Predicting breast cancer types on and beyond molecular level in a multi-modal fashion
Published in Cancer
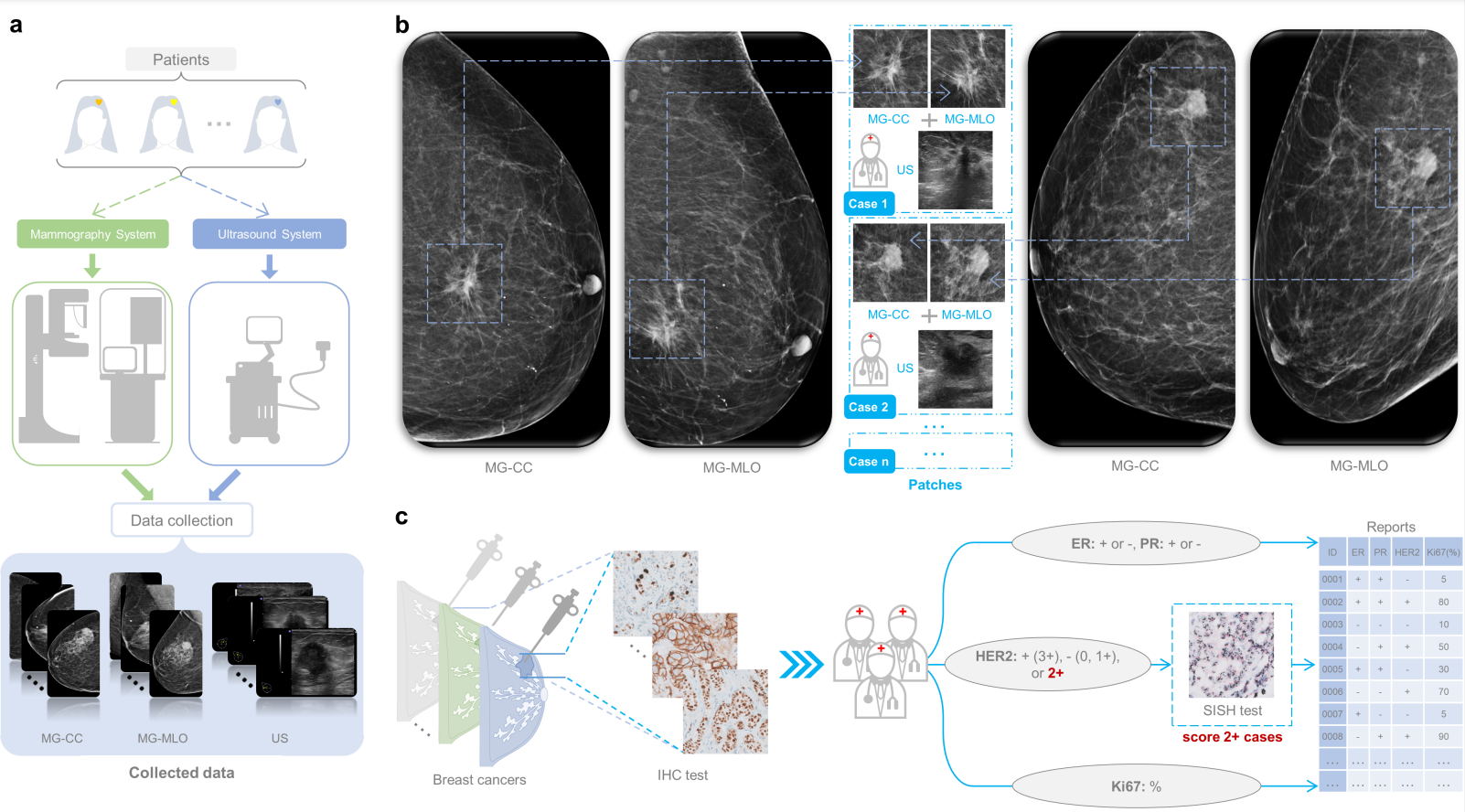
Breast cancer has a high degree of heterogeneity in terms of clinicopathological characteristics, prognosis, and response to treatment. Breast cancers can be divided into molecular subtypes, which is based upon the genetic profile of the cancers, but in practice is usually based upon the expression levels of the estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2) and Ki-67. The resulting molecular subtypes are known as Luminal A (ER+ and/or PR+, HER2-, low Ki-67), Luminal B (ER+ and/or PR+, HER2- with high Ki-67 or HER2+ with any Ki-67 status), HER2-enriched (ER-, PR-, HER2+) and Triple-negative (TN) breast cancer (ER-, PR-, HER2-). Accurately determining the molecular subtypes of breast cancer is important for the prognosis of breast cancer patients and can guide treatment selection. At present, the molecular subtypes of breast cancer are determined by immunohistochemistry (IHC) analysis of biopsy specimen, which is a surrogate for genetic testing, as the genetic analysis is quite costly. Unfortunately, the biopsy procedure limits the assessment to a small part of the tumor, which might prevent obtaining a full impression of the nature of the lesion. Differentiation within breast cancer may lead to subclones with different receptor expression, which may not be fully captured by analysis of core biopsies. Neither IHC nor in situ hybridization is available everywhere around the world, which may lead to substantial under- or overtreatment of patients. Therefore, there is a need for an effective technique to assist in the analysis of the entire breast lesions to accurately predict the molecular subtypes of breast cancer and provide decision support.
With the continuous development of artificial intelligence (AI), AI-based methods have been widely used in the field of breast imaging. In some studies, a combination of medical imaging and AI has been used to predict the molecular subtypes of breast cancer, however, most research efforts are based on traditional machine learning methods and provide limited accuracy. In addition, these previous studies only analyzed single-modality images, and did not integrate features from different imaging modalities. Moreover, obtaining breast MRI is not standard for all breast cancer patients in most countries, and is not uniformly done even within Europe.
Mammography (MG) and ultrasound (US) are routinely used during breast cancer screening, and are commonly used to identify, and characterize breast lesions and guide biopsy. Different than for breast MRI, these two modalities are virtually always and everywhere available at the time of cancer diagnosis. In this study, under the leadership of Dr. Ritse Mann, we develop a deep learning-based model for predicting the molecular subtypes of breast cancer directly from the diagnostic MG and US images.

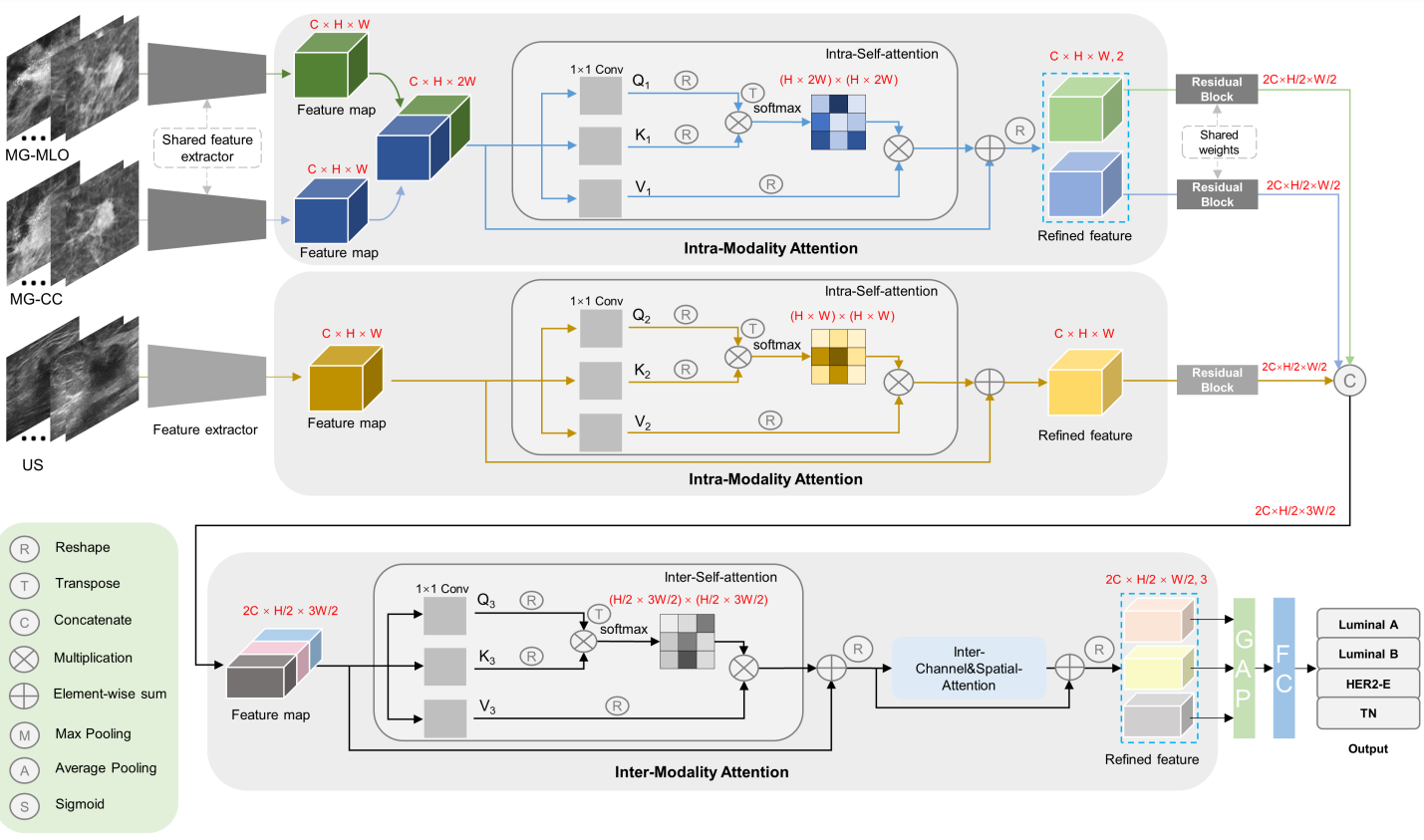
Multi-modal deep learning with intra- and inter-modality attention modules (MDL-IIA) is proposed to extract important relations between mammography and ultrasound for this task. The results of multi-modal deep learning models improve upon single-modal models. In addition, we specifically proposed the intra- and inter-modality attention modules to better integrate features of images from different modalities, further increasing the accuracy of the final result. MDL-IIA leads to the best diagnostic performance compared to other cohort models in predicting 4-category molecular subtypes.
Moreover, we conduct a reader study, comparing our multi-modal AI prediction with the prediction by experienced clinicians on the radiological images. These results of MDL-IIA significantly outperform clinicians’ predictions based on radiographic imaging. Radiologists identified some Luminal cases as TN cases, and had overall more errors in determining the 4-category molecular subtypes. This might be partly due to the fact that in clinical decision making the molecular subtypes are purely based upon pathological evaluation and radiologists are not really trained in this distinction. The higher performance of the MDL-IIA implies that more information is present. Therefore, radiologists may need more training in this area to improve their performance and could potentially also learn from the model output. Due to the inhomogeneity of cancers particularly results in which the output of the model and the pathological analysis are discrepant are of interest. Automated analysis of medical images could improve our understanding of the downstream impact of imaging features, and lead to new insights into representative and discriminative morphological features for breast cancer.
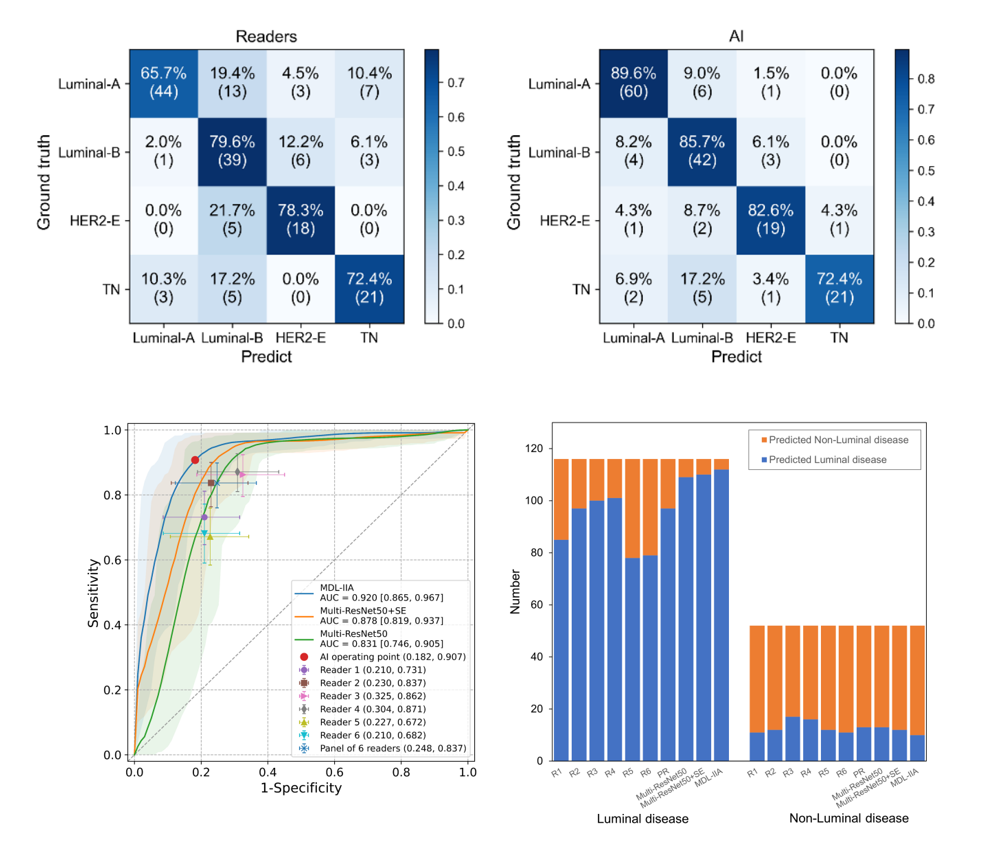
In conclusion, we have developed the MDL-IIA model, which can potentially be used to predict the molecular subtypes and discriminate Luminal disease from Non-Luminal disease of breast cancer, while being a completely non-invasive, cheap and widely available effective method. Beyond molecular-level test, based on gene-level ground truth, our method can bypass the inherent uncertainty from immunohistochemistry test. Multi-modal imaging shows better performance than single-modal imaging, and intra- and inter- modality attention modules are shown to further improve the performance of our multi-modal deep learning model. This supports the idea that combining multi-modal medical imaging may indeed provide relevant imaging biomarkers for predicting therapy response in breast cancer, thereby potentially guiding treatment selection for breast cancer patients.
“This is important as there is a large variability in histopathological assessment of breast lesions and even in simple tasks as receptor status assessment,” Ritse said. “Moreover, in many parts of the world receptor evaluation is not always performed due to costs. This algorithm enables virtually free determination of subtypes and may aid therapy selection even in places where pathologic evaluation is suboptimal.”
Follow the Topic
-
npj Breast Cancer
This journal publishes original research articles, reviews, brief communications, matters arising, meeting reports and hypothesis generating observations which could be unexplained or preliminary findings from experiments, novel ideas or the framing of new questions that need to be solved.
Related Collections
With Collections, you can get published faster and increase your visibility.
Advances in modeling
Publishing Model: Open Access
Deadline: Aug 06, 2026
Molecular Tumor Board in Breast Cancer
Publishing Model: Open Access
Deadline: Jul 22, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in