as Phenylalanine and Tyrosine, a resolution which had never been shown before.
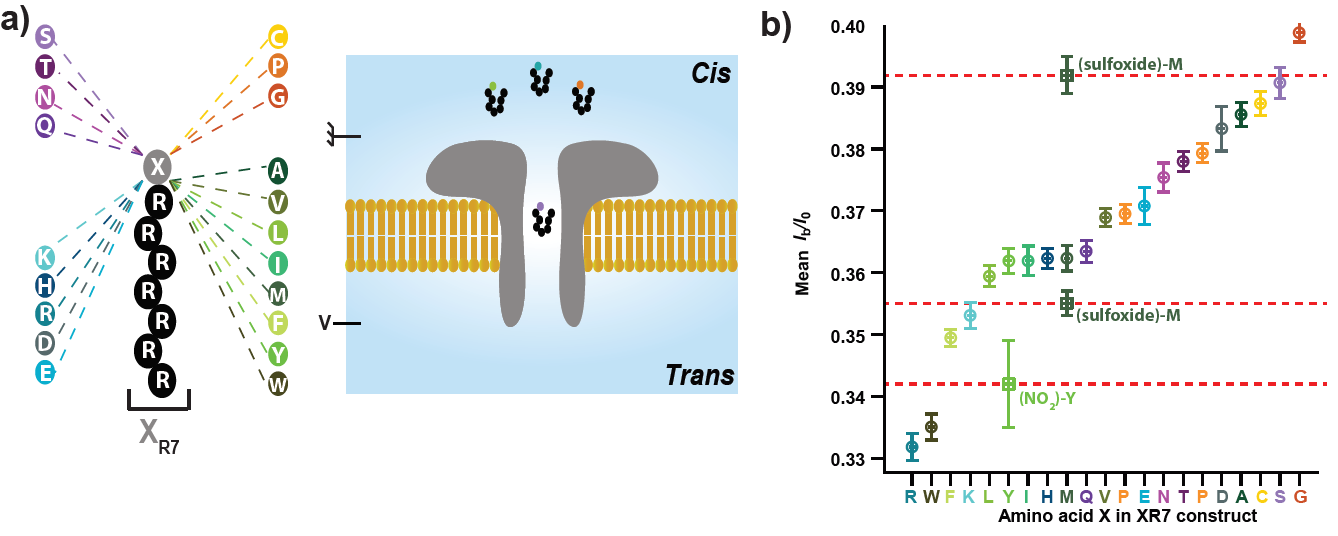
Figure 1. Identification of amino acids in the aerolysin nanopore. (a) Schematic of the peptide constructs used to investigate the current blockades produced by the twenty proteinogenic amino acids. A cationic carrier of seven arginine amino acids (R7) is chemically linked at the carboxyl terminus to the eighth amino acid, X, to form twenty XR7 peptides. (b) Experimentally determined mean relative residual current Ib/I0 value and its s.d. for all twenty XR7 peptides and for chemically modified XR7 peptides.
A few weeks later, we had the great chance to present our preliminary results at the first international symposium on single-molecule protein sequencing at Delft, Netherlands. In this conference, we met Prof. Aleksei Aksimentiev from the University of Illinois Urbana Champaign, USA, with whom we rapidly developed a collaboration directed at elucidating the physical principles underlying aerolysin detection and further improving detection resolution.
We also had the opportunity to work with the group of Pr. Jan C. Behrends, from University of Freiburg in Germany, where we spent several weeks working together to improve resolution of our measurements.
To understand the molecular mechanism enabling the exquisite sensitivity of the aerolysin nanopore, Pr. Aksimentiev and his PhD student, Kumar Sarthak, modelled the experimental measurements using molecular dynamics simulations. The simulations have shown that the high sensitivity of ionic current signals is caused by the very geometry of the aerolysin nanopore that act as a trap, holding the peptide for a prolonged time. The specific predictions of the model, i.e., the possibility of increasing the fidelity of peptide identification by increasing the residence time of the peptides inside the pore, was directly demonstrated by the high-resolution measurements realized together with Pr. Behrends and his PhD student Tobias Ensslen.
In order to extend the resolution of our approach even further, our strategy was to introduce chemical modifications on amino acids that were difficult to distinguish in their native form, namely Methionine and Tyrosine. The addition of chemical groups produced a shift of their ionic current signals, making them distinguishable from all other amino acids. These experiments have also shown that our nanopore sensing method can be used for identification of post-translational modifications.
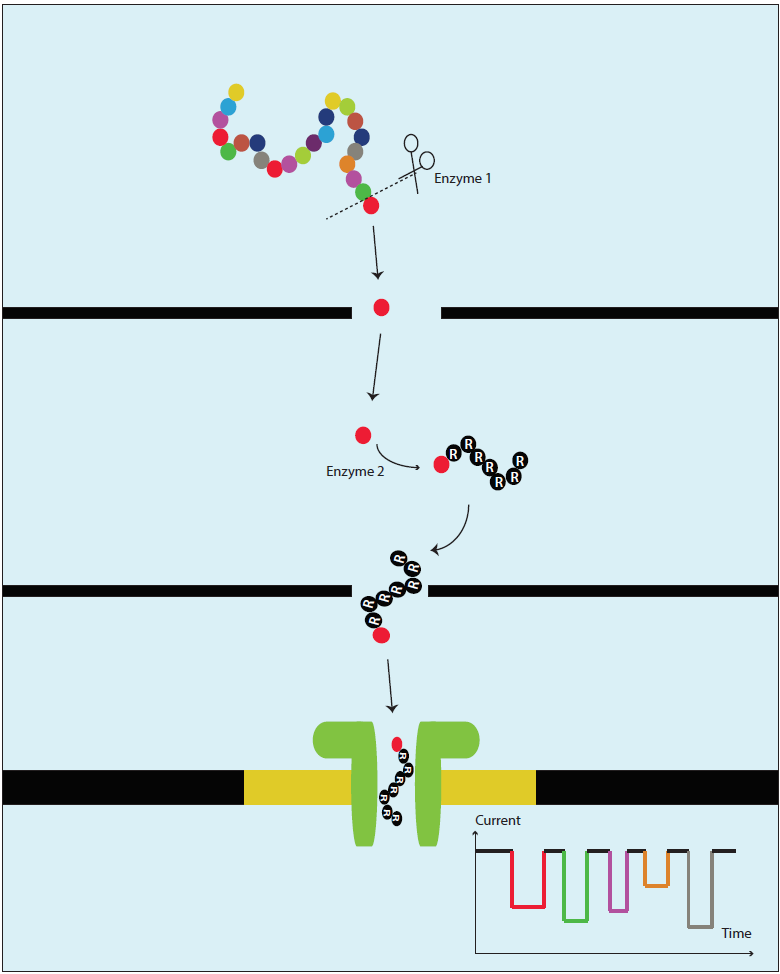
Figure 2. Schematic multistep approach envisioned for protein sequencing using a homopeptide carrier in the aerolysin nanopore
Overall, our results show a plausible route for the development of a nanopore-based protein sequencing. In this respect, our approach would require to first cleave a terminal amino acid from a target protein, ligate the cleaved amino acid to a polycationic carrier, and then analyze the complex using the aerolysin nanopore. Combining this amino acid recognition principle with instrumentation advances can open the road towards nanopore-based protein sequencing.
This paper is the result of a beautiful and fruitful collaboration between the groups of Dr. Oukhaled, Prof. Aksimentiev and Prof. Behrends.
I sincerely thank all the people who have made successful this work, especially Abdelghani Oukhaled, Aleksei Aksimentiev, Jan C. Behrends, Juan Pelta, Fabien Piguet, Kumar Sarthak, Tobias Ensslen, and Evely Sachou. I also thank the Direction Générale de l’Armement (French Defence Procurement Agency) and the Region Ile de France in the framework of DIM ResPore for giving me the opportunity to realize this PhD project.
Our paper is available at the following link: https://www.nature.com/articles/s41587-019-0345-2
The Aksimentiev group: http://bionano.physics.illinois.edu/
The Behrends group: http://www.physiologie.uni-freiburg.de/research-groups/membrane-physiology-and-technology/team/jan-c.-behrends
Follow the Topic
-
Nature Biotechnology
A monthly journal covering the science and business of biotechnology, with new concepts in technology/methodology of relevance to the biological, biomedical, agricultural and environmental sciences.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in