Ecosystem-wide antibiotic tolerance in the murine gut
Published in Microbiology

Maintaining an arsenal of effective antibiotics is an uphill battle. During the course of our lives, we will take many doses of antibiotics to eradicate one pathogen or another. The commensal bacteria that die in the wake of antibiotic treatment are collateral damage in the struggle to resolve the infection. This sets up an ongoing arms race between the bacteria in the gut and these frequently-applied antibiotics. Our gut bacteria learn to circumvent antibiotics by transferring resistance genes back and forth and employing a variety of tolerance-promoting mechanisms.
Many prior studies have characterized a single bacterial strain’s responses to a given antibiotic. Prior work in mice and humans has suggested that microbiome-wide responses to the same type and dose of an antibiotic can vary, with some individuals showing a minimal response (i.e. non-responders). However, these non-responder phenotypes have not been well characterized and remain poorly understood. A classical view of the drivers of these phenotypes might implicate the transfer of and enrichment for antimicrobial resistance genes among commensal strains. . Our initial foray into this topic involved the accidental discovery that these phenotypes appear to arise spontaneously, over very short timescales, and with fairly controlled/homogeneous initial conditions, which seemed to rule out the classical view. It all started with what looked like a failed experiment.
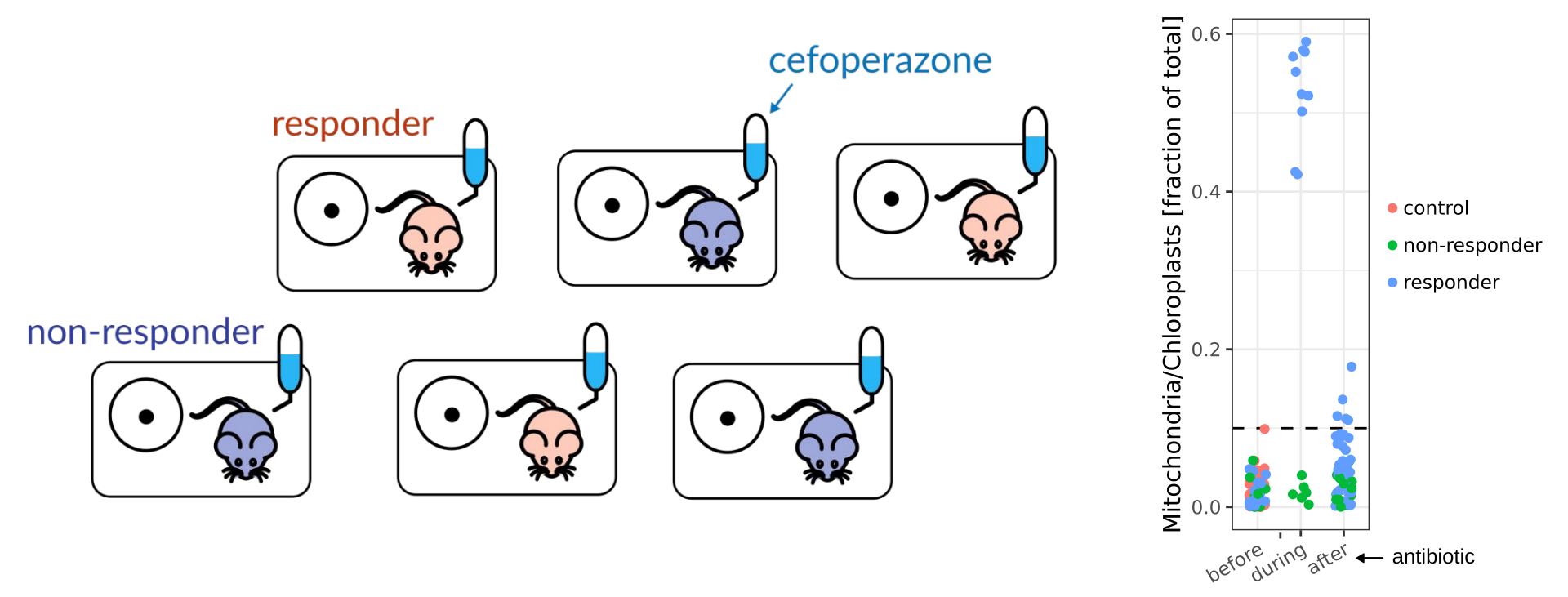
During his Postdoc at MIT Sean Gibbons, along with MIT graduate student Sean Kearney, studied how dietary components containing complex phytochemicals, such as seaweed, might modulate commensal susceptibility to a broad-spectrum antibiotic commonly used in experimental models of Clostridioides difficile infection in mice (cefoperazone). Their experimental setup was a bit different from what is usually done: their mice were singly-housed (after an initial period of co-housing). Usually, researchers house several mice together in a single cage because they are happier when co-housed. You need special experimental justification for single-housing. Because mice are coprophagic (poop-eating), co-housing them homogenizes their microbiota. The Seans were interested in removing this homogenization effect to better quantify inter-individual variability. To their surprise, they observed a binary response in the mouse gut microbiomes to antibiotic treatment. For ⅔ of the mice, the bacterial biomass in the gut totally collapsed (i.e. ‘responders). In the remaining ⅓ of mice the gut bacterial community composition looked almost identical to untreated controls (i.e. ‘non-responders’). The difference in biomass was massive. In responder mice, more than half of the total reads were annotated as mitochondria and chloroplasts, most likely derived from the mouse chow and host cells, whereas this abundance of organelle reads was less than 5% in non-responder and control mice. Quantitative PCR of the 16S rRNA gene confirmed that this enrichment for organelle reads was due to an almost complete collapse in bacterial biomass in responder mice. This non-responder effect was observed at a similar frequency across seaweed dietary treatments.
This result was very unexpected and somewhat suspicious, so in order to rule out experimental error, the Seans repeated the experiment with a new cohort of mice, where they varied the duration of cefoperazone treatment to see if that might influence the phenotype. The results were replicated, and they again observed the non-responder mice occurring at a similar frequency, independent of antibiotic duration. The mice shared the same mothers, were co-housed prior to beginning the experiments, and had had no prior history of antibiotic exposure. It was unlikely that resistance genes would have spread so quickly and unevenly. So what was going on? By this time, Sean G. was moving to start an Assistant Professor position at the Institute for Systems Biology (ISB), so he brought the samples and data with him to continue studying this mysterious phenomenon.
This is where Anna and I come into the story. Looking at the data in Seattle, Anna and I first thought that subtle differences in the bacterial composition between mice at the initial time point, prior to antibiotics, might explain the phenotype. We looked at 16S amplicon sequencing data from hundreds of mouse fecal samples, and even sequenced many more archived/frozen fecal pellets to look for pre-existing differences. Surprisingly, we found none. Before treatment, the composition of the gut microbiota between responders and non-responders was absolutely identical and had no predictive value in distinguishing responders from non-responders. So, we decided we needed to look beyond community composition and measure what genes were being expressed in responder and non-responder communities.
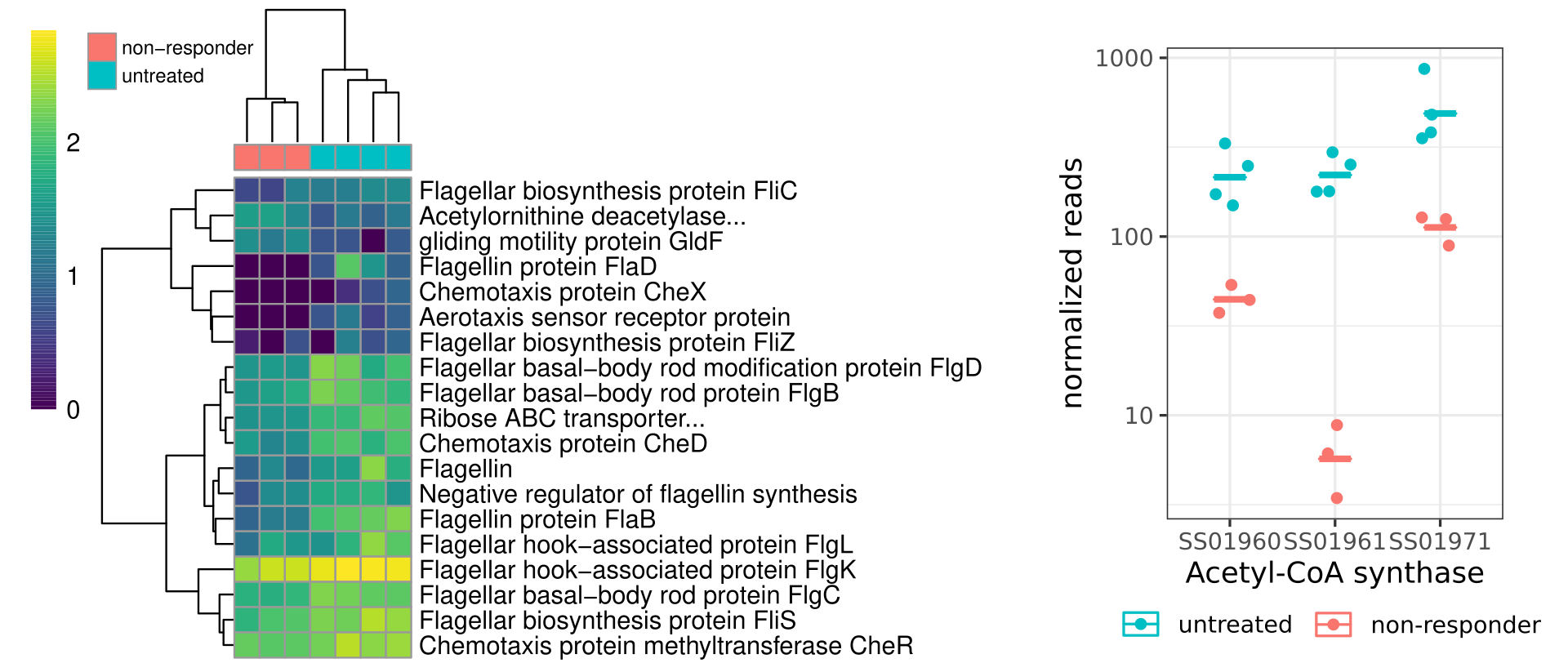

We performed RNA-seq on a subset of samples that we had archived and frozen that had already been classified as responders or non-responders before and during antibiotic treatment, along with some samples from untreated control mice. Because the responder mice had very little bacterial biomass, no RNA could be isolated from those samples that were taken during antibiotic treatment. Therefore, we first looked at whether untreated, responder, and non-responder mice showed similar transcriptional profiles prior to antibiotic treatment, and we then compared non-responder mice to untreated mice during the antibiotic treatment window. Before treatment, untreated controls, responder, and non-responder mice presented nearly identical RNA expression profiles, with fewer than 0.5% of genes showing differential expression. However, during antibiotic treatment more than ¼ of all detected genes differed significantly in expression between non-responders and controls. Here, we observed a global down-regulation of chemotaxis/motility genes and genes involved in biomass production and replication. In particular, three subunits of Acetyl-CoA synthase were down-regulated by an order of magnitude, and perfectly separated non-responders from controls. All this saved energy seemed to be redirected into the upregulation of efflux transporters and general defense mechanisms. Thus, community-wide tolerance to cefoperazone appears to be driven, in part, by the downregulation of growth and the upregulation of efflux transporters.
In summary, while non-responders showed very similar bacterial compositions to untreated mice, their transcriptional response was completely different and tailored towards surviving an antibiotic assault. This response seems to involve the majority of dominant bacteria in the microbiome and suggests that the microbiome somehow works in unison to protect itself against ecological damage and biomass collapse in the presence of antibiotic treatments. An intact and diverse commensal microbiota is an important buffer against opportunistic pathogens. We do not yet understand how microbiomes tip into responder and non-responder phenotypes, but we are excited to explore this further as a route to protecting commensal microbes from frequent antibiotic assaults.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
DNA repair and human disease
Publishing Model: Hybrid
Deadline: Oct 31, 2026
Cell death and inflammatory signalling
Publishing Model: Hybrid
Deadline: Oct 28, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in