Teaming up two ESKAPE: cross-feeding and cross-protection between Klebsiella pneumoniae and Acinetobacter baumannii
Published in Microbiology
When we think about having an infection most of us imagine a single microbe as the causative agent, and while this is true for some instances of disease, in reality it is often more complicated. The world we live in is not sterile, nor are we, polymicrobial interactions are occurring all around us, all the time. It therefore may not be surprising that there is mounting evidence that polymicrobial infections are much more common than once thought. Numerous studies have found that polymicrobial infections can lead to increased virulence and antimicrobial resistance of the infecting partners 1,2. We have found this to also be true of interactions between the deadly pathogens Acinetobacter baumannii (Ab) and Klebsiella pneumoniae (Kp).
Our study began with whole genome sequencing of a clinical isolate that we thought was Ab, however, upon analysing the data it was discovered that the reads were mapping to Kp. Subsequent digging revealed that the original culture was in-fact a mixed culture of Ab and Kp, which as it turns out isn’t all that uncommon. Co-infections of Ab and Kp can occur at rates of up to 40% of infections involving either species, and generally lead to worse outcomes for patients including increased mortality rates and increased resistance to antibiotics 3–5. The strain pair we investigated were co-isolated from a single lung infection.
Ab and Kp are formidable pathogens in their own right, both identified as part of the clinically relevant ESKAPE pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa and Enterobacter spp.), a group of microbes recognised for their ability to ‘escape’ antimicrobial treatment6. There is a wealth of information on Ab and Kp virulence and resistance factors in mono-infections, but previous studies have rarely focused on their interaction dynamics in co-culture. We wanted to know how these microbes might be interacting in co-infections at a molecular level, and what implications this may have for patient outcomes and treatment strategies.
We employed a number of techniques to investigate their co-interactions including sequencing and assembling their genomes, examining changes in gene expression in co-culture, determining the ratio of Ab:Kp cells in co-culture over time, and cross-feeding and cross-protection studies. Some of the highlights were that Kp could cross-feed Ab with its waste metabolites ethanol and lactate, which became a food source for Ab. Kp has 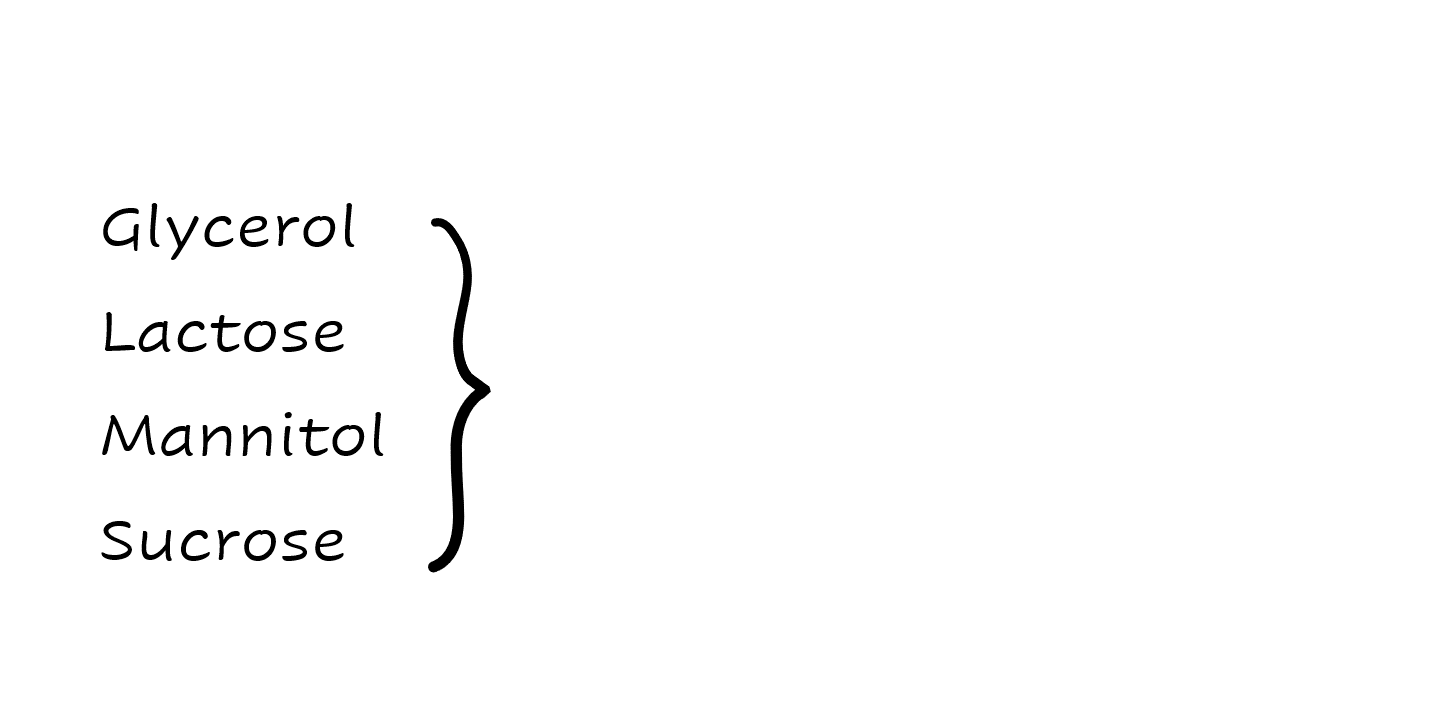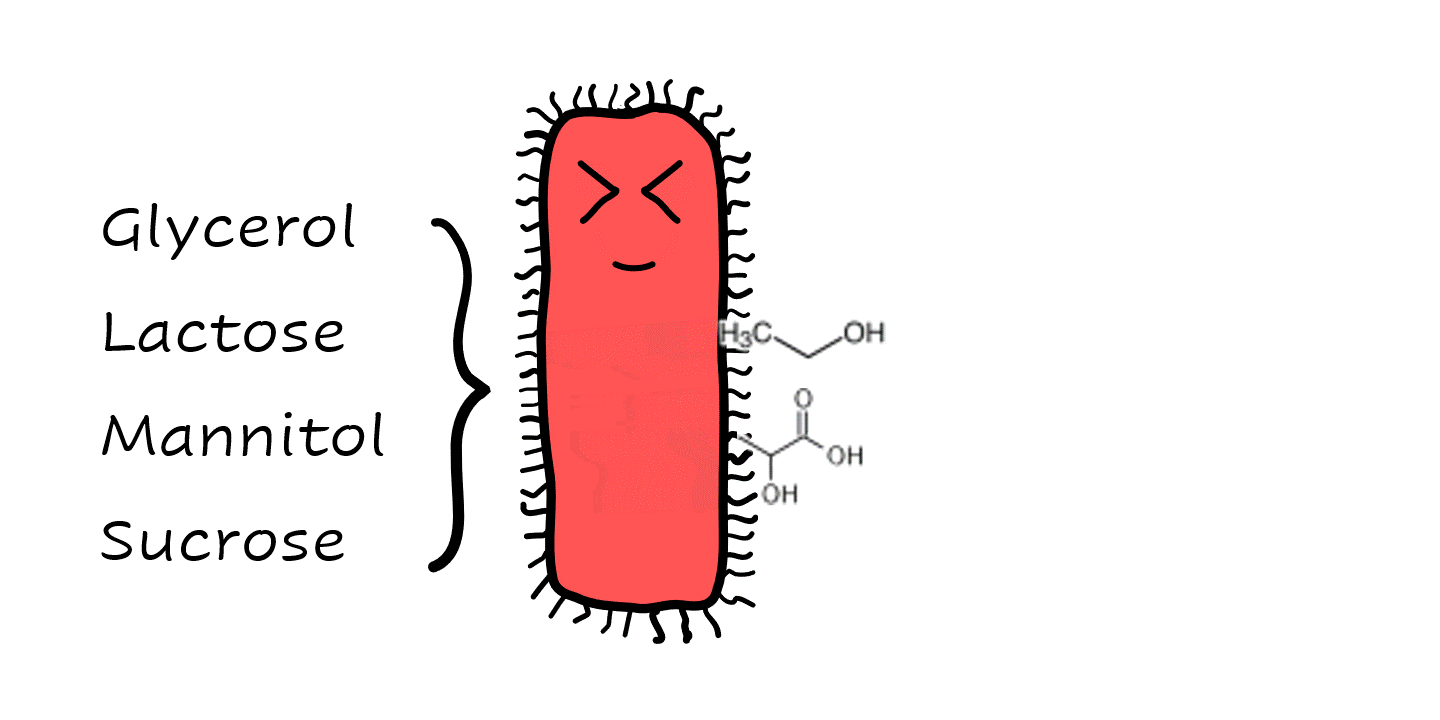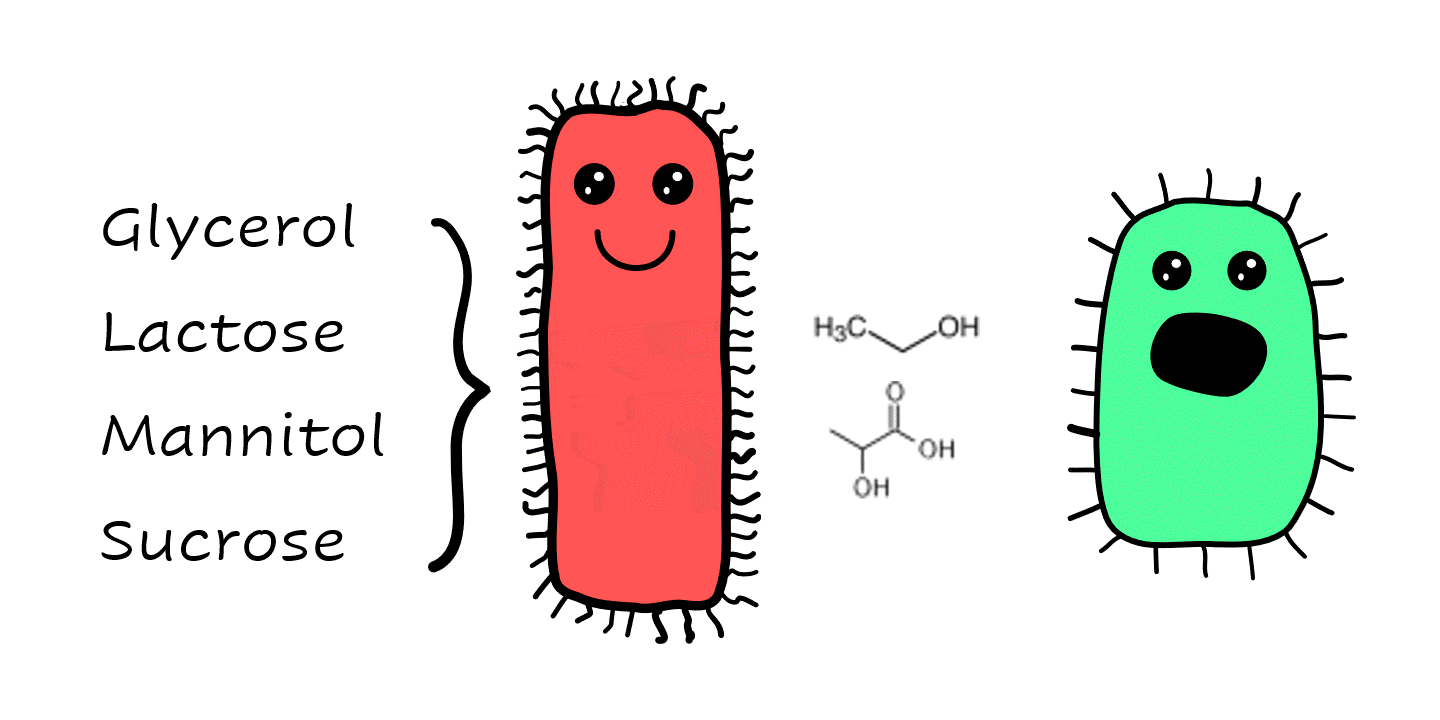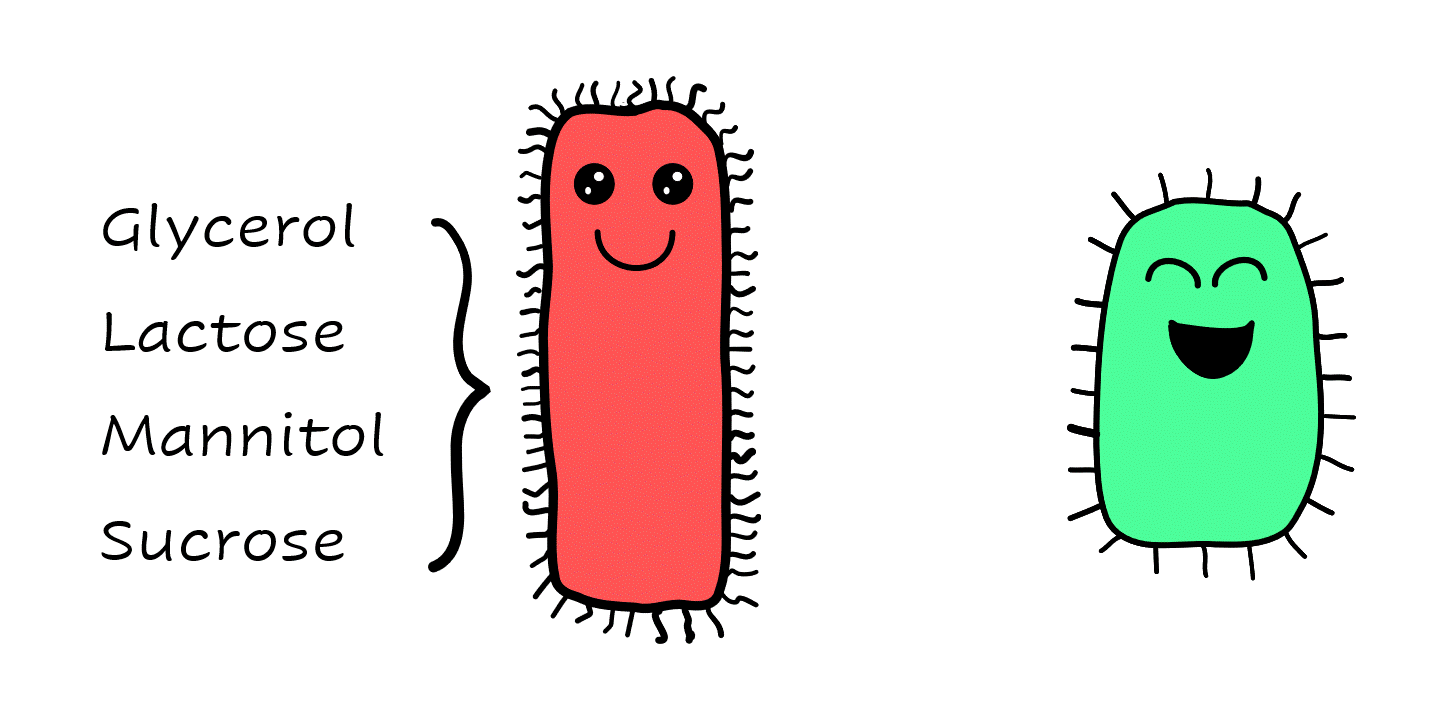 a much broader range of substrate utilisation than Ab, so it was able to grow on carbon sources that Ab cannot, such as glycerol, lactose, mannitol, and sucrose. This is important as cross-feeding allows the strain pair to co-exist in environments where they may not be able to otherwise.
a much broader range of substrate utilisation than Ab, so it was able to grow on carbon sources that Ab cannot, such as glycerol, lactose, mannitol, and sucrose. This is important as cross-feeding allows the strain pair to co-exist in environments where they may not be able to otherwise.
So, it looks like Ab may enjoy some benefits from co-existence, but what about Kp? In this particular strain pair, Ab had a more extensive antimicrobial resistance profile than Kp, so we were interested to see if Ab could provide cross-protection from antibiotics to Kp. Ab had a much higher (64 times higher!) minimal inhibitory concentration (MIC) against cefotaxime than Kp. Cefotaxime is a potent 3rd generation cephalosporin, susceptible to cephalosporinase degradation. When grown in co-culture, Kp could survive cefotaxime exposure at a concentration that was 32 times its mono-culture MIC. We discovered this was due to the secretion of cephalosporinases by Ab, as combined treatment with sulbactam, a β-lactamase inhibitor, significantly decreased cross-protection. So, it would seem that Kp can also benefit from interaction with Ab by receiving protection from at least one antimicrobial.
discovered this was due to the secretion of cephalosporinases by Ab, as combined treatment with sulbactam, a β-lactamase inhibitor, significantly decreased cross-protection. So, it would seem that Kp can also benefit from interaction with Ab by receiving protection from at least one antimicrobial.
We were also interested to see if this strain pair are more virulent in co-infections than mono-infections. For this, we used the model organism Galleria mellonella, an ethical, high-throughput alternative to mouse models7. Survival rates of caterpillars co-infected with Ab-Kp were significantly reduced compared to their mono-infected counterparts. The exact mechanism of increased virulence in co-infection is not yet known, however, it is intriguing and potentially dangerous in real world infection scenarios. The interactions between microbes become that much more complex in vivo as there is the added dimensions of how they are interacting with the host immune system and nutritional environment. This is an area we are interested in exploring further in the future.
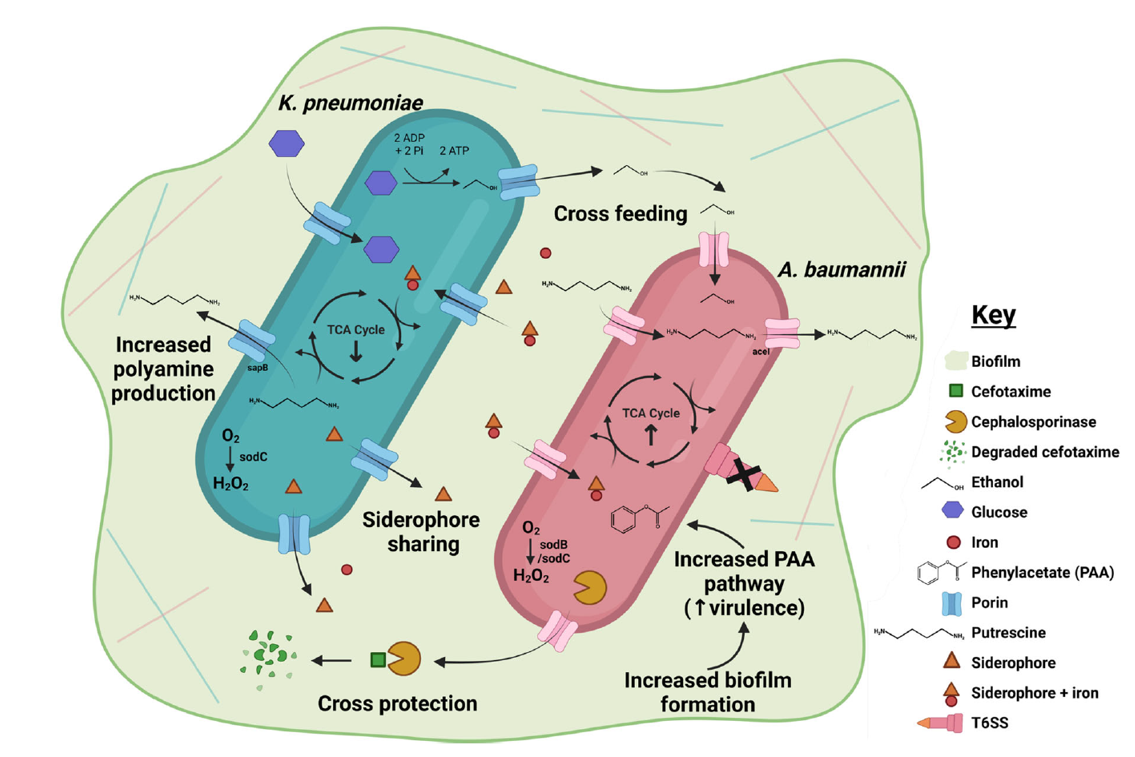
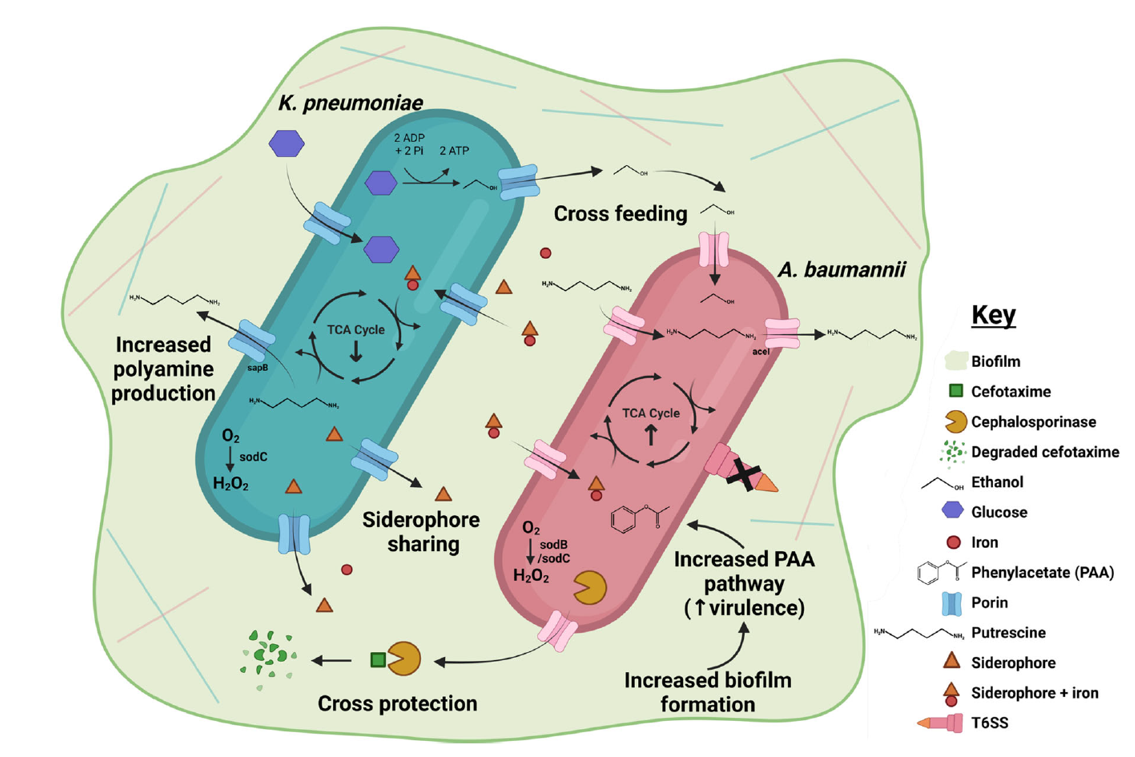
In a nutshell, this is the first time the potential interactions of Ab and Kp in co-infection have been investigated at the molecular level. We found evidence that Ab and Kp can participate in cross-feeding and cross-protection, and that co-infections are more lethal (at least in caterpillars). There were a number of other potential interactions that we uncovered in the full paper, graphically summarised below. There is a constant struggle in science between needing to simplify a system to control the variables and produce reproducible results, but also wanting to emulate the real-world setting as closely as possible, which is much more complex. There is so much more to learn about these two strains and how they might be interacting in an infection setting. I’m excited to know more about Ab and Kp interactions, and luckily for me, that is the focus of my PhD studies – so watch this space!
The Research Team

References
- Murray, J. L., Connell, J. L., Stacy, A., Turner, K. H. & Whiteley, M. Mechanisms of synergy in polymicrobial infections. J Microbiol 52, 188–199 (2014).
- Ibberson, C. B. & Whiteley, M. The social life of microbes in chronic infection. Current Opinion in Microbiology 53, 44–50 (2020).
- Aurora, A. et al. Recurrent bacteremia: A 10-year retrospective study in combat-related burn casualties. Burns 45, 579–588 (2019).
- Mammina, C. et al. Co-colonization with carbapenem-resistant Klebsiella pneumoniae and Acinetobacter baumannii in intensive care unit patients. Scandinavian Journal of Infectious Diseases 45, 629–634 (2013).
- Marchaim, D. et al. ‘Swimming in resistance’: Co-colonization with carbapenem-resistant Enterobacteriaceae and Acinetobacter baumannii or Pseudomonas aeruginosa. American Journal of Infection Control 40, 830–835 (2012).
- Rice, L. B. Federal Funding for the Study of Antimicrobial Resistance in Nosocomial Pathogens: No ESKAPE. The Journal of Infectious Diseases 197, 1079–1081 (2008).
- Dinh, H., Semenec, L., Kumar, S. S., Short, F. L. & Cain, A. K. Microbiology’s next top model: Galleria in the molecular age. Pathogens and Disease 79, ftab006 (2021).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in