What is in that vesicle?
Published in Protocols & Methods

Justifiably, many consider the cell the basic building block of higher organisms. By assembling a myriad of distinct cells into a tissue or organ, emergent properties coalesce to enable tissue function. Importantly, cells comprising tissue most often are chemically and morphologically diverse and may be responsible for different aspects of tissue or organ function. Due to the functional importance of cellular heterogeneity, there has been significant effort devoted to creating technologies to probe the spatial, temporal, and chemical nature of cells. However, approaches capable of creating the “parts list” of chemical compounds from an individual cell have lagged behind other characterization techniques, such as single-cell transcriptomics, making a comprehensive chemical inventory of the cell difficult to obtain.
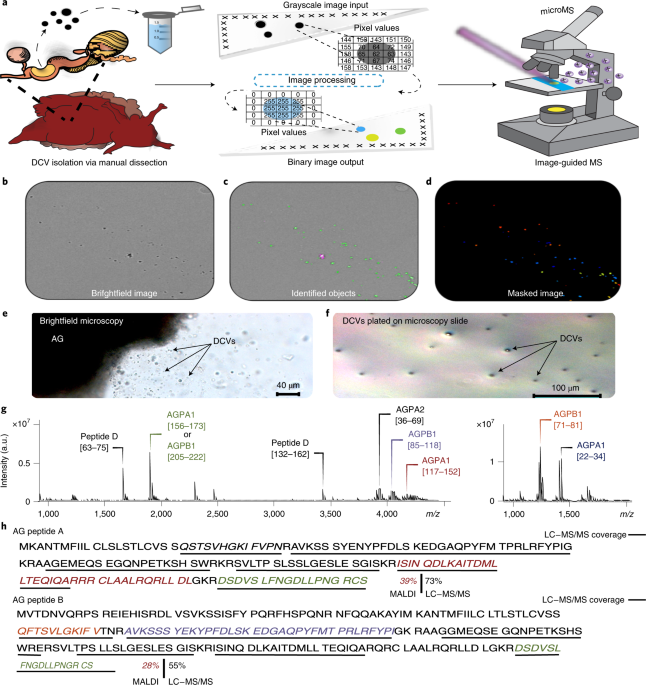
We understand that a cell is not a homogenous solution surrounded by its plasma membrane, but rather, contains a range of subcellular structures – organelles – that concentrate specific molecules into limited spatial domains, allowing coordinated control of cellular processes. Most organelles remain too small to characterize with mass spectrometry (MS). One important type of organelle found in endocrine, exocrine and neuronal tissues are the dense-core vesicles (DCVs), which are used to package and release a range of cell-to-cell signaling molecules. How small are they? As one example, the volume of a large DCV with an 800 nm diameter is about 300 aL; even for a molecule at the high concentration of 10 mM, the amount of material is only about 3 amol. That is not much material to start with when performing an MS measurement! Our biological model, Aplysia californica, provides a measurement advantage, in that the molecules of interest are contained by the vesicle membrane and thus, stay concentrated and localized throughout sample preparation. Using this model, our goal was to create a workflow approach that allows high-throughput sampling of individual vesicles for their bioanalytical characterization using MS.
Recently, we created a unique approach for assaying thousands of individual cells using matrix-assisted laser desorption/ionization (MALDI) MS (https://doi.org/10.1021/acs.analchem.9b01689). Briefly, we scatter the cells of interest onto a microscope slide, optically image the slide to map their physical locations, and then use MALDI to blast the cells and assay the resulting ions with a mass spectrometer. In our Nature Methods study (https://doi.org/10.1038/s41592-021-01277-2), we down-scaled this single-cell approach to measure single organelles – DCVs. This required optimizing several steps: improve our ability to target objects smaller than individual cells, optimize mass spectrometric sensitivity for subcellular analyte detection, and fine tune unsupervised data analysis workflows. Using our new high-throughput single-organelle MS approach, we detected both peptides and lipids in 0.5- to 2-μm-diameter DCVs and electron lucent vesicles (ELVs) isolated from the exocrine atrial gland and red hemiduct of Aplysia californica. Striking differences in the lipid content between the ELVs and DCVs were observed, as well as three chemically distinct types of DCVs.
One of the advantages of high-throughput single-organelle MS is that we don’t have to isolate and separate individual organelles. Rather, a population of isolated organelles can be deposited onto a glass slide and their specific spatial locations identified and selectively targeted according to their morphology using MS. We can select subpopulations on the slide for characterization based on their size, location, or fluorescence properties (for example, if they were labeled with a probe).
Now that we can measure objects that are smaller than a micron for their chemical contents, what is next? We will continue to optimize the approach to identify a greater range of molecules within these structures. From a measurement science perspective, we are moving to other subcellular structures such as mammalian extracellular vesicles and mitochondria. As the sensitivity of detection improves with next-generation mass spectrometers, we also expect to gain additional detail in the depth of chemical coverage, as well as throughput and flexibility in the subcellular MS analysis.
As a final comment, the individuals working on this project represent their home academic departments of Chemistry, Molecular and Integrative Physiology, and Bioengineering, demonstrating the unique contributions of each, ranging from sampling and measurement to data processing. This research was performed at the interdisciplinary Beckman Institute at the University of Illinois at Urbana -Champaign.
Follow the Topic
-
Nature Methods
This journal is a forum for the publication of novel methods and significant improvements to tried-and-tested basic research techniques in the life sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Methods development in Cryo-ET and in situ structural determination
Publishing Model: Hybrid
Deadline: Jul 28, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in