Targeting a Lineage-Specific PI3Kγ/ AKT Signaling Molecule in Acute Myeloid Leukemia Using a Heterobifunctional Degrader Molecule
Published in Cancer, Chemistry, and Genetics & Genomics
Background/ introduction
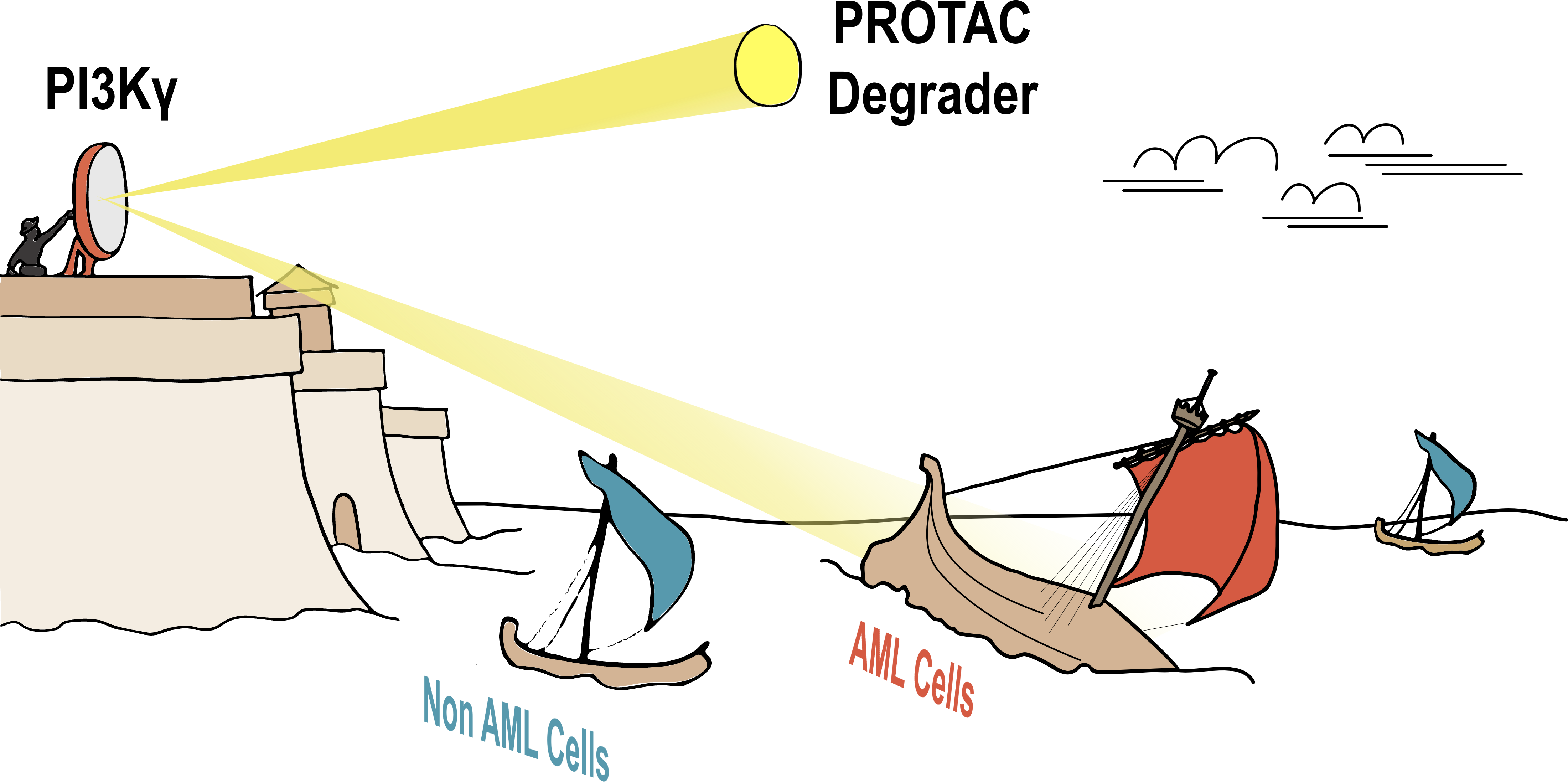
Acute myeloid leukemia is a cancer which derives from the myeloid lineage of the hematopoietic system. The 5-year survival rate for patients diagnosed before age 65 is over 50%; however, for patients over 65 years, the 5-year survival rate is significantly lower, falling below 10% (1). This disparity is partially driven by dose limiting toxicities of the standard-of-care chemotherapy which is poorly tolerated by elderly patients. To avoid off-target effects in non-malignant tissues caused by traditional chemotherapies, identifying a treatment that engages a lineage-specific target could improve tolerance to therapy and, consequently, survival outcomes in AML patients (2-5). A suitable lineage-specific target would thus need to: i) exhibit an expression pattern restricted to the myeloid compartment, ii) have pro-survival functions, and iii) be a druggable protein. In our article, we describe the identification of PI3Kγ as a lineage-specific target in AML. As a result, we developed a bifunctional degrader compound targeting PI3Kγ, which demonstrated superior potency in AML cells compared to existing small-molecule inhibitors (Figure 1).
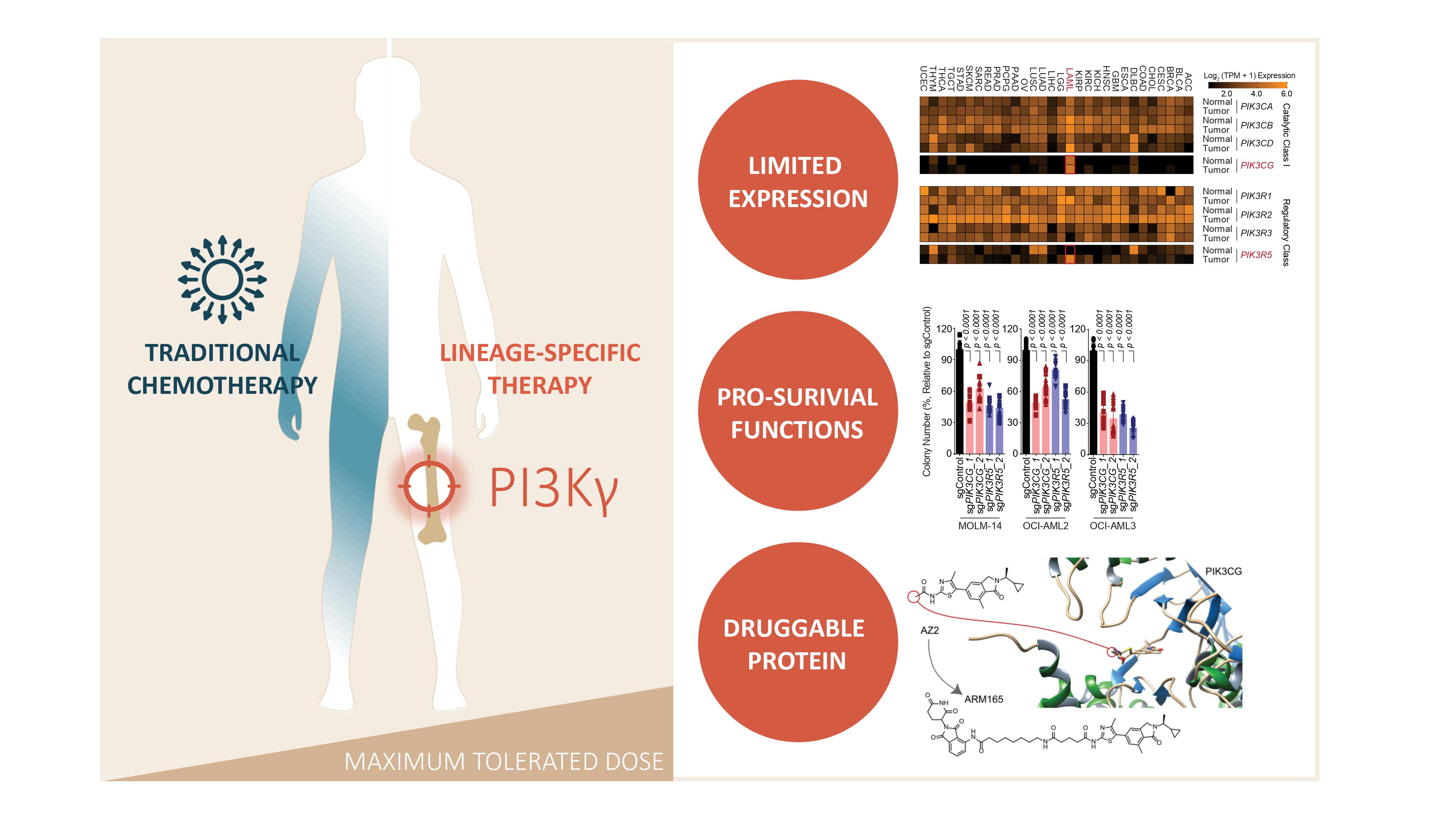
Key Findings
The myeloid-limited expression of PI3Kγ
In our study, we identified that PI3Kγ exhibits a restricted expression pattern in the myeloid compartment of the hematopoietic system, which translated to a dependency of acute myeloid leukemia (AML) cells on the expression of PIK3CG and PIK3R5, encoding the catalytic and regulatory subunit of PI3Kγ respectively (6-9). CRISPR-Cas9-mediated knockout of PIK3CG and PIK3R5 significantly dampened AML cell viability and colony forming capacity (Figure 1).
In our endeavor to investigate the potential of PI3Kγ as a target in AML, we used a publically available AML dataset to establish a positive correlation between high expression of PIK3CG and PIK3R5 and resistance to venetoclax. We therefore asked the question, could the targeting of PIK3CG and PIK3R5 increase sensitivity to venetoclax? We observed a substantial potentiation of the ability of venetoclax to reduce cell viability and colony forming capacity in combination with PIK3CG and PIK3R5 knockouts. These findings were ultimately validated in cell line-derived xenografts, where suppression of PIK3CG resulted in a significantly reduced disease burden, leading to a notable mouse survival advantage. This survival benefit was further enhanced by treatment with venetoclax.
PI3Kγ carries out essential pro-survival functions in AML
The next step was to identify the downstream cell signaling pathway regulated by the PIK3CG/R5 module in AML cells. We conducted a PIK3R5-based interactomic analysis to map PIK3CG/R5 interactors, revealing several nodes converging on AKT signaling. Using genetic tools to knockout or knockdown the expression of all the PI3K isoforms, we observed that AML cells rely strongly on PI3Kγ to activate the downstream AKT pathway. This led us to hypothesize that the ablation of PI3Kγ could reduce AML cell viability by disrupting downstream AKT activation.
To address this question, we examined the ability of existing small-molecule inhibitors to inhibit AKT signaling and reduce AML cell viability. Surprisingly, we found that existing PI3Kγ inhibitors failed to sustain long-term inhibition of both AKT signaling and AML cell growth. Consequently, an alternative therapeutic approach was required to ensure sustained disruption of PIK3CG/PIK3R5-mediated AKT signaling in AML cells.
PI3Kγ as a druggable protein: a proof-of-concept demonstrating superior cytotoxic capacity of a bifunctional degrader compound over existing small-molecule inhibitors of PI3Kγ.
In order to improve upon existing pharmacological agents targeting PI3Kγ, we designed a PIK3CG-targeting cereblon-based degrader molecule, or proteolysis targeting chimera (PROTAC), named ARM165. This degrader molecule was designed based on the structure of the existing PI3Kγ-targeting small-molecule inhibitor AZ2. We tested ARM165 for both specificity and efficacy. Unbiased proteomics and western blotting techniques revealed that ARM165 specifically degrades PIK3CG, leading to sustained ablation of AKT signaling. Furthermore, ARM165 significantly reduced the viability of AML cells compared to the parental AZ2 compound and an inactive degrader molecule. Importantly, ARM165 did not reduce the viability or colony-forming capacity of non-AML cell lines, further confirming the restricted dependency on PI3Kγ in AML cells.
To explore the clinical utility of ARM165, we tested its impact on viability and colony-forming capacity in primary patient cells. The results demonstrated its superior anti-leukemic capabilities compared to existing PI3Kγ small-molecule inhibitors. Based on our previous observation that targeting PIK3CG could potentiate the effect of the existing AML therapy venetoclax, we combined ARM165 with venetoclax and observed a similar potentiation effect in primary patient cells.
Finally, we employed two types of mouse models: one generated by injecting CBFB-Myh11-driven AML murine cells into syngeneic animals, and the other by xenotransplantation of primary AML patient cells into immunodeficient mice. In both models, ARM165 significantly reduced the disease burden in the bone marrow, spleen, and blood, while also enhancing the effects of venetoclax. These findings provide compelling proof-of-principle that targeting PI3Kγ offers a promising therapeutic strategy for AML. This work lays the groundwork for future studies aimed at enhancing the efficacy and delivery of PI3Kγ-targeting degrader molecules in vivo.
Conclusion
In summary, our research underscores the potential of targeting PI3Kγ specifically within the myeloid lineage, thereby minimizing off-target effects in non-malignant tissues. We elucidated the critical role of the PIK3CG-PIK3R5 signaling axis in activating downstream AKT signaling, highlighting its significance in AML cell growth. Moreover, our findings showcase the capacity of PI3Kγ-directed therapy to enhance the efficacy of current AML treatments, such as venetoclax. This study serves as a compelling demonstration of the potential of PROTAC degraders in disrupting PI3Kγ in AML, offering a promising avenue for enhancing the sustained targeting and efficacy of existing PI3Kγ-targeting small-molecule inhibitors. Through the development of this degrader, we have demonstrated the superior anti-leukemic efficacy achieved by targeting PI3Kγ compared to its conventional inhibition using small-molecule inhibitors.
References
- Shimony, S., Stahl, M. & Stone, R. M. Acute myeloid leukemia: 2023 update on diagnosis, risk-stratification, and management. 901 American journal of hematology 98, 502-526, doi:10.1002/ajh.26822 (2023).
- Burger, J. A. Treatment of Chronic Lymphocytic Leukemia. The New England journal of medicine 383, 460-473, doi:10.1056/NEJMra1908213 (2020).
- Jordan, V. C. & O'Malley, B. W. Selective estrogen-receptor modulators and antihormonal resistance in breast cancer. Journal of clinical oncology : official journal of the American Society of Clinical Oncology 25, 5815-5824, doi:10.1200/JCO.2007.11.3886 (2007).
- Watson, P. A., Arora, V. K. & Sawyers, C. L. Emerging mechanisms of resistance to androgen receptor inhibitors in prostate cancer. Nature reviews. Cancer 15, 701-711, doi:10.1038/nrc4016 (2015).
- Yu, A. L. et al. Anti-GD2 antibody with GM-CSF, interleukin-2, and isotretinoin for neuroblastoma. The New England journal of medicine 363, 1324-1334, doi:10.1056/NEJMoa0911123 (2010).
- De Henau, O. et al. Overcoming resistance to checkpoint blockade therapy by targeting PI3Kgamma in myeloid cells. Nature 539, 443-447, doi:10.1038/nature20554 (2016).
- Foubert, P., Kaneda, M. M. & Varner, J. A. PI3Kgamma Activates Integrin alpha(4) and Promotes Immune Suppressive Myeloid Cell Polarization during Tumor Progression. Cancer immunology research 5, 957-968, doi:10.1158/2326-6066.CIR-17-0143 (2017).
- Schmid, M. C. et al. Receptor tyrosine kinases and TLR/IL1Rs unexpectedly activate myeloid cell PI3kgamma, a single convergent point promoting tumor inflammation and progression. Cancer cell 19, 715-727, doi:10.1016/j.ccr. (2011).
- Kaneda, M. M. et al. PI3Kgamma is a molecular switch that controls immune suppression. Nature 539, 437-442, doi:10.1038/nature19834 (2016).
Follow the Topic
-
Nature Cancer
This journal aims to provide a unique forum through which the cancer community will learn about the latest, most significant cancer-related advances across the life, physical, applied and social sciences.
Your space to connect: The Myeloid cell function and dysfunction Hub
A new Communities’ space to connect, collaborate, and explore research on Clinical Medicine and Cell Biology!
Continue reading announcementRelated Collections
With Collections, you can get published faster and increase your visibility.
Cancer Neuroscience: from mechanisms to therapy
Publishing Model: Hybrid
Deadline: Jan 30, 2027



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in