The genetic architecture and biology of youth-onset type 2 diabetes -- the
Published in Protocols & Methods, Genetics & Genomics, and General & Internal Medicine
Back in 2009, soon after the first genome wide association studies (GWAS) were published and as exome sequencing was becoming cost-effective, complex trait geneticists began to turn their attention to understanding the role of rare coding variants in common disease. Some researchers were more bullish on the value of rare variants than were others, but nearly everybody was interested in them. It soon became apparent, however, that "high impact" rare variant risk factors for common disease were very hard to find, and, as GWAS continued to produce more and more common variant associations for more and more traits, the field shifted gradually but steadily to a model in which common variant risk factors explained the bulk of common disease heritability. For type 2 diabetes (T2D), we were at last able to identify *some* rare variant associations ten years after we started looking for them and with 100x more samples than we at first thought we would need, but even then the message was that rare variants (while potentially useful for providing "genetic support" for therapeutic targets) seemed to play a bit part in common disease.
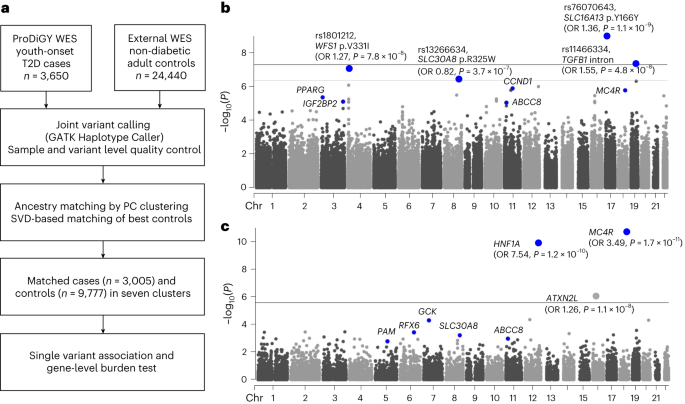
It was within this context that we began our analysis of ~3,600 exomes for youth-onset T2D as part of the ProDiGY study. Youth-onset T2D is an important disease in its own right, as (by definition) it affects children, particularly those from minority groups, and is rapidly rising in prevalence. It is also of particular interest to a geneticist, because it's related to two more widely studied diseases: adult-onset T2D and Maturity Onset Diabetes of the Young (MODY). Adult-onset T2D is the most common form of diabetes and typically presents after the age of 40 due to both environmental and genetic risk factors; it's been extensively studied by GWAS, and it's very heterogeneous both biologically and genetically, being influenced by thousands of common genetic variants spread across numerous genes and biological pathways. In fact, it's so heterogeneous that we actually think of it now as a collection of disease subtypes, each with a distinct clinical presentation, prognosis, and genetic cause.
MODY lies on the other "extreme" as a monogenic form of diabetes that presents much earlier in life and is caused by single (very rare) mutations in one of ~10-15 genes involved in beta-cell development and insulin secretion. MODY is distinct from T2D but symptoms do overlap and are sometimes misdiagnosed, and for most MODY genes we have also found variants that are associated with T2D. So these two forms of diabetes clearly overlap to some extent, although exactly how widely is unclear. Notably, the ProDiGY study excluded cases suspected as having MODY, and at any rate the youth-onset T2D phenotype is quite distinct from MODY -- cases are typically obese rather than lean (as is typical for MODY).
The obvious genetic hypothesis regarding youth-onset T2D is that it is somehow "in between" MODY and adult-onset T2D. If true, this would suggest that perhaps adult-onset T2D subtypes extend beyond those we've identified thus far to forms that present early on in life, and that perhaps these forms eventually blur into monogenic diseases like MODY. This would be a model consistent with the hopes of personalized medicine to move beyond broad descriptions of disease based on symptoms to individual descriptions of disease based on the specific genetic insults and disrupted biological processes. It would also imply that perhaps we might not only use our relatively good understanding of the genes and pathways involved in MODY to help anchor our understanding of adult-onset T2D (which we basically already do), but also that we might be able to apply our advanced statistical methods and large genomic datasets for adult-onset T2D to better diagnose and treat rare forms of diabetes.
Based on the genetic data we analyzed, this is exactly the case. The first striking finding was several strong rare variant association signals, with significance levels exceeding those that we observed in 10x as many cases for adult-onset T2D. These signals heavily overlap known genes for adult-onset T2D and MODY and cluster very clearly within pathways previously implicated in both adult-onset T2D and MODY. This is evidence of a strong genetic overlap between youth-onset T2D and both adult-onset T2D and MODY.
Furthermore, as a whole, the youth-onset cases are definitively more genetically extreme than are adult-onset cases. They are more likely to carry ultra-rare MODY variants and they have an excess of common variant risk factors *and* an excess of rare variant risk factors. In absolute terms, most of their risk is due to common variants -- as is the case for adult-onset T2D -- but in relative terms, the distribution of their risk factors skews toward rare variants more so than is the case for adult-onset T2D. This is strongly consistent with the "in between" genetic model for youth-onset T2D.
The above picture about rare and common variant risk factors holds *in average* across all samples. But, this doesn't tell us much about individual cases. It could be the case that *all* samples look roughly the same genetically with a comparable mix of common and rare variant risk factors, or it could be the case that some samples have risk due almost entirely to rare variants and some samples have risk due almost entirely to common variants, with these heterogeneities averaging out across the population. Both models are consistent with the population-level data, but the latter model is more promising for personalized medicine, as it suggests that individual cases might be "simpler" genetically than is the population.
When we looked at individual cases, we found evidence for the second model. Some cases have genetic risk almost entirely due to rare variants, and on the other extreme some cases have genetic risk almost entirely due to common variants. In between are cases with a mixture of risk factors, with no clear dividing line between "rare" and "common" variant cases. But -- and most intriguingly -- phenotypic presentation correlates with the frequency of genetic risk factor: common variant risk correlates with an adult-onset T2D-like phenotype, ultra-rare variant risk correlates with a MODY-like phenotype, and rare variant risk correlates with an intermediate phenotype (earlier age of onset than adult-onset T2D but not as lean as MODY). This is unlikely to be useful in personalized medicine, because the biological pathway impacted by a variant is likely more important clinically than its frequency, but a dependence between variant frequency and variant effect is consistent with models in which rare variants are more likely to impact "core" genes than are common variants.
So what does this all mean? It emphasizes that, as has been recognized for years, we should keep studying patients with extreme phenotypes, particularly those in the sweet spot with severe enough disease to expect to find strong signals but common enough disease to be amenable to large-scale statistical studies. It suggests that perhaps it is worthwhile seeing how blurred the line really is between rare and common forms of disease. And, it helps us understand youth-onset T2D as a condition closer to adult-onset T2D than to a monogenic condition, but one in which nearly every type of genetic risk factor for adult-onset T2D is observed at elevated frequencies.
From another (perhaps more narrow) perspective, it represents a satisfying next step in the story of rare variants and T2D. While the picture that emerged from this study is perhaps unsurprising, the clarity with which it was painted was something that we did not expect based on our years of work -- and trials -- in this area.
Follow the Topic
-
Nature Metabolism
This journal publishes work from across all fields of metabolism research that significantly advances our understanding of metabolic and homeostatic processes in a cellular or broader physiological context, from fundamental cell biology to basic biomedical and translational research.
Related Collections
With Collections, you can get published faster and increase your visibility.
The expanding therapeutic landscape of GLP 1 receptor agonists
Publishing Model: Hybrid
Deadline: Jan 23, 2027
Microbiome and energy metabolism
Publishing Model: Hybrid
Deadline: Dec 06, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in