Random barcoding enables the study of plant infection by a bacterial pathogen
Published in Ecology & Evolution
The puzzle
Have you ever wondered how plants fight against bacterial pathogen invaders? Or, how do plants prevent the spread of infection across plant tissues? And how do bacteria adapt to overcome plant defenses? These are some of the questions that motivated our research, published in Nature Communications 1. In brief, where we developed a novel technology to track the population dynamics and intraspecific diversity of bacterial pathogen invaders during plant tissue infection.
Our solution
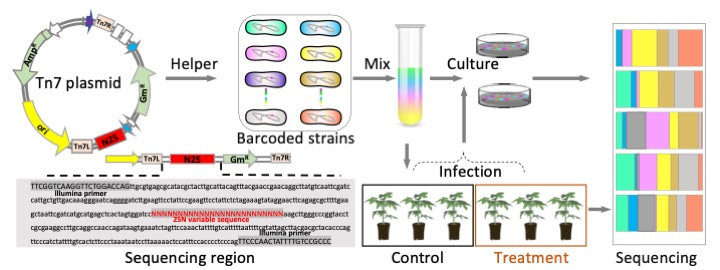
Our technology is based on the Tn7 transposon to create the bacterial invader populations carrying unique chromosomal barcodes (Figure 1a). In our technology, we insert a random barcode in a neutral location of the bacterial chromosome, which acts as a unique identifier for each bacterial cell, just like a tag. By sequencing this barcode, we can determine the abundance and diversity of the invader population at the strain-level within plant tissues (Figure 1b). Since bacteria divide asexually, the abundance of each barcode comes from initially a single cell, so we can track entire cell lineages within complex environments such as plant tissues. This approach allows us to quantify general processes such as genetic drift, or aspects of population dynamics such as bottleneck effects, at an unprecedented level of precision.
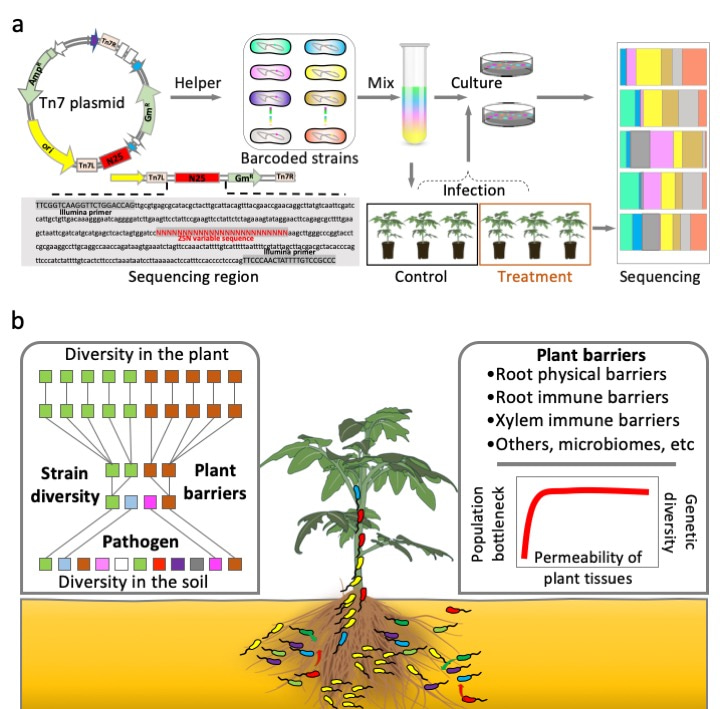
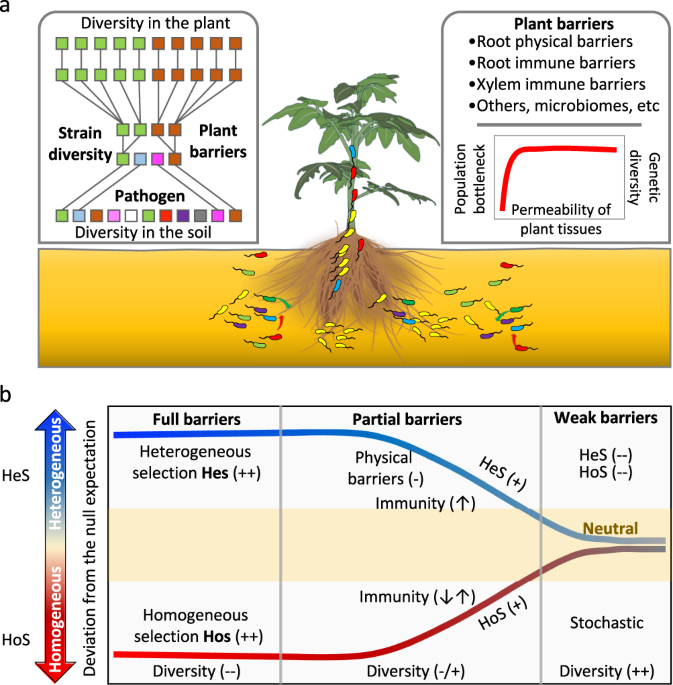
Figure 1 | The developed strategy used to investigate the effects of plant barriers on phytopathogen invader populations. a) Random barcoding-based phytopathogen infection tracking system. b) Schematic representation of the invasion processes of phytopathogens moving from the soil toward internal plant tissues.
Model system
We applied our method to investigate the infection of tomato plants by the pathogen Ralstonia solanacearum. This pathogen has impacts on diverse plant species – causing wilt disease in many crops worldwide, and particular in China (Figure 2). We infected tomato plants with a mixture of R. solanacearum populations carrying unique barcodes, and collected samples from the aerial part of the plants. We then sequenced the barcodes and analyzed the data using distinct models. We found that plant root permeability, the difficulty of pathogens to invade plant tissues due to plant physical or immunological barriers, acts as the main bottleneck to pathogen invasion affecting intraspecific diversity. The bacterial population in planta was diverse, undergoing drastic changes in size and composition during the infection of internal plant tissues. These changes were mostly driven by stochastic processes such as random drift, rather than by natural selection or adaptation.
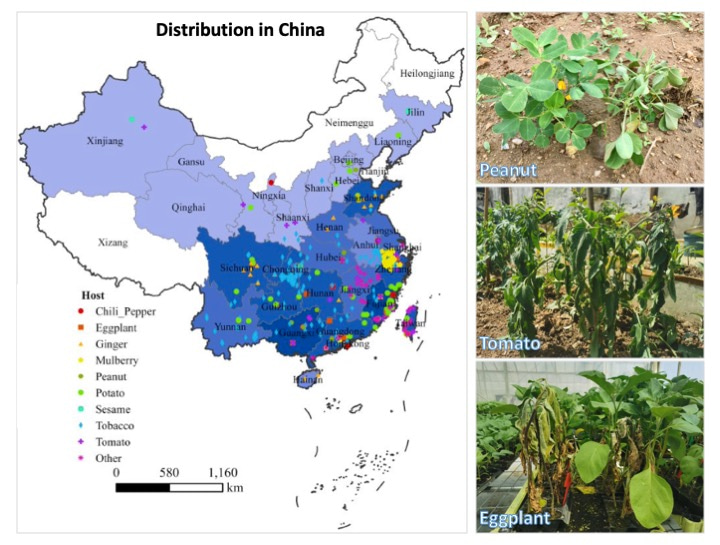
Figure 2 | Geographical distribution of bacterial wilt caused by Ralstonia solanacearum in China. Color depth indicates the number of records in each province of China 2. Typical symptoms of wilt disease on peanuts, tomato, and eggplant. Pictures Credit: Gaofei Jiang.
The end
Our study demonstrates the broad applicability of random barcoding to explore plant-pathogen interactions at an unprecedented level of detail. We hope that our work will inspire other researchers to use this method to develop other prospective studies on distinct phytopathogens, plant hosts and microbiomes 3. Such an effort will likely lead to novel insights into the ecology and evolution of phytopathogens. We expect our study to raise awareness in the public about the importance and complexity of plant diseases, and the need for developing innovative, effective, and sustainable solutions to protect our crops and ecosystems from diverse phytopathogens, most of which are expected to increase under current and future climate change scenarios 4.
Reference
- Gaofei Jiang, et al., Effects of plant tissue permeability on invasion and population bottlenecks of a phytopathogen. Nature Communications, 2024, 15, 62.
- Gaofei Jiang, et al., Bacterial wilt in China: history, current status, and future perspectives. Frontiers in Plant Science, 2017, 8, 1549.
- Zhong Wei, et al., Rhizosphere immunity: targeting the underground for sustainable plant health management. Frontiers of Agricultural Science and Engineering, 2020, 7: 317‒328
- Brajesh K. Singh, et al. Climate change impacts on plant pathogens, food security and paths forward. Nature Reviews Microbiology, 2023, 21, 640–656.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Healthy Aging
Publishing Model: Open Access
Deadline: Dec 31, 2026

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in