Unique Transcriptional Signatures Correlate with Behavioral and Psychological Symptom Domains in Alzheimer’s Disease
Published in Neuroscience, Protocols & Methods, and General & Internal Medicine
What are Behavioral and Psychological Symptoms of Dementia (BPSD) and why are they important?
When it comes to dementia, most scientists are focused on the mechanisms, progression, and treatment of cognitive decline – and appropriately so, as cognitive deficits are distressing to patients and families. What a lot of people don’t know is that over 90% of patients with neurodegenerative dementias develop new-onset, not present-at-a-younger-age BPSD that can include anxiety, apathy, depression, agitation, aggression, hallucinations, and delusions. And this makes a lot of sense: If the brain is degenerating, and cognition is emergent from neural circuitry, then it’s no surprise that BPSD would also develop during neurodegeneration, as neural circuits also facilitate processes such as emotional reactivity, motivation and reward processing, and behavioral inhibition. BPSD are debilitating for patients, families, and caregivers. Independent of cognitive decline, these symptoms hasten morbidity and mortality, increase healthcare costs up to 50%, and speed up transition from independent living to assisted living facilities and memory units.
Despite the ubiquity and high burden of BPSD, molecular and cellular mechanisms of BPSD remain vastly understudied compared to cognition. This is concerning not only from a treatment standpoint – as only one medication, brexpiprazole, is FDA-approved for any BPSD – but also because BPSD can be the earliest presenting symptoms of neurodegenerative disease. By not studying BPSD early and often, we are missing opportunities to detect molecular and cellular changes towards the beginning of the disease process. With this newest publication, our group developed a pathway to investigate BPSD on the transcriptomic level, which we hope will inspire new lines of investigation into BPSD mechanisms.
Our Approach to Studying BPSD
In general, if a neuropsychiatric symptom can arise in younger adults, it can also arise de novo during neurodegeneration as BPSD. However, epidemiological studies have noted that BPSD tend to cluster in terms of presentation. For instance, mood symptoms like anxiety and depression co-present more often than depression and hallucinations.
We utilized a cohort from a memory clinic at Rush Alzheimer’s Disease Center (n ~200) who were diagnosed with Alzheimer’s Disease (neuropathologically confirmed) and had their BPSD measured by a trained psychiatric provider using a standardized questionnaire that was completed by conversing with a caregiver or loved one of the patient. The questionnaire was designed to track the frequency and severity of multiple behaviors related to BPSD. This approach is a substantial advantage compared to most larger cohorts (i.e. NACC-UDS, some ADNI phases, etc) and subsequent biobanks that rely on the NPI-Q, which has single question about each BPSD. Using an unbiased clustering analysis across these individual questions, we found symptoms grouped into four categories: Affective (depression and anxiety), Psychosis (predominantly hallucinations), Agitation (including aggression), and Apathy Domains. We created domain subscales, scoring all the questions that clustered together in terms of frequency and severity, and created a final score.
Then, we designated participants with the highest scores (>70th percentile) as cases and those with the lowest scores (<30th percentile) as controls in each domain. We did not include those with ambiguous scores between the 30th and 70th percentiles. With this design, an individual participant could be classified as a case in one domain and a control in another.
Once grouped, we isolated post-mortem tissue from the Anterior Cingulate Cortex, a region that has been linked to all four BPSD domains and Alzheimer’s Disease in neuroimaging studies. We chose tissue samples from 60 participants with the evenest mix of cases and controls, balanced for gender. We performed bulk RNA-seq and analyzed the data to pick out differentially expressed genes (DEGs), establish functional pathway enrichments, interrogate gene network co-expression, and explore if ‘hidden’ proteins might be affecting the flow of information through the transcriptional network.
What we found
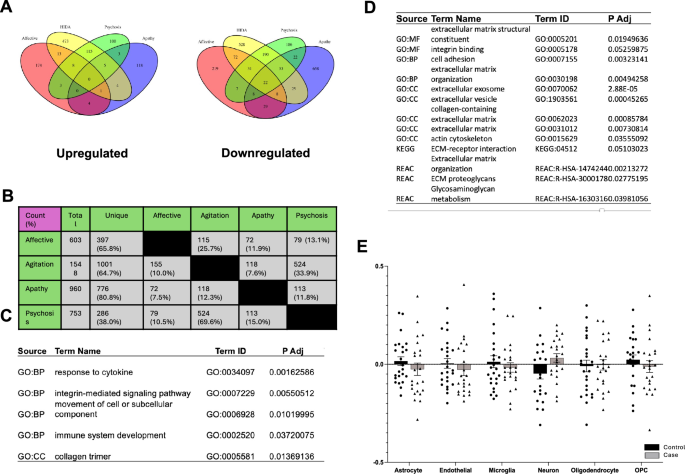
Despite our overlapping design, we found that the majority of the DEGs were unique to each domain, with the exception of agitation and psychosis, which are known to be highly co-morbid clinically. Though Rank-Rank Hypergeometric Overlap analyses suggested some general agreement in transcription across domains, our study design still allows for some differentiation of transcriptional signatures . Overall, all 4 BPSD domains were correlated with a ‘downregulation’ of gene expression.
Because there were four domains to analyze, there are a lot of details in the main paper, exploring Agitation predominantly, and the supplement, exploring the Affective, Apathy, and Psychosis Domains. Below, we describe some of the more interesting findings that we believe have the greatest potential for further study.
First, we performed cellular deconvolution to estimate the abundances of 6 cells types, and we noted a change for only one domain: a decrease in microglia associated with apathy cases. This difference in immune function was echoed in our co-expression analysis, where a module of co-transcribed genes was highly enriched for the innate immune system, including AD-related pathways such as TYROBP and the complement cascade. While these immune findings were striking, it was even more notable that 78 out of 80 known ribosomal subunits were downregulated. To our knowledge, no one has explored protein homeostasis nor innate immunity with apathy in Alzheimer’s Disease.
When analyzing information flow through the transcriptional network, ESR1 was noted as the highest rated impact gene mediating agitation. Coding for Estrogen Receptor-alpha, ESR1 is an important marker of cells in the periaqueductal grey and hypothalamus that facilitate aggression – which has been reproduced consistently. However, little is known about how ESR1 itself might facilitate aggression or agitation nor how its expression in the Anterior Cingulate Cortex might relate to these behaviors.
Finally, one of the transcriptional modules most strongly associated with the Agitation and Psychosis Domains was for extracellular matrix proteins. One interesting finding from another very well-done proteomic study found that changes in the extracellular matrix, which they called the “matrisome”, was associated with ApoE status and Alzheimer’s Disease pathology but NOT cognition. Whether this is a reinforcement of our findings or coincident is unknown.
Conclusion
Our study represents the first -omics study to associate molecular changes with the Affective, Apathy, and Agitation Domains in dementia. Despite its novelty, we would advocate that this study is best described as hypothesis-generating, as relatively low statistical power and the overlapping participant design should lead to caution in interpreting our findings. Our hope is that this study inspires others to look more critically at molecular and cellular mechanisms of BPSD and consider that dementia is much more than just cognitive decline.
Follow the Topic
-
Translational Psychiatry

This journal focuses on papers that directly study psychiatric disorders and bring new discovery into clinical practice.
Related Collections
With Collections, you can get published faster and increase your visibility.
From mechanism to intervention: translational psychiatry of childhood maltreatment
Publishing Model: Open Access
Deadline: Jun 30, 2026
Moving towards mechanism, causality and novel therapeutic interventions in translational psychiatry: focus on the microbiome-gut-brain axis
Publishing Model: Open Access
Deadline: May 19, 2026
Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in