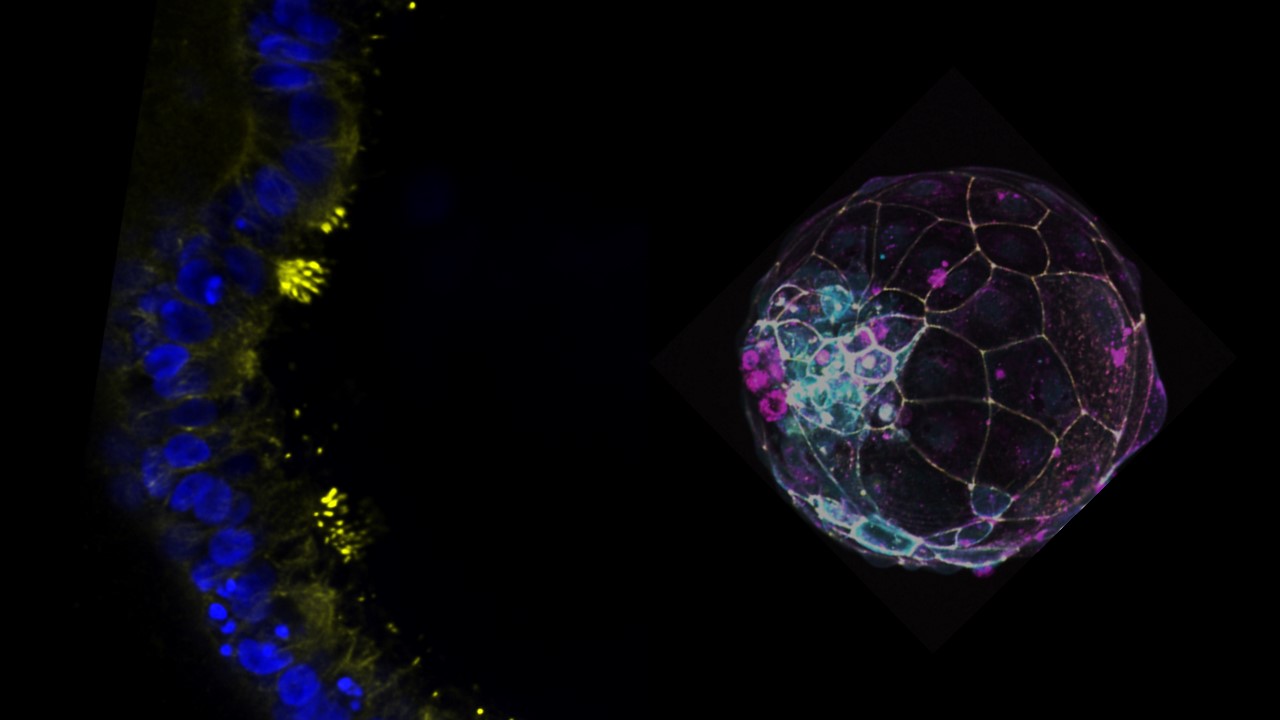
The enigmas of early human development
The early stages of human development are crucial in setting the stage for embryonic and adult life but remain difficult to study1,2. For decades, such studies have relied on donated embryos produced by in vitro fertilization (IVF). It has enabled tremendous advances in our knowledge about early development and pregnancy, and continues to provide new insights into developmental biology. These embryos, however, are scarce, difficult to manipulate, and have ethical implications. As an alternative, embryo models created from stem cells, a new field that aims to reproduce snapshots of the development, provide exciting opportunities for advancing the fundamental understanding of human development, as well as reproductive medicine.
Modeling with stem cells
In our lab, the Rivron lab at the IMBA - Institute for Molecular Biotechnology, Austrian Academy of Sciences -, our research focuses on early mammalian development and how embryos implant into the womb. Previously, we developed mouse blastoid, a model of the whole mouse blastocyst generated entirely from stem cells3. This mouse blastoid morphologically and transcriptionally resembled the blastocyst stage, generated the 3 founding lineages (trophectoderm (TE), epiblast (EPI), and primitive endoderm (PrE), and implanted into the womb. In spite of the fact that mouse blastoids have never been able to form a mouse, this model offers a scalable and experimentally flexible platform for utilizing multimodal approaches such as genetic manipulations and high-throughput imaging for the study of early development and self-organization principles. However, there are certain aspects of development that are not conserved between mice and humans. A prime example is the way mouse and human blastocysts pattern and implant into the uterus: In humans, it is the polar trophectoderm cells, the ones that are in close contact with the epiblast cells (EPI, future body), that first interact with the uterus lining, the endometrium. In contrast, in mouse embryos, it is the cells that are diametrically opposite the polar trophectoderm, the so-called mural trophectoderm cells, that establish this first contact4. Thus, understanding human development requires a human model, especially one that is scalable and allows for high-throughput and mechanistic analyses.
Human blastoids: Simple, reproducible and extremely efficient
Human blastocysts develop at a specific pace and following a specific sequence, and this allows for the formation of cells that are specific to that stage. It takes about 3.5 days for a human morula (day 4) to mature into a blastocyst (day 7)5,6. During this phase, the embryo first generates the trophectoderm (TE), which forms an epithelial cyst with a fluid-filled cavity (blastocoel) and an inner cluster known as the Inner Cell Mass (ICM). Then, in a second step, the primitive endoderm (PrE) and EPI form from the ICM, and the polar and mural regions of the TE are differentiated based on their distance from this EPI5. When modeling the blastocyst with stem cells, it is probably crucial that this natural sequence of cellular commitment, morphogenesis, and patterning events is recapitulated, and in a timely manner, because it allows for the generation of cells that closely resemble the blastocyst. Overall, closely matching developmental events allows forming a predictive model enabling scientific and biomedical discoveries. In our original publication, starting with naive state human pluripotent stem cells (hPSCs) cultured in PXGL medium, blastocyst-like structures were efficiently formed when aggregated hPSCs were exposed to a triple inhibition using lysophosphatidic acid (LPA), A83-01, and PD03259017. In our original publication, we used non-adherent hydrogel microwell plates to aggregate the cells through a technology developed by our lab8,9 and others many years ago. Now, in this Nature Protocol, we also use a wide range of commercially available plates, such as Elplasia Corning, 3DGrid SunBioscience, and AggreWell. With the Elplasia Corning and 3DGrid SunBioscience plates, experiments are performed in 96-well plates, which makes human blastoids more amenable for screening. As part of this protocol, we have also added a piece of data showing that human blastoids can not only form in stem cell media (N2B27), but also in three commercially available IVF media (G2 from Vitrolife, Continuous Single Culture-NX from Fujifilm, ORIGIO® Sequential Blast™ from CooperSurgical Fertility), making clinically relevant screens possible. Finally, we also showed that blastoids can be frozen/defrosted using a routine IVF vitrification procedure, thereby allowing for shipping.
Benchmarking the model: a challenge for the field
Forming blastoids is a challenge and, in this protocol, we explained in details our current understanding (1) of the initial parameters (e.g., initial cell state, initial aggregate properties, mix of molecules necessary and sufficient to stimulate the process) and (2) of the methods to evaluate the results (e.g., obtaining efficient morphogenesis and lineages specification, matching the pace and sequence of blastocyst development, etc…, see figure 3). As of now, the most powerful method for performing end-point assessments of the cell's reflected stage is to combine single cell RNA sequencing (scRNA sequencing) data of the model with a reference map generated from various stages of human embryos (pre-blastocyst, blastocyst, and post-implantation stages). For the purpose of benchmarking, a consortium of international scientists has developed an extensive reference map that can be used to project and evaluate the stage of the generated cells10. This enables the community to determine the degree of similarity of our model to blastocysts. Though this reference map should be improved, it uses state-of-the-art methods for merging cells harvested from human embryos at different stages. Future evaluation of blastoid cells' developmental stage(s) could be refined by analyzing transposon expression, DNA methylation, and X chromosome status. Based on our measurement and that of others, blastoids generated with this protocol contain over 97% of cells with transcriptomes similar to those of the blastocyst7,10.
Our attempt to provide a practical, efficient, and broadly applicable protocol for blastoids
Sharing this protocol with a broad group of users gives us the opportunity to also present our current understanding of 'how to form a bona fide model of the blastocyst', and we are very interested in exchanging ideas about it. Several manuscripts using this method have now been published7,11–13 and others are being prepared. This protocol is currently being used by technicians, PhD students, and postdoctoral fellows in our lab, and by our collaborators as well, and we hope that both the quality of the model and the platform's broad availability and applicabilities will make it easy to adopt by other researchers, thereby accelerating research on early human development.
References;
- Gerri, C. et al. Initiation of a conserved trophectoderm program in human, cow and mouse embryos. Nature 587, 443–447 (2020).
- Stamatiadis, P. et al. TEAD4 regulates trophectoderm differentiation upstream of CDX2 in a GATA3-independent manner in the human preimplantation embryo. Hum. Reprod. 37, 1760–1773 (2022).
- Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
- Siriwardena, D. & Boroviak, T. E. Evolutionary divergence of embryo implantation in primates. Philos. Trans. R. Soc. Lond. B Biol. Sci. 377, 20210256 (2022).
- Meistermann, D. et al. Integrated pseudotime analysis of human pre-implantation embryo single-cell transcriptomes reveals the dynamics of lineage specification. Cell Stem Cell 28, 1625–1640.e6 (2021).
- Radley, A., Corujo-Simon, E., Nichols, J., Smith, A. & Dunn, S.-J. Entropy sorting of single-cell RNA sequencing data reveals the inner cell mass in the human pre-implantation embryo. Stem Cell Reports (2022) doi:10.1016/j.stemcr.2022.09.007.
- Kagawa, H. et al. Human blastoids model blastocyst development and implantation. Nature 601, 600–605 (2022).
- Rivron, N. C. et al. Tissue deformation spatially modulates VEGF signaling and angiogenesis. Proc. Natl. Acad. Sci. U. S. A. 109, 6886–6891 (2012).
- Vrij, E. et al. Directed Assembly and Development of Material-Free Tissues with Complex Architectures. Advanced Materials vol. 28 4032–4039 Preprint at https://doi.org/10.1002/adma.201505723 (2016).
- Zhao, C. et al. Reprogrammed blastoids contain amnion-like cells but not trophectoderm. bioRxiv 2021.05.07.442980 (2021) doi:10.1101/2021.05.07.442980.
- Pham, T. X. A. et al. Modeling human extraembryonic mesoderm cells using naive pluripotent stem cells. Cell Stem Cell 29, 1346–1365.e10 (2022).
- Zhao, C. et al. Single-cell multi-omics of human preimplantation embryos shows susceptibility to glucocorticoids. Genome Res. 32, 1627–1641 (2022).
- Zijlmans, D. W. et al. Integrated multi-omics reveal polycomb repressive complex 2 restricts human trophoblast induction. Nat. Cell Biol. 24, 858–871 (2022).
Follow the Topic
-
Nature Protocols
This journal publishes secondary research articles and covers new techniques and technologies, as well as established methods, used in all fields of the biological, chemical and clinical sciences.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in