Molecular porosity and permeability of a viral capsid
Published in Bioengineering & Biotechnology

The world has become increasingly aware of viruses during the last year, while for us virus particles have been the focus of our investigations for many years. A few thousand viruses have been studied, many of which assemble to form a protein coat or capsid and many of these have been structurally characterized due to advances in x-ray crystallography and cryo-electron microscopy. Virus capsids are porous to some molecules but the static structural models we have do not always tell us about the effective pore size and what influences which molecules get in and which do not; we were interested in directly measuring this. Some viruses undergo morphological transformations such as reversible swelling and irreversible expansion, suggesting that viruses exhibit significant elasticity.1 There have been studies indicating that dynamic fluctuations can result in transient large-scale deformations of the virus capsids and influence their shape, resembling a “breathing” mechanism. Do virus particles really “breath,” and if so, are the fluctuations so drastic that molecules diffusing across exceed the pore sizes estimated from structural models?
Not long after beginning this project, I realized that porosity goes beyond our inquiry into virus capsid dynamics. Porous barriers and diffusion across them are recurring themes found throughout nature and synthetically derived systems. Pores provide controlled permeability of molecules based on their physical properties such as size, charge, and chemical makeup. For example, carboxysomes are proteinaceous compartments found within bacterial cells and architecturally resemble icosahedral virus capsids.2 The porous shell controls diffusion across and is thought to discriminate between O2 and CO2 while simultaneously protecting densely packed enzymes that catalyze metabolic reactions. The carboxysome has inspired many designs in our lab; we postulated that the carboxysome pores would be analogous to those of virus particles, where they would resemble molecular sieves.
The Douglas Lab has developed a remarkable system using virus-like particles (VLPs) derived from bacteriophage P22 that can encapsulate a wide range of biomacromolecules as cargo, including enzymes.3The design of our experiment was simple - we used this system to encapsulate an enzyme, alcohol dehydrogenase-D (AdhD), in the P22 VLPs and probed the differences in activity using substrates of varying size or charge (Figure 1). We were able to explore pore dynamics and predict the effective pore sizes for two different P22 morphologies using this approach.
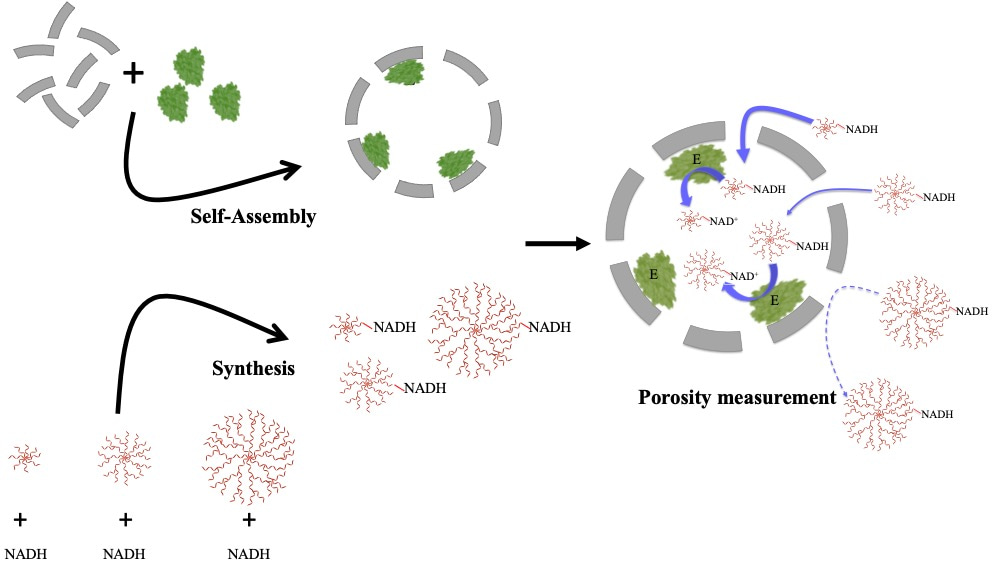
Figure 1: Scheme of our experiments: P22 particles self-assembled with coat protein (CP) and scaffold protein (SP) -enzyme fusion (AdhD-SP). NADH was synthesized to yield NADH-dendrimer conjugates. The porosity of P22 particles was probed by using the various NADH-dendrimer molecules as bait for the AdhD enzyme. If the “bait” accessed the interior of the particle, we saw the redox reaction (NADH-->NAD+).
One of the most surprising findings was that negative substrates traversed across the negatively charged P22 VLPs more readily than neutral substrates, sending us down an investigative path into the electrostatic effects of both the virus particles and the synthesized substrates. Since the zeta potential of P22 VLPs is about -30 mV, our initial hypothesis was that electrostatic repulsion would affect the negative substrates, and their translocation across the pores would be inhibited. Since our experimental results suggested otherwise, we generated an electrostatic map of the P22 asymmetric unit that revealed a complex distribution of charges around the pore, with negative charges lining the exterior regions and positive charges lining the interior (Figure 2a), resulting in an electrostatic gradient from outside to inside. The electric field lines provided the likely path that charged substrates might take when entering the electrostatic field, leading us to hypothesize that negative substrates encounter the gradient and are effectively driven inside. To explore this phenomenon, we designed two different mutants of P22 in which the charge distribution was altered by several point mutations. The results showed that removing the positive charges lining the interior pore (and disrupting the gradient) leads to a significant decrease in the ability of negative substrates to access the interior of the capsid. In this mutant, the negative and neutral substrates access the interior to the same extent.
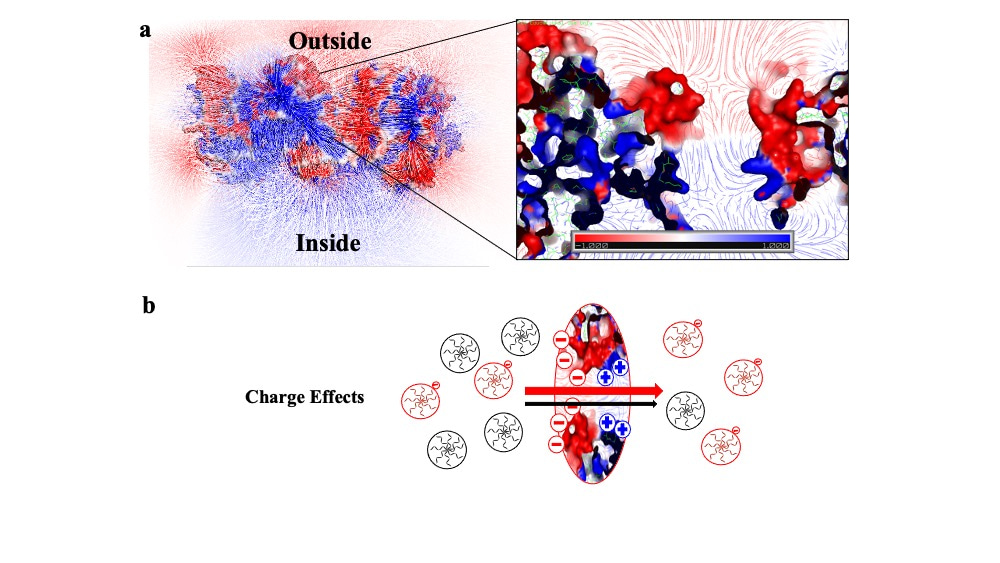
Figure 2 shows electrostatic maps of P22 pore. a) Electrostatic surface potential of the wild type P22 asymmetric unit with electric field lines with negative and positive surface potential in red and blue, respectively. Units are kT/e. b) Cartoon of dendrimers diffusing across the P22 pore where black and red represent neutral and negative molecules respectively.
Going back to our original question- do virus particles undergo dynamic fluctuations? My short answer is yes, however, our data suggest that there is a rigid size/charge cutoff for molecules diffusing across P22. Viruses come in different shapes and sizes, and therefore, it is very likely they exhibit drastically different dynamics. The dynamics can be explored with the methods provided in this elegant design. For those interested in nanotechnology, this work demonstrated the versatility of P22 VLP-based nanoreactors as permeable containers whose pore sizes can be adjusted through several morphologies, mutants which alter the pore electrostatics, and further tuned by changing the charge of the diffusants.
1 Bruinsma, R. F., Wuite, G. J. L. & Roos, W. H. Physics of viral dynamics. Nat Rev Phys, 1-16, doi:10.1038/s42254-020-00267-1 (2021).
2 Tanaka, S. et al. Atomic-level models of the bacterial carboxysome shell. science 319, 1083-1086 (2008).
3 O'Neil, A., Reichhardt, C., Johnson, B., Prevelige, P. E. & Douglas, T. Genetically programmed in vivo packaging of protein cargo and its controlled release from bacteriophage P22. Angew Chem Int Ed Engl50, 7425-7428, doi:10.1002/anie.201102036 (2011).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in